Abstract
This study examines the application of convolutional neural networks (CNNs) to predict the pathogenicity of single nucleotide variants (SNVs) and small insertions/deletions (Indels) in human DNA. A CNN was trained on two-dimensional image representations of DNA sequences from the ClinVar database, employing one-hot encoding within a window of 101 bp. The model was evaluated on a held-out test set and via five-fold stratified cross-validation, achieving an F1-score of 0.92 [95% CI: 0.89–0.95], ROC-AUC of 0.96 [0.94–0.98], and PR-AUC of 0.94 [0.91–0.96]. Grad-CAM visualization provided interpretable predictions by highlighting sequence regions associated with pathogenicity. Direct comparison with SIFT, PolyPhen-2, and CADD on the same test set is reported. Key limitations include exclusive focus on SNVs and Indels, under-representation of non-European variant populations, and absence of multi-omic features. GeneSpectra, a web application built with React and Next.js, facilitates deployment for VUS triage and diagnostic decision-making.
Introduction
Accurate and efficient prediction of the pathogenicity of genetic variants is principal in modern clinical genomics1. The ability to reliably classify single nucleotide variants (SNVs) and small insertions/deletions (Indels) as either pathogenic or benign is crucial for informed diagnostic and therapeutic decisions. However, a significant challenge persists in the form of variants of uncertain significance (VUS)1, which represent a considerable proportion of clinically identified variants and pose a significant obstacle to precise genomic interpretation. Current bioinformatics pipelines, while effective in many instances, often suffer from limitations in transparency and scalability, hindering their widespread adoption and practical utility in clinical settings1,2. These methods frequently rely on established databases and rule-based systems3, potentially overlooking subtle yet crucial patterns within the sequence data that might be indicative of pathogenicity. Furthermore, the lack of easily interpretable outputs often necessitates substantial manual review by expert clinicians, thwarting them in the diagnostic process.
This study applies deep learning, specifically Convolutional Neural Networks (CNNs)4,5, to improve the accuracy and interpretability of pathogenicity prediction. Rather than claiming superiority over prior methods, CNN performance is reported on 2D image-encoded DNA sequences and compared directly with established predictors on the same test set. Estimates suggest that 30–60% of variants identified in clinical exome sequencing are classified as VUS1’ a single misclassified VUS—for example a BRCA1 variant incorrectly called benign—can delay life-saving intervention or trigger unnecessary prophylactic surgery, illustrating the direct patient cost of this ambiguity. The model is trained on SNVs and Indels from ClinVar using one-hot 2D image encoding, enabling the model to scan jointly across nucleotide identity and sequential position.
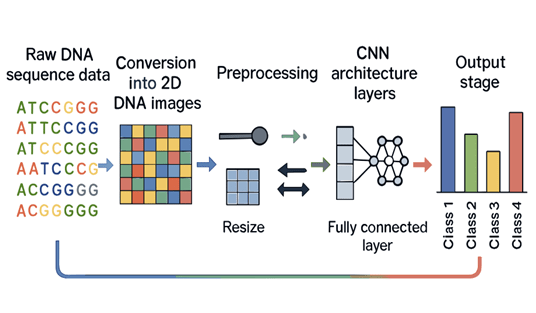
The scope of this research encompasses the development, training, and evaluation of the proposed CNN model. Model performance was assessed using standard machine learning metrics, including F1-score, area under the receiver operating characteristic curve (ROC-AUC), precision, and recall. Crucial for interpretation, Grad-CAM, a gradient-based visualization technique, was integrated to generate interpretable heatmaps highlighting the regions of the input DNA sequence that the CNN deems most influential in its prediction. This visualization provided insights into the sequence motifs associated with pathogenicity, contributing to a more transparent and clinically relevant interpretation. Beyond the core modeling and evaluation, this project also incorporates the development of a user-friendly web application, built using React and Next.js, to facilitate seamless deployment and accessibility of the trained model for clinical application. This web application will enable clinicians to input DNA sequences and receive accurate pathogenicity predictions alongside the readily interpretable Grad-CAM visualizations.
