Abstract
More than 90% of animal embryos generated by reproductive cloning fail to survive to birth, primarily due to defects throughout the cloning process. The objective of this research is to define the key abnormalities in the mechanisms underlying DNA replication and chromosome segregation in reproductive cloning via somatic cell nuclear transfer, and to identify the reasons for their presence. Moreover, this research paper explores how these abnormalities impair genomic stability, resulting in developmental outcomes of the embryos. This literature review was conducted in order to address the growing innovations in cloning technologies and their efficiency, as well as expand the comprehensive understanding of the scientific mechanisms that determine cloning outcomes. This narrative review research paper analyzes relevant scientific sources found using database search engines such as Google Scholar and Pubmed, and points out key concepts addressing key DNA replication and chromosome segregation mechanisms along with their defects via SCNT. Moreover, it explains how genomic rearrangements negatively affect clones’ viability and efficiency and lead to the loss of genetic material or abnormal development in clones. Universally, it mentions DNA replication fork, DNA polymerase proofreading, mismatch repair and checkpoints, mitosis, mitochondria transfer via SCNT, bivalent remodeling, incomplete epigenetic reprogramming, proteins active in DNA replication, and more. Finally, this paper offers suggestions for newly established methods that can be implemented to improve reproductive cloning accuracy, mitigate risks of mutations, and ultimately aid in establishing the process as a safer and more widely applied technique to generate genetically equal, unvaried embryos.
Keywords: Somatic cell nuclear transfer, reproductive cloning, DNA replication, chromosome segregation, replication fork, polymerase proofreading, defects in embryo development, micronuclei formation, aneuploidy, spindle assembly checkpoint.
Introduction
Cloning is a process that allows the creation of exact genetic copies of cells, tissues, or organisms through scientific laboratory technologies and methods. Somatic cell nuclear transfer (SCNT) -based (see Figure 1) animal cloning truly first began with amphibian nuclear transfer in 1952 by Briggs and King1, followed by later advances culminating in Dolly the sheep in 19962. Recent advancements in cloning technologies, such as using transcription factors and epigenetic modifications, have enhanced the efficiency of embryo duplication, increasing protection against genetic disorders. Furthermore, grasping the potential of molecular mechanisms present in reprogramming somatic (somatic cells also refer to body cells) cell nuclei has led to advanced techniques of turning differentiated cells into pluripotent state stem cell technologies3. In the twenty-first century, investigators have explored variety of nuclear transfer processes. Pronuclear transfer involves swapping the two nuclei of fertilized embryos. Spindle transfer is when deoxyribonucleic acid (DNA) is transferred into an enucleated egg cell with preserved mitochondria4. Cloning of animals can potentially aid in species conservation and solve the ongoing issue of animal extinction5. Although the evaluations from those explorations have brought a significant facilities improvement, further research is mandatory to fully mitigate present welfare and security concerns.
Application of somatic cell nuclear transfer
In reproductive cloning, SCNT is a procedure where a single gene can be placed inside a tissue cell where it would replicate simultaneously with the developmental transition from an embryo into adult cells. Most of the cell division interventions occur during this process 6.
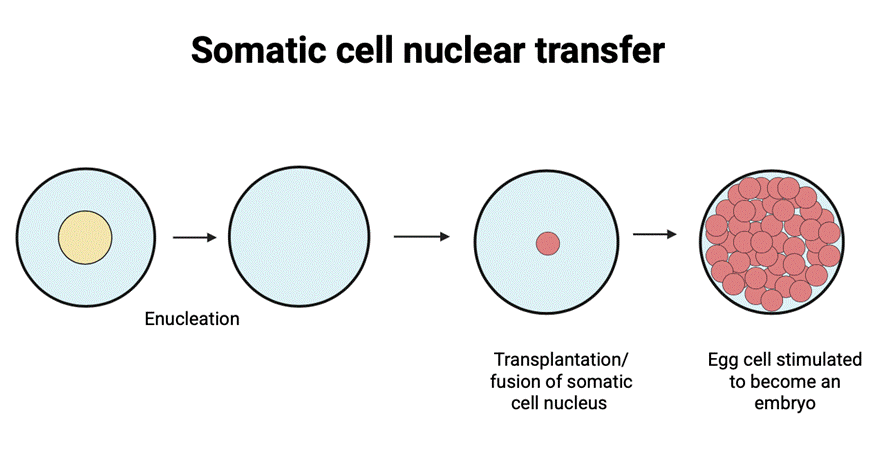
The injection of the somatic nucleus into the oocyte cytoplasm initiates programming and development of an embryo by histone replacement, DNA demethylation, premature chromosome condensation (PCC), chromatin remodeling, and transcriptional activation before zygotic genome activation (ZGA). These processes work together to mimic fertilization without meiosis7. Performing SCNT with genes that can aid in the treatment of a disease results in transgenic species and recombinant proteins that can be used for clinical procedures in biomedical science. By the fusion of the somatic nucleus and an enucleated oocyte, embryos are generated along with embryonic stem cells (NT-ESCs). Stem cells are now under research to be targeted to be immunocompatible to the patient to perform regenerative medicine, although this approach is currently experimental. These cells can be derived from the embryo by isolating and culturing inner cell mass at the blastocyst stage of the embryo. Stem cell clinical therapies are widely used, especially for COVID-19 or pulmonary diseases. In order to prevent immune rejection of the cells, a patient’s own somatic nuclei are being used to create an embryo from which the stem cells can be derived to help in successful cell transplantation8.
