Abstract
Messenger ribonucleic acid (mRNA) vaccines have shown significant development in the past few years, specifically with the COVID-19 mRNA vaccine, which has paved for other mRNA vaccines. This review examines and compares the development of the COVID-19 mRNA vaccine and the cancer mRNA vaccine. The analysis includes the foundation, key factors, and limitations of the COVID-19 and cancer mRNA vaccines. Findings demonstrate the adaptability of mRNA technology and its uses for different purposes. However, significant barriers remain such as mRNA stability, delivery system limitations, and the individualization of vaccines.
Keywords: mRNA vaccine, COVID-19, Cancer.
Introduction
The COVID-19 pandemic marked a turning point for mRNA vaccines. Although the concept of basing a vaccine on mRNA was first conceived in the 1960s, developing such a vaccine has faced many challenges1. For example, although researchers sought to develop mRNA vaccines for diseases such as Zika, influenza, and human immunodeficiency virus (HIV), those attempts were ultimately unsuccessful. Concurrently, researchers endeavored to utilize mRNA vaccines for non-infectious diseases, but the use of this type of vaccine was not widespread.
It was not until the COVID-19 pandemic that this type of vaccine had its breakthrough, illustrating the potential of mRNA vaccines for widespread application. Even though the foundations of mRNA vaccines have been known for three decades, demonstrating the maturity of this technology, their use was not widespread until the COVID-19 mRNA vaccine was developed rapidly and approved in less than a year. On the other hand, mRNA vaccines for diseases like cancer are still in clinical trials. Therefore, the rapid development of the COVID-19 vaccine raises a question: how does the development of an mRNA vaccine for an infectious disease such as COVID-19 compare with that for an individualized, non-infectious disease such as cancer?
The aim of this study is to analyze and compare the development of mRNA vaccines, specifically for COVID-19 and cancer, in terms of biological, technological, and strategic differences, specifically the differences in mechanisms and development pipelines of the mRNA vaccines. Therefore, the vaccines will be evaluated across four factors: antigen selection, vaccine design, delivery system, and their limitations.
Methodology
A comprehensive literature review was performed using published articles and peer-reviewed research articles from Google Scholar to identify relevant research to provide a comprehensive understanding of the similarities and differences in mRNA vaccine development. Studies were included if they focused on the development pipelines and the mechanisms of COVID-19 mRNA vaccine and cancer mRNA vaccine. Moreover, to ensure sources are relevant to addressing the development of the COVID-19 mRNA vaccine and cancer vaccine, the following key words were used in the search: mRNA vaccine development, COVID-19 mRNA vaccine mechanisms, cancer mRNA vaccine mechanisms, mRNA design, and mRNA delivery systems.
mRNA Vaccine Design
Foundation of mRNA Vaccines
Unlike traditional vaccines, which uses a weakened or inactivated antigen to produce an immune response, mRNA vaccines use mRNA to produce a specific protein found in the pathogen, which stimulates the immune system to recognize and respond to the disease. Specifically, mRNA vaccines deliver the genetic code for producing a protein unique to the target pathogen. Once in contact with host cells, the mRNA strand is detected as foreign, corresponding to the protein. In essence, the mRNA strand provides a molecular blueprint for antigen production. The detection of the foreign protein signals to the body to produce antibodies. Thereafter, the strand of mRNA is destroyed following the development of antibodies to develop one’s immunity against the pathogen, ensuring a short, safe, and effective response2,3.
As stated before, the COVID-19 vaccine had unprecedented success in its implementation during the pandemic. These vaccines prevent and weaken the reaction when coming in contact with the pathogen by replicating the spike protein found on the surface of the SARS-CoV-2 virus. The development and function of the COVID-19 vaccine are illustrated in Figure 1. As depicted, the process begins with identifying the spike protein, which is located on the surface of the SARS-CoV-2 virus. Through genome sequencing, a complete strand of DNA can be replicated and used as a basis for the vaccine. A synthetic mRNA strand is then formulated and incorporated into the vaccine using lipid nanoparticles (LNPs)—small fat-based carriers that help protect the mRNA during delivery. Once injected into the body, the LNPs deliver the mRNA to host cells where it escapes the endosome, a critical step that prevents degradation and allows translation into spike protein. The presence of the foreign mRNA activates the pattern-recognition receptors (PRRs) such as Toll-like receptors (TLRs), which trigger the innate immune response and cytokine release. These signals enhance dendritic cell (DC) activation, leading to effective antigen processing and presentation of major histocompatibility complexes (MHCs). DCs then prime CD8+ cytotoxic T cells to directly kill infected cells and CD4+ T helper cells to support B-cell antibody production. Therefore, this builds immunity by training the immune system to recognize and neutralize SARS-CoV-24. Currently, COVID-19 mRNA vaccines are widely approved and have been administered globally.