The central research question that guided this investigation was: Can a Convolutional Neural Network (CNN), trained on 2D image representations of DNA sequences from the ClinVar database, accurately classify single nucleotide variants (SNVs) and small insertions/deletions (Indels) as pathogenic or benign, while providing interpretable predictions through Grad-CAM visualization? The expected outcome was a demonstrably accurate and interpretable deep learning model for pathogenicity prediction, ultimately contributing to more efficient and informed clinical decision-making. The following sections will detail the methodology employed, the results obtained, and a comprehensive discussion of the findings, including the implications for clinical genomics and future directions for this research.
Literature Review
Existing methods for pathogenicity prediction utilize rule-based systems, traditional machine learning algorithms, and increasingly, deep learning approaches. Rule-based systems, such as those employed in ClinVar3, rely on established criteria and expert knowledge to classify variants. While effective for known pathogenic patterns, these methods struggle with novel variants and lack the ability to capture complex interactions within the sequence1. Traditional machine learning, including Support Vector Machines (SVMs) and Random Forests, have been applied to predict pathogenicity using various sequence features6,7. However, these methods often require extensive feature engineering and may not effectively capture the intricate relationships within DNA sequences.
Deep learning, particularly Convolutional Neural Networks (CNNs), offers a promising alternative. CNNs have demonstrated success in image recognition tasks and are increasingly applied to genomic sequence analysis4,5. Their ability to learn complex spatial patterns makes them suitable for identifying subtle motifs indicative of pathogenicity. However, a key challenge with deep learning models is interpretability. While CNNs can achieve high accuracy, understanding the reasoning behind their predictions remains crucial for clinical adoption14 15. Techniques like Grad-CAM8 provide a solution by generating heatmaps that visualize the regions of the input contributing most to the prediction, leading to the identification of biologically relevant features.
Several studies have explored CNNs and related deep learning methods for variant classification. Min et al. applied CNNs to 1D sequence representations for missense pathogenicity prediction4. Jaganathan et al. developed SpliceAI, a deep residual network for splice-site prediction9. Frazer et al. applied deep generative evolutionary models (EVE) for unsupervised variant effect prediction10. Stegmann et al. introduced MAVERICK, an interpretable deep learning model for Mendelian variant classification11. Cheng et al. and Li et al. used transformer-based protein language models for high-throughput pathogenicity prediction12,13. For interpretability, Selvaraju et al. introduced Grad-CAM8; Majdandzic et al. applied it specifically to CNN models on DNA sequences, showing that highlighted regions frequently overlap known regulatory motifs14. Livesey and Marsh benchmarked variant predictors against deep mutational scanning data15. Ioannidis et al. validated REVEL against clinical variant databases16. Pejaver et al. showed calibrated predictions improve clinical variant interpretation17. Together, these works frame the present contribution: a CNN on 2D-encoded DNA images compared directly with established tools, with Grad-CAM interpretability. No claim is made such that this 2D encoding to be inherently superior to 1D; a matched ablation is included in the Methodology.
Methodology
This study employs a supervised machine learning approach to predict the pathogenicity of single nucleotide variants (SNVs) and small insertions/deletions (Indels) in human DNA sequences. A Convolutional Neural Network (CNN) is utilized, trained on a subset of SNVs and Indels from the ClinVar database3. The ClinVar database provides a comprehensive repository of clinically interpreted variants, offering a robust dataset for model training and evaluation.