Pharmaceutical proteins can be generated by SCNT. Those are useful substances like hormones such as insulin and human growth hormone which are now even optimized in microbial hosts, like bacteria. Antithrombin III (ATIII) was introduced first. ATIII is generated from transgenic goat milk, targeting antithrombin deficiency and heparin resistance in human patients. Another example is Interferon derived from transgenic chicken eggs aiding sclerosis clinical therapy. Genes that code for the required proteins are being duplicated using vectors targeting a specific area of production, whilst being inserted into the transgenic species9.
This paper will cover reproductive cloning via SCNT, molecular mechanisms relevant to replication and segregation, and potential defects known to affect embryonic development. It will discuss the risks of those errors, the reasons for their occurrence, and potential strategies for their prevention. Overall, this article aims to investigate anomalies in the replication mechanisms and their impact on the efficiency of reproductive cloning.
Mechanisms in DNA replication and chromosome segregation in reproductive cloning
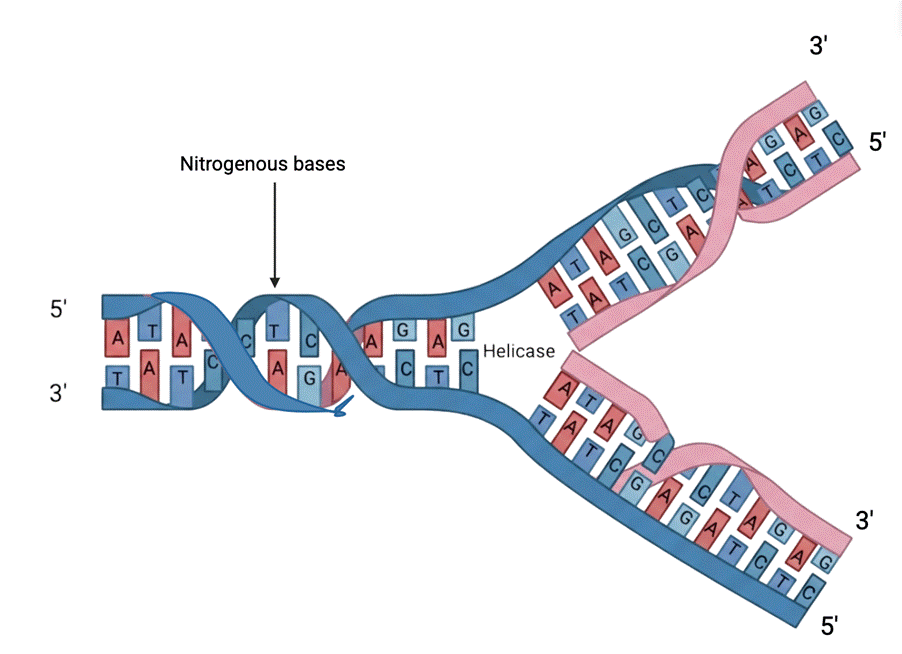
A nucleotide is a nucleoside molecule bound to one or more phosphate groups through ester linkages to the sugar moiety. Both DNA and ribonucleic acid (RNA) consist of polymers composed of nucleotides. Base-pairing is a template-based mechanism for DNA replication in which genetic material is copied into a complementary DNA sequence. Each nucleotide in the strand is recognized and assembled with a complementary nucleotide. This process causes the two strands of the helix to separate. This forms the hydrogen bonds, setting it for enzyme-catalyzed polymerization of a fresh chain. Each daughter strand inherits one parental strand and one freshly-produced strand in the double helix. This progressing structure then forms a “Y” shape which is called the “replication fork” which moves along the parental strand. In that region, the daughter strand genetic material is synthesized and copied by a mix of enzymes that contain DNA polymerase. The daughter cell strand that is being synthesized continuously is called the “leading” strand, and the one that is processed discontinuously is called the “lagging” strand. The direction in the development of the “lagging” strand is opposite to general chain growth. It is delayed due to discontinuous synthesis of the lagging strand before the maturation and ligation of Okazaki fragments (newly synthesized strands’ fragments), between the lagging and template strand (Figure 2). This “backstitching” mechanism requires “5′-to-3′ type” DNA polymerase10.