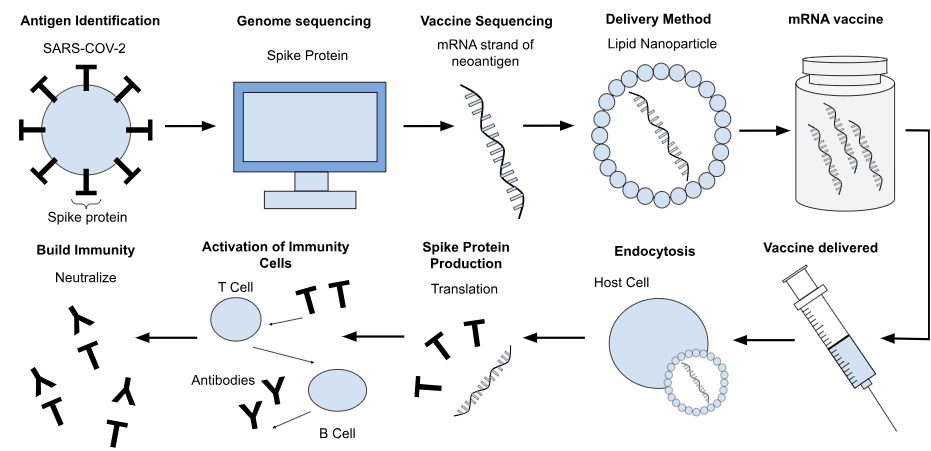
Moreover, the development of the COVID-19 vaccine has rapidly accelerated the development of mRNA vaccines as a whole, not only for infectious diseases, but also for non-infectious diseases. In contrast to the traditional vaccine, which is prophylactic and administered to healthy individuals to prevent a specific illness, cancer mRNA vaccines are therapeutic, meaning that they treat rather than prevent. They are customized for an individual patient in order to target cancerous cells in a patient’s immune system. Similar to the COVID-19 mRNA vaccine design, the cancer vaccine starts with a sample of a pathogen, in this case, an individual’s tumor and peripheral blood, as illustrated in Figure 2. This is used to analyze and compare both the cancerous cells, found in the tumor, and the healthy cells, found in the peripheral blood. Thereafter, neoantigens—a protein found in cancerous cells caused by mutations in the tumor’s DNA—are identified to be encoded into mRNA strands, which are delivered through LNPs, creating the personalized vaccine7. After injection, the process is largely similar to the COVID-19 mRNA vaccine, where the mRNA is recognized by PRRs on antigen-presenting cells, activating the innate immune system. Tumor antigens are translated by DCs and processed and presented on MHCs. This primes the CD8+ T cytotoxic cells, which are most important for cancer vaccines, as these cells are crucial for killing tumor cells, and CD4+ T helper cells, thereby activating an immune response from the body8,9. Furthermore, certain cancers have higher tumor mutational burden (TMB), the total number of somatic mutations present within a tumor genome, which correlates to a higher probability of producing neoantigens that can trigger a strong T-cell immune response. This affects how suitable different types of cancer can be for developing an individualized cancer vaccine treatment. The individualized nature of the cancer vaccine precludes its mass production, since they are tailored not only for the type of cancer, but also for the individual receiving treatment. However, there are cases where a shared vaccine can be developed when individuals share common mutations such as the KRAS or TP53, which make mass production possible10.
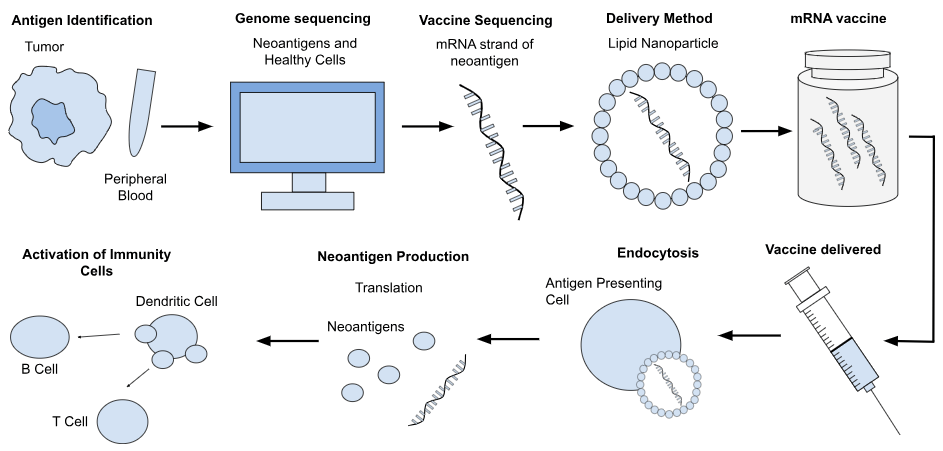
To sum up, while both mRNA vaccines rely on the same principles when developing an mRNA vaccine, their applications and scope differ due to being inherently different. The COVID-19 mRNA vaccine prevents individuals from contracting the SARS-CoV-2 virus, while the cancer mRNA vaccine targets existing cancerous cells.