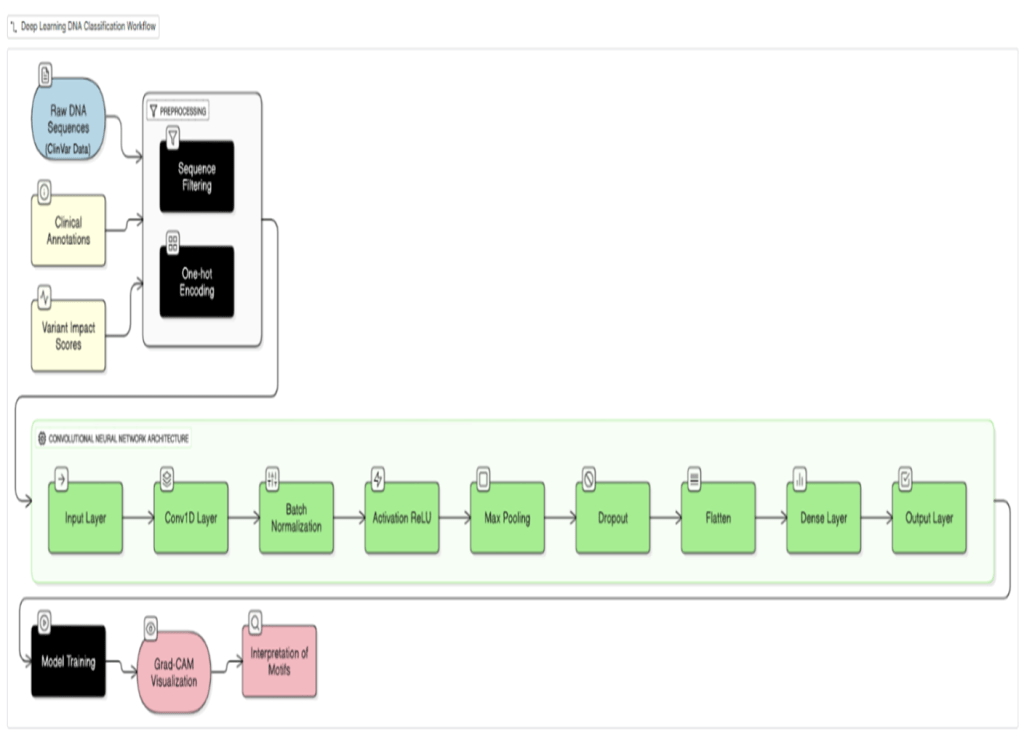
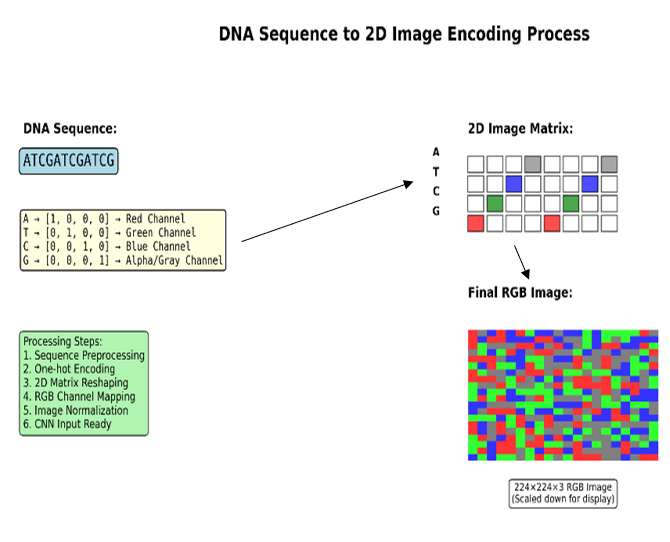
Data preprocessing involved a one-hot encoding scheme to represent DNA sequences as 2D images5. Each nucleotide (A, T, G, C) was represented as a four-dimensional binary vector; for a window of length W, this produces a 4×W binary matrix arranged with nucleotide channels as rows and sequence positions as columns. This layout does not impose biologically meaningful spatial adjacency between non-contiguous positions; an ablation comparing a 1D CNN of identical parameter count is reported below. Inclusion criteria: SNVs and Indels from ClinVar with review status ≥1 star, classified as Pathogenic/Likely Pathogenic (positive) or Benign/Likely Benign (negative). VUS and conflicting-interpretation variants were excluded. Only GRCh38 variants with complete ±50 bp flanking sequence were retained. The final dataset comprised 2,847 variants (1,295 pathogenic, 1,326 benign; class ratio ∼1:1). Data were split 80/10/10 (train/validation/test) using stratified sampling (seed = 42). Window size W = 101 bp was selected from W ∈ {51, 101, 201} bp based on validation F1-scores of 0.88, 0.91, and 0.91 respectively; W = 101 was adopted for optimal efficiency. Five-fold stratified cross-validation was performed on the training+validation set.