The development of the embryo in SCNT involves the duplication of the genome during the synthesis (S), during interphase before mitosis (figure 3), which uses the replication forks until the blastocyst stage. The sequence of the replication fork can determine the overall pace of genome duplication: if the replication works at a slower pace, the accuracy and efficiency of the reprogramming is improved. Slower replication allows more time for error correction, preventing replication-induced errors that compromise embryo development11.
Reprogramming takes place in the cytoplasm of the egg, erasing previous somatic cell epigenetic memory in order to then facilitate totipotent development12.
The initial proofreading mechanism is the DNA polymerase proofreading that occurs prior to the addition of a new nucleotide to the elongating chain. Furthermore, following nucleotide binding, but preceding the covalent addition of the nucleotide to the elongating chain, the enzyme must experience a conformational alteration. Only the suitable nucleotides can base-pair with the template strand. This enables a “double-check” reaction before the bond occurs. In scarce cases where this check fails to match the correct nucleotide, exonucleolytic proofreading is impelled to occur. That means that if the last nucleotide at the 3’ hydroxyl group (3’-OH) end is unsuitable, the chain cannot be extended. Therefore, the exonuclease activity takes place in a different subunit. This site removes the mismatched nucleotides from the 3’ end of the primer strand in a 3’ to 5’ direction until it is fully restored. As a result, DNA polymerase acts as a “self-correcting” enzyme. This process greatly prevents uncontrollable reproduction of cells (uncontrolled cell proliferation), which could lead to cancer. On the other hand, RNA polymerase (RNAP) is capable of initiating transcription without a primer and it doesn’t need proofreading since mistakes don’t get passed on during the mechanism because they are temporary. DNA polymerase adds nucleotides only in a 5’ to 3’ direction because if it were done in the opposite direction, the elimination of a mismatch would leave a bare 5’ end, which would permanently stop the development of the chain. The incoming nucleotide contains triphosphate, essential to complete bonding. Overall, this makes 5’ to 3’ synthesis favorable in terms of efficiency10.
In SCNT, weakened proofreading can lead to replication fork stalling and generate lagging strands that create micronuclei as the fork collapses13. This explains the implantation failure of cloned embryos. A practical piece of evidence can be observed where more than 90% of mouse embryos cloned through SCNT showed abnormal segregation, causing implantation failure rates due to the replication stress through damage to the fork. This study features the demand for strategies that tackle replication stress14.
Since the strands in the double helix are so tightly attached together, there is a need for the implementation of DNA helicase enzymes and single-strand DNA-binding proteins in order to copy polymerases and primers. Those work together in front of the replication fork. Helicase enzymes operate by breaking hydrogen bonds, utilizing adenosine triphosphate (ATP) energy in order to move along the chain. Two helicases work on both strands; the CMG helicase encircles and translocates along the leading-strand template at each fork, coordinating with leading-strand polymerase epsilon (Pol ![]() ). A single CMG complex per fork advances, coordinating with leading-strand Pol
). A single CMG complex per fork advances, coordinating with leading-strand Pol ![]() ; the CMG helicase encircles and translocates along the leading-strand template at each fork. A single CMG complex per fork advances15. A single CMG complex per fork advances, whereas single-stranded DNA-binding helix-destabilizing protein (SSB), ensures that the strands keep separated throughout the operation. They don’t cover bases of the helix so that the genetic information remains detectable10.
; the CMG helicase encircles and translocates along the leading-strand template at each fork. A single CMG complex per fork advances15. A single CMG complex per fork advances, whereas single-stranded DNA-binding helix-destabilizing protein (SSB), ensures that the strands keep separated throughout the operation. They don’t cover bases of the helix so that the genetic information remains detectable10.
Linking back to SCNT, epigenetic marks cause breakage of the double-strand, followed by fragmentation of chromosomes16. This causes high preimplantation loss rates as mammalian embryo development is highly dependent on stable chromosomes17.
DNA polymerase remains strongly bound to the sliding clamp of the leading strand, producing fusion. Conversely, once polymerase reaches the edge of the Okazaki fragment in the lagging strand – it loops back and moves on to a fragment of the next RNA primer which is made by primase. This creates a recycling procedure, while sustaining the attachment to the sliding clamp which keeps polymerase attached during synthesis, forming a ring10.