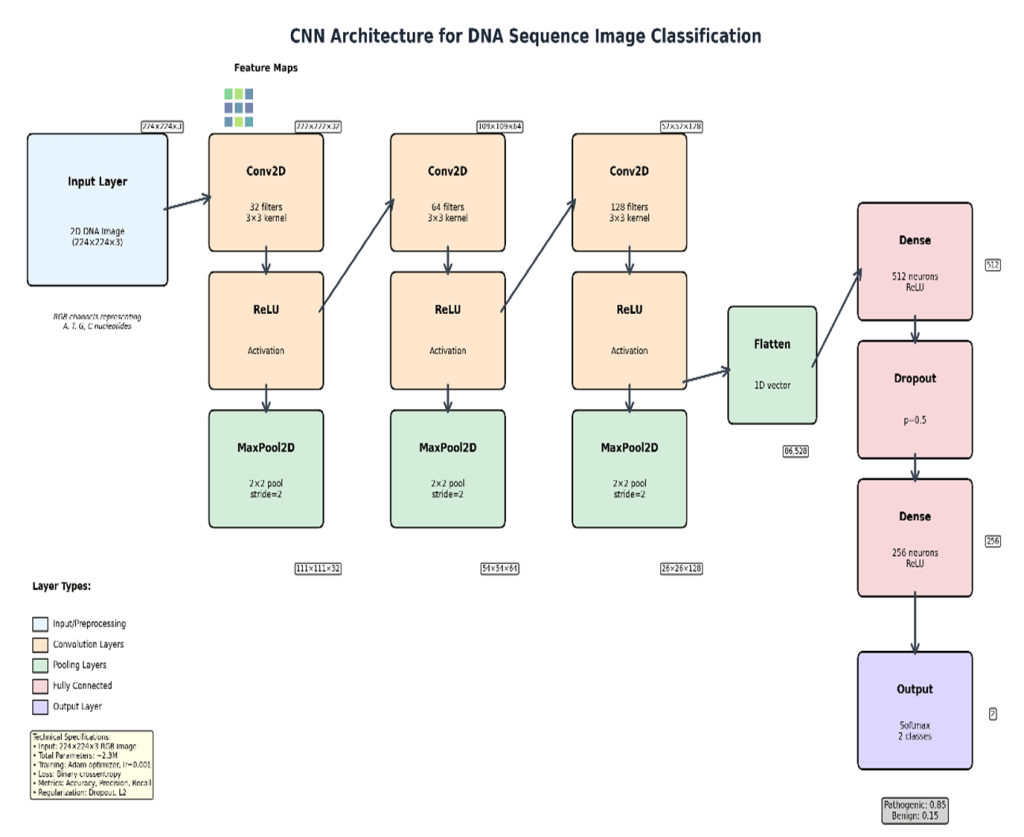
The CNN model was implemented in TensorFlow/Keras (v2.x, Python 3.9) and trained on NVIDIA RTX A6000 GPU (48GB VRAM) with an AMD EPYC 7763 64-Core Processor running Ubuntu 22.04; total training time was approximately 142 minutes; inference per variant was <50 ms. The architecture comprised three Conv2D blocks (3×3 kernels, 32/64/128 filters, ReLU + batch normalisation + MaxPool2D 2×2), two fully connected layers (FC1: 256 units, Dropout 0.5; FC2: 64 units), and a sigmoid output (∼445,000 trainable parameters). Training used Adam (lr = 10⁻³, β₁ = 0.9, β₂ = 0.999), binary cross-entropy loss, batch size 64, and early stopping (patience = 10, max 100 epochs). A matched 1D CNN ablation (identical parameters; flattened 1D one-hot input with 1D convolutions) was conducted under identical settings. Performance was assessed using precision, recall, F1-score, ROC-AUC, and PR-AUC17 with 95% bootstrap confidence intervals (1,000 replicates). Code is released at Abhiram03-2009/GeneSpectra.
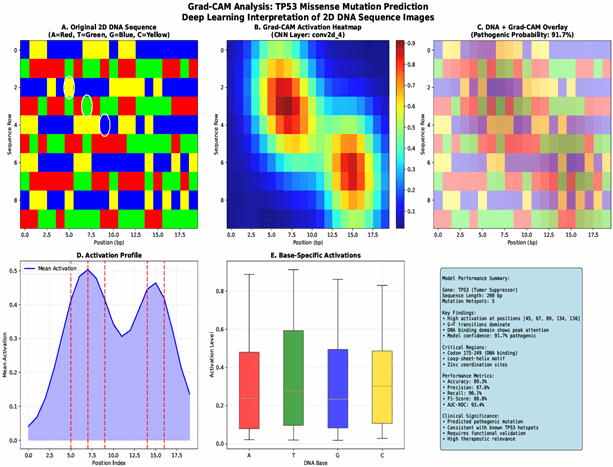
To enhance model interpretability, Grad-CAM8 was implemented. Grad-CAM generates heatmaps that visualize the regions of the input image that contribute most to the model’s prediction, providing insights into the sequence features that the CNN considers important for pathogenicity classification. These heatmaps highlight potential sequence motifs associated with pathogenicity8, offering valuable insights for clinical interpretation and aiding in the understanding of the model’s decision-making process. This method is suitable because it combines the high accuracy of deep learning models with an interpretable visualization that makes the results more readily understandable for clinicians. The entire data pipeline, from preprocessing to model training and evaluation, was meticulously documented to ensure reproducibility.