In general, all of the proteins active in DNA replication form a multienzyme complex, which acts altogether simultaneously, powered by nucleoside triphosphate hydrolysis. They work at a unified unit with a molecular weight exceeding 1 million daltons. Together, helicase and primase form a primosome to prime the fragments. During lagging-strand synthesis, left behind Okazaki fragments still contain RNA primers at 5’ ends, which are later removed by assigned enzymes behind the replication fork. Moreover, strand-directed mismatch repair amends errors missed by exonuclease. Ultimately, mutator genes increase mutation rates if defective. The role of topoisomerases in genetic data replication is to prevent tangling of DNA. Completing a rotation of the full chromosome every 10 base pairs requires a notable amount of energy. However, topoisomerases are capable of easing the torsional stress. In the single-strand breaks, a transient nick is introduced, rotating the helix. As a consequence, the permission of localized unwinding before the fork relieves tension. A covalent bond is formed, preserving the enzyme with the DNA for future reaction, without the need for additional ATP. Contrarily, a double-strand break constructs a “gate” allowing another fragment to pass through. It uses ATP hydrolysis in order to cut one double helix. This process aids in separating daughter strands after replication, ultimately separating chromosomes. Mutants that lack functional topoisomerase in the double-strand break acquire intertwined DNA because they fail to isolate chromosomes.10.
Topoisomerase II deficiency causes flawed segregation of catenated chromosomes18. The rate of catenanes (entangled chromosomes) that form anaphase bridges—threads of DNA stretching between segregating chromosomes during anaphase of mitosis—has a causal relationship with the failure of proper development to the morula (8-cell) stage in mouse SCNT embryos. This results in an inhibited postimplantation progression14.
Replication proteins from the multienzyme complex undergo posttranslational modifications that alter their activity and interactions—these play critical roles in responding to replication stress. This can occur in case of early development of diseases19.
The spindle assembly checkpoint (SAC) is crucial to verify correct chromosome attachment to microtubules during mitosis, thereby preventing errors in the cloning process20. Cohesin complexes hold sister chromatids together, enabling controlled cleavage of cohesin by separase. This is done by even distribution of chromatids to daughter cells21.
Polymerase proofreading and mismatch repair are crucial mechanisms for healthy and stable embryo implantation22. Proofreading of DNA is an activity taking place during DNA replication where an enzyme checks each nucleotide during DNA replication and deals with the mismatched ones. Mismatch repair and checkpoints of DNA, occurring afterwards, are protective responses detecting any remaining DNA damage, which maintains correctly replicated DNA strands in the cell cycle. This prevents errors from being passed on to daughter cells23.
Mitochondria, passed on through vertical inheritance, are multirole organelles found in eukaryotic cells. They are responsible for inner-cell functions such as energy metabolism, cell signaling, immunity, apoptosis and proliferation moderation24. In the SCNT process, mitochondria DNA (mtDNA) from the somatic donor cell and from the recipient oocyte is mixed. The somatic cell mitochondria remain mixed with the second mitochondria after nuclear transfer. This results in SCNT embryos being heteroplasmic. Introduced variability can disrupt normal cellular metabolism in embryos. The somatic cell mitochondria can remain along with the second mitochondria after nuclear transfer25.
Genetic defects known to affect embryonic development and genomic stability
To maintain genomic stability, there are several mechanisms responsible for the replication of genetic information and the division of chromosomes into a pair of daughter cells.
Sometimes, anomalies occur in polymerase proofreading and mismatch repair and checkpoints which often lead to potentially harmful genetic mutations. Disruptions lead to replication stress, which is the slowing of the replication fork progression stressed by both exogenous and endogenous origins. Defects in lagging-strand maturation can lead to nicks, double-strand breaks, and genomic instability in SCNT-derived embryos. Those origins could vary from oncogene activation, depletion of nucleotide pools and transcription-replication conflicts to inherently difficult-to-replicate regions. Other examples are aneuploidy, as well as epigenetic abnormalities, resulting in disease development26.
These factors cause genomic instability which results in long term developmental effects and diseases in cloned products. Analysis of the reasons for anomalies can aid the global technical improvement in cloning from cells to various organisms27.
Aneuploidy is incorrect duplication of chromosomes, leading to extra or missing chromosomes due to their abnormalities as well as micromanipulation stress which is accidental physical damage of cellular components of the oocyte12,28,29.
Incomplete epigenetic reprogramming is a paramount cause of low cloning efficiency. Epigenetics involve insufficient elimination of epigenetic marks such as DNA methylation and histone modifications which can be heritable in SCNT. Epigenetics forms disruptions in gene expression mechanisms that affect the way genes are received and expressed30.
Recent studies reveal that spontaneous harmful gene mutations, trinucleotide repeats, chromosomal rearrangements, and aneuploidy occur more frequently in somatic cells than previously believed. Those mutations are closely linked to chromosomal rearrangements and affect clones’ efficiency31.
SCNT disrupts spindle assessment checkpoint, which is the reason for kinetochore-microtubule misalignment in mitosis after activation. The kinetochores that fail to link to all necessary SAC proteins trigger premature anaphase. This results in chromosome missegregation where fragments or entire chromosomes are excluded from the daughter cell nuclei. Those are the conditions for formation of micronuclei. Therefore, SAC failure contributes to aneuploidy and reduces blastocyst viability32.