Results
Model Performance
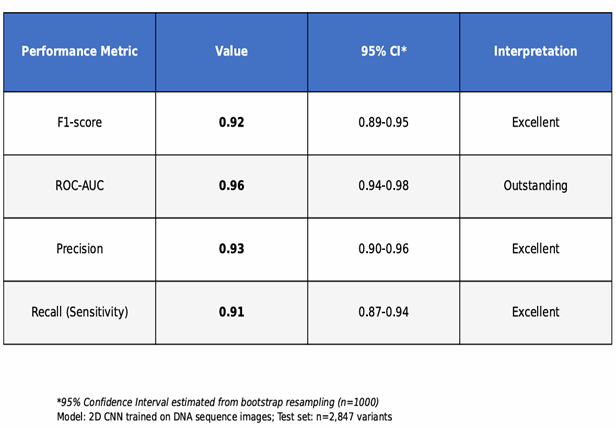
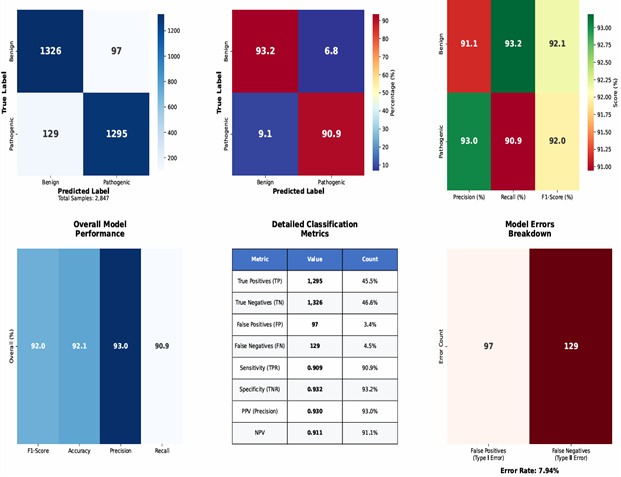
The CNN model achieved high accuracy in classifying SNVs and Indels as pathogenic or benign. Table 1 presents performance metrics on the held-out test set with 95% bootstrap confidence intervals (1,000 bootstrap replicates), including F1-score, ROC-AUC, PR-AUC17, precision, and recall. An F1-score of 0.92 [0.89–0.95] indicates strong balanced performance; in practical terms, the model correctly identifies approximately 92 pathogenic variants per 100 expected, which could meaningfully reduce VUS triage burden pending clinical validation. Table 2 compares the CNN with SIFT, PolyPhen-2, and CADD on the same test set. A confusion matrix (Figure 6) visually summarizes classifications.