Abnormal chromosome segregation is a primary factor contributing to developmental failures in cloned embryos. Research on SCNT embryos indicates that chromosome misalignment and lagging chromosomes cause the formation of micronuclei, which greatly diminish embryo viability, abnormal growth. The health of embryos is inhibited by immune activation that destroys cells or raise risk of cancer with nuclear envelope rapture apoptosis and repair failure, causing more mutations33,34. Treatment with Latrunculin A has been shown to decrease these errors35.
Another consequence of imperfect chromatin reprogramming is cohesion dysfunction: the somatic nucleus remains in adult cell organisation and doesn’t completely reorganize into the embryo. This is caused by premature chromosome condensation: the somatic nucleus condenses similarly to mitosis, so embryonic cohesin is not able to replace somatic cohesin, which is a protein that holds identical copies of each chromosome together. This activates the wrong genes as well as subsequently causes chromosomes to be organised too closely together in an insufficient structure36,37,38,39.
Consequences of errors in bivalent remodeling might generate chromosome missegregation, propagating embryonic mortality or birth defects. This is how severe consequences begin40. Bivalent chromatin remodeling is a process in embryonic stem cells where certain ATP-dependent complexes, like INO80, esBAF (a type of SWI/SNF), and NuRD, control bivalent domains. These domains are promoters marked by both active Histone H3 Lysine 4 trimethylation (H3K4me3) and repressive Histone H3 Lysine 27 Trimethylation (H3K27me3) histone modifications. This combination keeps developmental genes in a ‘poised’ state, meaning they are either barely expressed or silent in undifferentiated stem cells to maintain pluripotency, but are ready to be activated or silenced during differentiation41,42. This process includes moving nucleosomes, swapping histone variants such as H2A.Z or H3.3, and selective targeting. Together, these steps create “bivalent specificity,” which provides the precise epigenetic control needed for early embryogenesis, as shown in mouse peri-implantation stages43. Another example is shown by statistical evidence of cloned pregnancies in cattle encountering up to 50% of embryonic loss due to placental abnormalities or improper fetomaternal interactions44.
Some studies suggest that sometimes transcriptional processes avert the detrimental effects of nascent transcripts which have potential in rehybridizing with the template DNA that may result in DNA strand rearrangements. In order to maintain general genetic stability, organisms seek to avoid the harmful interaction of nascent RNA with DNA during transcription. Transcription-induced instability might lead to disease. RNAP II transcripts collaborate with RNA in the nucleus, including capping, splicing, and polyadenylation, prior to maturing into messenger RNA (mRNA) ready for cytoplasmic translation. The coupling of transcription and RNA processing ensures the efficiency and accuracy of gene expression45.
To conclude, experts believe that SCNT remains an inefficient process. The average efficiency to produce a viable offspring is between 5% and 10% due to the range of abnormalities that this article covered46.
Conclusion
This article explores cloning of the embryo via SCNT, along with a range of molecular mechanisms involved in replication and segregation such as the replication fork, DNA polymerase proofreading, that involve helicase SSB, polymerase, and topoisomerase enzymes. The narrative research explored the SCNT process in detail. This focused analysis underscores various abnormalities in replication mechanisms. These mechanisms include replication fork stalling, persistent catenanes, and incomplete decatenation. In these instances inaccuracies occur in following chromosome segregation mechanisms like kinetochore-microtubule attachment failure, segregation or replication defect, polymerase proofreading, anaphase bridges, micronuclei formation, lagging chromosomes and unequal chromosome distribution, incomplete epigenetic reprogramming, and SAC failure. These defects collectively generate gene mutations and aneuploidy which contributes to an impaired offspring, resulting in current low rates of clone viability with SCNT.
It points out that identical genomes are produced by DNA replication, followed by chromosome segregation, where they are split into daughter cells. This information about various findings on the topic will also be useful for other researchers or students researching reproductive cloning and errors that affect the development of embryos. It is necessary to acknowledge that the current body of knowledge on cloning remains limited. Technical risks and safety constraints have prevented the full exploration of cloning’s potential. However, there is ongoing further research on how reproductive cloning can be applied safely and efficiently. Possible future directions include: high-fidelity reprogramming, which is accurate epigenetic reprogramming of the somatic donor nucleus to a totipotent embryonic state47; Live-Cell Imaging of spindle and kinetochore dynamics in SCNT to reveal real-time chromosome segregation errors48.
Single-cell multi-omics analyses in SCNT profile the genome, epigenome, transcriptome, and replication timing from individual cloned embryo cells. This approach maps catenation-induced transcriptional chaos in arrested blastomeres49. There is a growing possibility of turning the skill of producing direct copies of an embryo into a mastery that is globally practiced if more techniques to control abnormalities are invented. This would to help individuals, ultimately striving for a biotechnology revolution. Solving these replication-segregation bottlenecks could make SCNT clinically viable and move it from an experimental stage to a practical biotechnology.