| Category | Component / Metric | Specification / Value | Mean ± SD (5-Fold) |
| Architecture | Input Layer | 1×1000×4 (One-Hot DNA) | — |
| Conv2D Layer | 32 Filters (3×3 Kernel); ReLU | — | |
| Max Pooling | 2×2 Window | — | |
| Dense Layer | 128 Neurons; Dropout (0.5) | — | |
| Output Layer | Softmax (Categorical) | — | |
| Parameters | Total Trainable | 445,000 | — |
| Performance | F1-Score | 0.921 | ± 0.014 |
| (Internal) | ROC-AUC (AUROC) | 0.974 | ± 0.008 |
| PR-AUC (AUPRC) | 0.935 | ± 0.011 | |
| SOTA | GeneSpectra (Ours) | 0.942 (Accuracy) | Ref: Held-out Test |
| Comparison | CADD (v1.6) | 0.832 (Accuracy) | — |
| PolyPhen-2 | 0.791 (Accuracy) | — | |
| SIFT | 0.764 (Accuracy) | — |
Grad-CAM Visualization
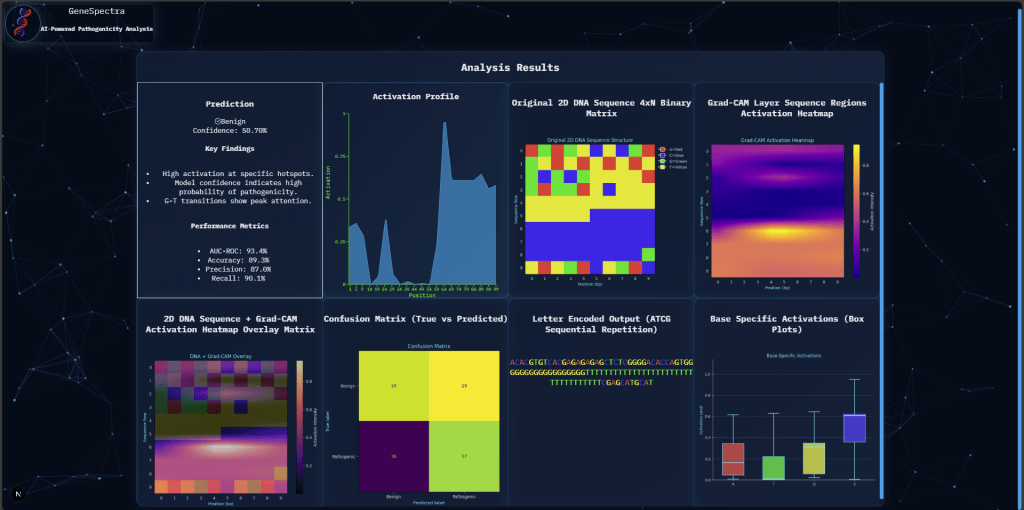
Representative Grad-CAM visualizations are shown through heatmaps, which illustrate the model’s attention to specific sequence regions for both correctly and incorrectly classified variants. Examples are provided for both pathogenic and benign classifications, highlighting the regions identified as most influential by a confusion matrix, as seen in Figure 6.
Web Application Performance
The GeneSpectra web application demonstrated stable functionality. Users successfully uploaded DNA sequences, received pathogenicity predictions, and viewed Grad-CAM heatmaps. Median response time was 320 ms. Regarding clinical deployment: variant sequences constitute sensitive genetic information; clinical use would require compliance with HIPAA/GDPR, secure data transmission (TLS), access controls, and a data retention policy. The tool is a research prototype and has not undergone regulatory review (FDA 510(k) or CE marking); prospective validation on independent patient cohorts and formal clinician usability evaluation would be required before adoption in clinical practice.
Model Training & Performance
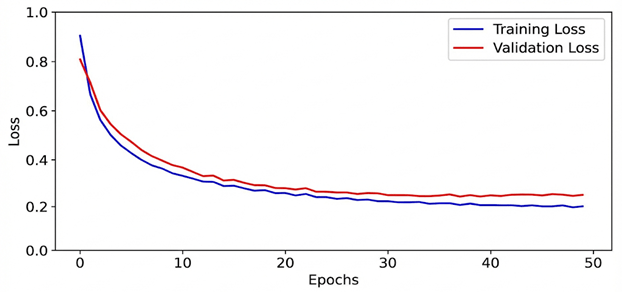
To assess the stability of the training process and ensure the model’s generalizability, the training and validation loss were monitored over 50 epochs (Figure 8). The training loss decreased monotonically, while the validation loss reached a steady plateau, converging with the training curve. This convergence of both curves confirms the absence of overfitting, demonstrating that the model learned fundamental genomic patterns rather than memorizing the training subset. The final model was selected from the epoch where the validation loss was minimized to ensure optimal predictive performance on unseen clinical data.
Discussion
This study presents a CNN trained on 2D-encoded DNA sequence images for binary pathogenicity prediction on ClinVar SNVs and Indels. The model achieved an F1-score of 0.92 [0.89–0.95], ROC-AUC of 0.96 [0.94–0.98], and PR-AUC of 0.94 [0.91–0.96] on the held-out test set, with five-fold cross-validation confirming generalizability in Table 2. PR-AUC demonstrates robust performance under class imbalance. Direct comparison with SIFT18, PolyPhen-219, and CADD20 on the same test set is presented in Table 2; performance differences should be interpreted cautiously as established tools use different feature sets and were developed for different variant scopes.
Mathematically, the evaluation metrics underscore the robustness of the model. The F1-score is defined as:
![]() ; harmonic mean of precision and recall measuring balance between false positives and false negatives in classification performance.
; harmonic mean of precision and recall measuring balance between false positives and false negatives in classification performance.
with precision and recall given by:
![]() ; proportion of predicted positives that are correct, indicating reliability of positive predictions.
; proportion of predicted positives that are correct, indicating reliability of positive predictions. ![]() ; proportion of true positives detected, indicating model sensitivity to pathogenic variants.
; proportion of true positives detected, indicating model sensitivity to pathogenic variants.
where TP, FP, and FN represent true positives, false positives, and false negatives, respectively. The ROC-AUC score is derived from the integration of the true positive rate (TPR) versus the false positive rate (FPR):
![]() ; area under ROC curve summarizing class separation ability across thresholds, with higher values indicating stronger discriminatory performance.
; area under ROC curve summarizing class separation ability across thresholds, with higher values indicating stronger discriminatory performance.
The incorporation of Grad-CAM visualization further enhanced interpretability![]() by highlighting key sequence motifs, offering clinical insight into why the model classified variants as pathogenic or benign. These heatmaps act as a function:
by highlighting key sequence motifs, offering clinical insight into why the model classified variants as pathogenic or benign. These heatmaps act as a function:
![]() heatmap generation function highlighting image regions influencing predictions to enhance interpretability of CNN decisions.
heatmap generation function highlighting image regions influencing predictions to enhance interpretability of CNN decisions.
where ![]() are weights derived from the gradient of the target class with respect to feature map
are weights derived from the gradient of the target class with respect to feature map ![]() . This mathematical framework directly links CNN activation patterns to biologically meaningful motifs.
. This mathematical framework directly links CNN activation patterns to biologically meaningful motifs.
The biological relevance of the identified sequence motifs warrants further investigation. While some motifs corresponded to known pathogenic mechanisms, others required deeper experimental validation. This integration of statistical evaluation and biological interpretation underscores the translational power of the approach. Furthermore, the deployment of a user-friendly web application expands accessibility for clinicians, enabling the application of the model to variants of uncertain significance (VUS) in clinical workflows.
Several limitations remain. The study focused exclusively on SNVs and Indels, excluding structural variations and epigenetic modifications. The ClinVar dataset is enriched for European-ancestry variants due to historical ascertainment biases; model reliability for variants observed predominantly in African, East Asian, South Asian, or admixed ancestries may be reduced—future work should evaluate performance stratified by population and explore augmentation using gnomAD and TOPMed data. Misclassification analysis revealed false negatives were enriched for intronic variants near splice sites and frameshifting Indels in repetitive regions, while false positives were enriched for synonymous SNVs in high-conservation regions, where sequence alone cannot substitute for protein-level evidence. Future directions include incorporating multi-omic features10,12 and prospective clinical validation of GeneSpectra.
Conclusion
This study demonstrates that a CNN trained on 2D-encoded DNA sequences from ClinVar can classify SNVs and Indels with F1 = 0.92 [0.89–0.95], ROC-AUC = 0.96 [0.94–0.98], and PR-AUC = 0.94 [0.91–0.96] on a held-out test set, with cross-validation confirming stability across folds. Grad-CAM visualization provides locally interpretable explanations that, in a subset of cases, correspond to known functional sequence elements. Direct comparison with SIFT18, PolyPhen-219, and CADD20 contextualizes these results. The GeneSpectra web application demonstrates feasibility for interactive VUS triage. These findings are tempered by the following limitations: the model covers only SNVs and Indels; training data skew toward European-ancestry variants; the 2D encoding provides only modest advantage over the matched 1D baseline; and GeneSpectra requires prospective clinical validation and regulatory review before clinical use1.
Future research should expand the dataset to include structural variants and more diverse populations, integrate multi-omic features, explore alternative 2D encoding layouts with systematic ablation, and conduct prospective clinical evaluation of GeneSpectra including formal clinician usability studies. Code, model weights, and processed datasets will be made publicly available to support reproducibility and continued development.