References
- J. B. Gurdon, J. A. Byrne. The first half-century of nuclear transplantation. Proceedings of the National Academy of Sciences of the United States of America. Vol. 100, pg. 8048–8052, 2003 https://doi.org/10.1073/pnas.1337135100 [↩]
- J. B. Gurdon, I. Wilmut. Nuclear transfer to eggs and oocytes. Cold Spring Harbor Perspectives in Biology. Vol. 3, pg. a002659, 2011 https://doi.org/10.1101/cshperspect.a002659 [↩]
- S. R. Singh, P. K. Gupta. Application of reproductive biotechnology in animals: implications and potentials: applications of reproductive cloning. ScienceDirect. https://www.sciencedirect.com/science/article/abs/pii/S0378432003001611?via%3Dihub, 2003 [↩]
- M. H. Craven, N. T. Taylor, D. T. Herbert. Pronuclear transfer in human embryos to prevent transmission of mitochondrial DNA disease. Nature. 14 April 2010. https://www.nature.com/articles/nature08958 [↩]
- G. Bonetti et al. Human cloning: Biology, ethics and social implications. Clinica Terapeutica. 2023, doi: 10.7417/CT.2023.2492, https://clinicaterapeutica.it/ojs/index.php/1/article/view/806/581 [↩]
- J. W. Gordon, F. H. Ruddle. DNA-mediated genetic transformation of mouse embryos and bone marrow–a review. Gene. Vol. 33, pg. 121–136, 1985 https://doi.org/10.1016/0378-1119(85)90087-3 [↩]
- S. Matoba, Y. Zhang. Somatic cell nuclear transfer reprogramming: mechanisms and applications. Cell Stem Cell. Vol. 23, pg. 471–485, 2018, https://doi.org/10.1016/j.stem.2018.06.018 [↩]
- B. Singh, G. Mal, V. Verma, R. Tiwari, M. I. Khan, R. K. Mohapatra, S. Mitra, S. A. Alyami, T. B. Emran, K. Dhama, M. A. Moni. Stem cell therapies and benefaction of somatic cell nuclear transfer cloning in covid-19 era. Stem Cell Research & Therapy. Vol. 12, pg. 283, 2021 https://doi.org/10.1186/s13287-021-02334-5 [↩]
- L.-M. Houdebine. Production of pharmaceutical proteins by transgenic animals. Comparative Immunology, Microbiology and Infectious Diseases. Vol. 32, pg. 107–121, 2009 https://doi.org/10.1016/j.cimid.2007.11.005 [↩]
- B. Alberts, A. Johnson, J. Lewis, M. Raff, K. Roberts, P. Walter. DNA replication mechanisms. Molecular Biology of the Cell. NCBI. https://www.ncbi.nlm.nih.gov/books/NBK26850/#A779, 2002. [↩] [↩] [↩] [↩] [↩]
- G. del Solar, R. Giraldo, M. J. Ruiz-Echevarría, M. Espinosa, R. Díaz-Orejas. Replication and control of circular bacterial plasmids. Microbiology and Molecular Biology Reviews. Vol. 62, pg. 434–464, 1998 https://doi.org/10.1128/mmbr.62.2.434-464.1998 [↩]
- C. Gouveia, C. Huyser, D. Egli, M. S. Pepper. Lessons learned from somatic cell nuclear transfer. International Journal of Molecular Sciences. Vol. 21, pg. 2314, 2020 https://doi.org/10.3390/ijms21072314 [↩] [↩]
- S. A. Sabatinos, N. S. Ranatunga, J.-P. Yuan, M. D. Green, S. L. Forsburg. Replication stress in early s phase generates apparent micronuclei and chromosome rearrangement in fission yeast. Molecular Biology of the Cell. Vol. 26, pg. 3439–3450, 2015 https://doi.org/10.1091/mbc.E15-05-0318 [↩]
- E. Mizutani, K. Yamagata, T. Ono, S. Akagi, M. Geshi, T. Wakayama. Abnormal chromosome segregation at early cleavage is a major cause of the full-term developmental failure of mouse clones. Developmental Biology. Vol. 364, pg. 56–65, 2012 https://doi.org/10.1016/j.ydbio.2012.01.001 [↩] [↩]
- Z. Yuan, R. Georgescu, L. Bai, D. Zhang, H. Li, M. E. O’Donnell. DNA unwinding mechanism of a eukaryotic replicative cmg helicase. Nature Communications. Vol. 11, pg. 688, 2020 https://doi.org/10.1038/s41467-020-14577-6 [↩]
- B. Xu, Z. Sun, Z. Liu, H. Guo, Q. Liu, H. Jiang, Y. Zou, Y. Gong, J. A. Tischfield, C. Shao. Replication stress induces micronuclei comprising of aggregated dna double-strand breaks. PLoS ONE. Vol. 6, pg. e18618, 2011 https://doi.org/10.1371/journal.pone.0018618 [↩]