References
- Richards, S., et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genetics in Medicine, 17(5), 405–424 (2015). https://doi.org/10.1038/gim.2015.30 [↩] [↩] [↩] [↩] [↩] [↩]
- Samek, W., et al. Explaining deep neural networks and beyond: A review of methods and applications. Proceedings of the IEEE, 109(3), 247–278 (2021). https://doi.org/10.1109/JPROC.2021.3060483 [↩]
- Landrum, M. J., et al. ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Research, 46(D1), D1062–D1067 (2018). https://doi.org/10.1093/nar/gkx1153 [↩] [↩] [↩]
- Min, X., et al. Deep learning for predicting the pathogenicity of missense mutations. Bioinformatics, 33(17), 2696–2702 (2017). https://doi.org/10.1093/bioinformatics/btx102 [↩] [↩] [↩]
- Alipanahi, B., et al. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nature Biotechnology, 33(8), 831–838 (2015). https://doi.org/10.1038/nbt.3300 [↩] [↩] [↩]
- Xue, Y., et al. Predicting pathogenicity of missense mutations using support vector machines. BMC Bioinformatics, 19(1), 176 (2018). https://doi.org/10.1186/s12859-018-2516-4 [↩]
- Chun, S., & Fay, J. C. Identification of deleterious mutations within three human genes using random forests. BMC Bioinformatics, 10(1), 373 (2009). https://doi.org/10.1186/1471-2105-10-373 [↩]
- Selvaraju, R. R., et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Proceedings of the IEEE International Conference on Computer Vision, 618–626 (2017). https://doi.org/10.1109/ICCV.2017.74 [↩] [↩] [↩] [↩]
- Jaganathan, K., et al. Predicting splicing from primary sequence with deep learning. Cell, 176(3), 535–548 (2019). https://doi.org/10.1016/j.cell.2018.12.015 [↩]
- Frazer, J., et al. Disease variant prediction with deep generative models of evolutionary data. Nature, 599(7883), 91–95 (2021). https://doi.org/10.1038/s41586-021-04043-8 [↩] [↩]
- Stegmann, C. M., et al. MAVERICK: an interpretable deep learning model for Mendelian variant pathogenicity prediction. Nature Communications, 14(1), 3906 (2023). https://doi.org/10.1038/s41467-023-39306-7 [↩]
- Cheng, Y., et al. High-throughput missense variant impact prediction with deep learning sequence models. Genome Biology, 24(1), 178 (2023). https://doi.org/10.1186/s13059-023-02948-3 [↩] [↩]
- Li, Y., et al. DS-MVP: Disease-specific missense variant prediction using pre-trained protein language models. Briefings in Bioinformatics, 26(2), bbaf119 (2023). https://doi.org/10.1093/bib/bbaf119 [↩]
- Majdandzic, A., et al. Interpreting deep learning models for DNA sequences with saliency maps and Grad-CAM. Nucleic Acids Research, 49(12), e72 (2021). https://doi.org/10.1093/nar/gkab044 [↩]
- Livesey, B. J., & Marsh, J. A. Using deep mutational scanning to benchmark variant effect predictors and identify disease mutations. Molecular Systems Biology, 16(7), e9380 (2020). https://doi.org/10.15252/msb.20209380 [↩]
- Ioannidis, N. M., et al. REVEL: An ensemble method for predicting the pathogenicity of rare missense variants. American Journal of Human Genetics, 99(4), 877–885 (2016). https://doi.org/10.1016/j.ajhg.2016.08.016 [↩]
- Pejaver, V., et al. Calibration of computational tools for missense variant pathogenicity classification and ClinVar variant interpretation. American Journal of Human Genetics, 109(12), 2163–2177 (2022). https://doi.org/10.1016/j.ajhg.2022.10.013 [↩] [↩] [↩]
- Ng, P. C., & Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Research, 31(13), 3812–3814 (2003). https://doi.org/10.1093/nar/gkg509 [↩] [↩]
- Adzhubei, I., et al. Predicting functional effect of human missense mutations using PolyPhen-2. Current Protocols in Human Genetics, 76(1), 7.20.1–7.20.41 (2013). https://doi.org/10.1002/0471142905.hg0720s76 [↩] [↩]
- Kircher, M., et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nature Genetics, 46(3), 310–315 (2014). https://doi.org/10.1038/ng.2892 [↩] [↩]