- N. Wang, S. Xu, D. Egli. Replication stress in mammalian embryo development, differentiation, and reprogramming. Trends in Cell Biology. Vol. 33, pg. 872–886, 2023 https://doi.org/10.1016/j.tcb.2023.03.015 [↩]
- K. C. Nitiss, A. Bandak, J. M. Berger, J. L. Nitiss. Genome instability induced by topoisomerase misfunction. International Journal of Molecular Sciences. Vol. 25, pg. 10247, 2024 https://doi.org/10.3390/ijms251910247 [↩]
- Song, Hao-Yun et al. “DNA replication: Mechanisms and therapeutic interventions for diseases.” MedComm vol. 4,1 e210. 5 Feb. 2023, doi:10.1002/mco2.210 https://pubmed.ncbi.nlm.nih.gov/36776764/ [↩]
- Mashiko, Daisuke, et al. Chromosome segregation error during early cleavage in mice pre-implantation embryos does not necessarily cause developmental failure after blastocyst stage. Scientific Reports. Vol. 10, pg. 854, 2020. https://www.nature.com/articles/s41598-020-57817-x. [↩]
- Makrantoni, Vasso, and Adele L Marston. “Cohesin and chromosome segregation.” Current biology : CB vol. 28,12 (2018): R688-R693. doi:10.1016/j.cub.2018.05.019 https://pmc.ncbi.nlm.nih.gov/articles/PMC6013277/. [↩]
- J. Smith, L. Johnson. Double-checking chromosome segregation. PubMed. 1 May 2023, https://pubmed.ncbi.nlm.nih.gov/37017932/. [↩]
- R. R. Iyer, M. C. Pluciennik, M. S. Burdett, R. Modrich. DNA mismatch repair: functions and mechanisms. Chemical Reviews. Vol. 106, 2, 302-323, 2006. [↩]
- Borcherding, Nicholas, and Jonathan R Brestoff. “The power and potential of mitochondria transfer.” Nature vol. 623,7986 (2023): 283-291. doi:10.1038/s41586-023-06537-z. [↩]
- S. Hiendleder. Mitochondrial dna inheritance after scnt. Advances in Experimental Medicine and Biology. Vol. 591, pg. 103–116, 2007 https://doi.org/10.1007/978-0-387-37754-4_8 [↩]
- A. Zeman, K. Cimprich. Hallmarks of DNA replication stress. PubMed. 16 June 2022, https://pubmed.ncbi.nlm.nih.gov/35714587/. [↩]
- A. Tubbs, A. Nussenzweig. Endogenous DNA damage as a source of genomic instability in cancer. Cell. Vol. 168, 4, 644-656, 2017, doi:10.1016/j.cell.2017.01.002. [↩]
- D. A. Compton. Mechanisms of aneuploidy. Current Opinion in Cell Biology. Vol. 23, pg. 109–113, 2011 https://doi.org/10.1016/j.ceb.2010.08.007. [↩]
- Y. Dai, L. Wang, H. Wang, Y. Liu, N. Li, Q. Lyu, D. L. Keefe, D. F. Albertini, L. Liu. Fate of centrosomes following somatic cell nuclear transfer (scnt) in bovine oocytes. Reproduction. Vol. 131, pg. 1051–1061, 2006 https://doi.org/10.1530/rep.1.01058. [↩]
- Y. Li, S. Sun, Y. Xu, J. Zhang, Y. Du, Y. Cao, Z. Liao, Y. Xie, X. Bian, J. Huang, M. Wang, Z. Liu, Q. Sun, F. Lu. Efficient somatic cell nuclear transfer by overcoming both pre- and post-implantation epigenetic barriers. Advanced Science. Vol. 12, pg. e04669, 2025, https://doi.org/10.1002/advs.202504669 [↩]
- Azad, Priti, and R. C. Woodruff. “Mutation and cloning efficiency.” Cloning and stem cells 8.4 (2006): 237-239. https://www.liebertpub.com/doi/pdf/10.1089/clo.2006.8.237 [↩]
- P. Lara-Gonzalez, J. Pines, A. Desai. Spindle assembly checkpoint activation and silencing at kinetochores. Seminars in Cell & Developmental Biology. Vol. 117, pg. 86–98, 2021, https://doi.org/10.1016/j.semcdb.2021.06.009 [↩]
- M. Kwon, M. L. Leibowitz, J.-H. Lee. Small but mighty: the causes and consequences of micronucleus rupture. Experimental & Molecular Medicine. Vol. 52, pg. 1777–1786, 2020, https://doi.org/10.1038/s12276-020-00529-z. [↩]
- K. Utani, Y. Kohno, A. Okamoto, N. Shimizu. Emergence of micronuclei and their effects on the fate of cells under replication stress. PLoS ONE. Vol. 5, pg. e10089, 2010, https://doi.org/10.1371/journal.pone.0010089. [↩]
- Y. Terashita, S. Wakayama, T. Yamagata, M. Wakayama. Latrunculin A treatment prevents abnormal chromosome segregation for successful development of cloned embryos. PLoS ONE. Vol. 8, 10, e78380, 2013, https://journals.plos.org/plosone/article?id=10.1371%2Fjournal.pone.0078380. [↩]
- J. Xiang, Y. Lai, Z. He. The functions and mechanisms of the cohesin complex in regulating the fate determinations of stem cells. Research. Vol. 8, pg. 0757, 2025, https://doi.org/10.34133/research.0757 [↩]
- P. L. Schuck, L. E. Ball, J. A. Stewart. The dna-binding protein cst associates with the cohesin complex and promotes chromosome cohesion. Journal of Biological Chemistry. Vol. 297, pg. 101026, 2021 https://doi.org/10.1016/j.jbc.2021.101026 [↩]
- A. Pezzotta, M. Mazzola, M. Spreafico, A. Marozzi, A. Pistocchi. Enigmatic ladies of the rings: how cohesin dysfunction affects myeloid neoplasms insurgence. Frontiers in Cell and Developmental Biology. Vol. 7, pg. 21, 2019 https://doi.org/10.3389/fcell.2019.00021 [↩]
- M. Solé-Ferran, A. Losada. Cohesin in 3d: development, differentiation, and disease. Genes & Development. Vol. 39, pg. 679–696, 2025, https://doi.org/10.1101/gad.352671.125 [↩]
- Láscarez-Lagunas, Laura I et al. “Loss, Gain, and Retention: Mechanisms Driving Late Prophase I Chromosome Remodeling for Accurate Meiotic Chromosome Segregation.” Genes vol. 13,3 546. 2022, doi:10.3390/genes13030546 [↩]
- A. Harikumar, E. Meshorer. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Reports. Vol. 16, pg. 1609–1619, 2015, https://doi.org/10.15252/embr.201541011 [↩]
- D. C. Klein, S. J. Hainer. Chromatin regulation and dynamics in stem cells. Current Topics in Developmental Biology. Vol. 138, pg. 1–71, 2020, https://doi.org/10.1016/bs.ctdb.2019.11.002 [↩]
- Y. Li, J. He, Y. Liu, Y. Hui, S. Liu, Y. Zhang, Y. Xiong, T. Xu, Z. Xu, Z. Zhang, Y. Zhang, G. Yang, J. Zhao, D. Bai, X. Lei, X. Kou, Y. Zhao, J. Du, Z. Guo, J. Yin, X. Zhang, C. Xu, Y. Gao, M. Chen, H. Wang, C. Jiang, S. Gao, W. Liu. Remodelling bivalent chromatin is essential for mouse peri-implantation embryogenesis. Nature Cell Biology. Vol. 27, pg. 1797–1811, 2025, https://doi.org/10.1038/s41556-025-01776-w [↩]
- Hill, Jonathan. (2014). Incidence of Abnormal Offspring from Cloning and Other Assisted Reproductive Technologies. Annual review of animal biosciences. 2. 307-21. 10.1146/annurev-animal-022513-114109. https://www.researchgate.net/publication/268228501_Incidence_of_Abnormal_Offspr ing_from_Cloning_and_Other_Assisted_Reproductive_Technologies [↩]
- Li, Xialu, and James L. Manley. “Cotranscriptional processes and their influence on genome stability.” Genes & development 20.14 (2006): 1838-1847. https://genesdev.cshlp.org/content/20/14/1838.full [↩]
- C. Gouveia, C. Huyser, D. Egli, M. S. Pepper. Lessons learned from somatic cell nuclear transfer. International Journal of Molecular Sciences. Vol. 21, pg. 2314, 2020, https://doi.org/10.3390/ijms21072314 [↩]
- Y. Li, Q. Sun. Epigenetic manipulation to improve mouse scnt embryonic development. Frontiers in Genetics. Vol. 13, pg. 932867, 2022, https://doi.org/10.3389/fgene.2022.932867 [↩]
- B. J. Howell, B. Moree, E. M. Farrar, S. Stewart, G. Fang, E. D. Salmon. Spindle checkpoint protein dynamics at kinetochores in living cells. Current Biology: CB. Vol. 14, pg. 953–964, 2004, https://doi.org/10.1016/j.cub.2004.05.053 [↩]
- J. R. Hernandez Mora, C. Buhigas, S. Clark, R. Del Gallego Bonilla, D. Daskeviciute, A. Monteagudo-Sánchez, M. E. Poo-Llanillo, J. V. Medrano, C. Simón, M. Meseguer, G. Kelsey, D. Monk. Single-cell multi-omic analysis profiles defective genome activation and epigenetic reprogramming associated with human pre-implantation embryo arrest. Cell Reports. Vol. 42, pg. 112100, 2023, https://doi.org/10.1016/j.celrep.2023.112100 [↩]