Key factors
Vaccine Structure
The design of a vaccine is dependent on its purpose, which shapes certain development factors such as immune focus and target cells. Most commonly produced vaccines are for prophylactic use; however, this is not always the case, as some illnesses can be treated therapeutically. Regardless of whether the vaccine is prophylactic or therapeutic, the structural design itself remains consistent. As illustrated in Figure 3, the standard mRNA vaccine consists of five elements: the 5’ cap, 5’ and 3’ untranslated regions (UTR), open reading frame (ORF), and 3’ poly(A) tails. The 5’ cap, located at one of the ends of the mRNA vaccine, assists in translation efficiency and mRNA stability, playing a critical role in protein production and immunogenicity12. The 5’ and 3’ UTRs are regions vital for maximizing gene expression and subcellular localization13,14. Elements such as the length of the structure and sequence all affect the functions of the mRNA. The ORF, also known as the coding region for the selected antigen, is the one aspect of the vaccine that is modified and tailored to the specific illness and contributes to the expression of the antigen 15. Lastly, the 3’ poly(A) tails are crucial for the translation process and stability of the mRNA molecule16.
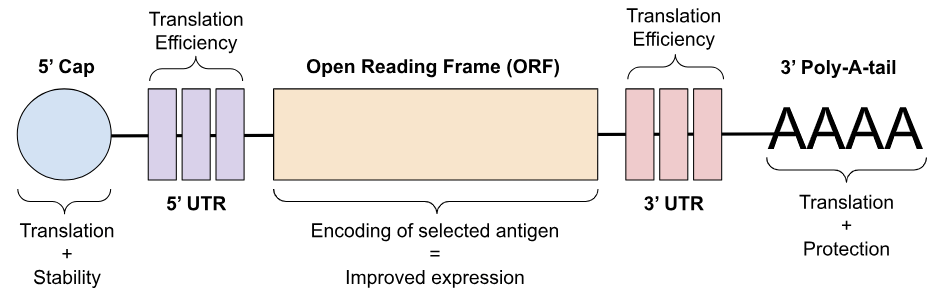
Vaccines for infectious diseases such as COVID-19 mimic a component of the selected pathogen, which primes the immune system to recognize and react to the foreign pathogen. By expressing an antigen for the spike protein, the vaccine induces an antibody response through B cell activation, along with CD4⁺ T helper cells and CD8⁺ T cytotoxic cells, which help build long-term immunity. The mRNA vaccine design contains the five standard elements, but the ORF is modified to encode the spike protein18. While this varies between vaccines produced by different companies, Pfizer and BioNTech both use a similar coding sequence, as seen in Figure 4. The ORF contains the full length spike protein and two proline substitutions, K986P and V987P, which improve expression and prefusion stability19. The ORF has two parts, S1 and S2, which are protein subunits. S1 contains the N-terminal domain (NTD) and the receptor binding domain (RBD) and functions as a receptor binder. S2 facilitates cell entry and membrane fusion, and contains the fusion peptide (FP), internal fusion peptide (IFP), heptad-repeat domains 1 and 2 (HR), transmembrane domain (TM), and c-terminal domain (CT)20. There is also a nucleoside modification with the substitution of a N1-methylpseudouridine to increase stability and low immunogenicity21. A full explanation of these regions is addressed in other studies, but is outside the scope of this review.
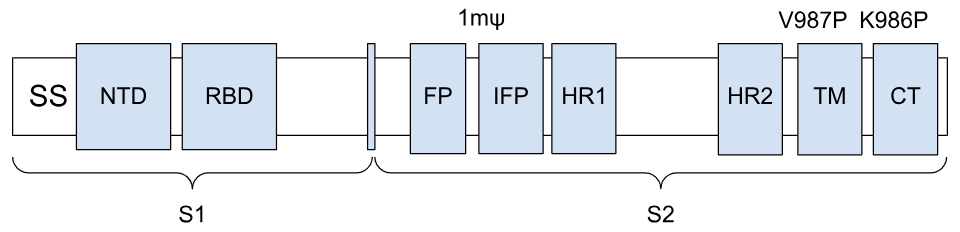
On the other hand, cancer vaccines are aimed to be therapeutic. These vaccines also induce a response through CD4⁺ T helper cells and CD8⁺ T cytotoxic cells. An example is illustrated in Figure 5, where a general illustration of the sequence in the coding region of the cancer vaccine is shown, as various modifications are made for specific cancers. Neoantigens are encoded into the ORF and usually around 20 neoantigens are used. However, this number may vary depending on the vaccine and individual, meaning some vaccines may only have seven neoantigens, while others may have up to forty neoantigens encoded in the ORF. A larger number of neoantigens allows for more to be targeted at once22,23.
Overall, it is clear that the structure of both mRNA vaccines, specifically their ORF, follows a similar foundational structure. In the case of the COVID-19 vaccine, the ORF encodes a spike protein to prime the immune system for future infections, whereas the cancer vaccine uses neoantigens to trigger an immune response. Despite targeting different diseases, they induce responses through the T cells, the common cells that build a strong immune system.
Antigen Selection
Antigen selection is critical in developing a vaccine as it is the basis of the vaccine itself. Antigens are markers that trigger a response in the human body when a foreign species, such as a pathogen, is detected by the immune system. In mRNA vaccines, antigens are the key basis of the vaccine as they are the component that is encoded into the strand of mRNA. The host’s cells will then use this mRNA to produce the antigen in native conformation, preparing the body to recognize and eliminate the pathogen.
In the case of the COVID-19 vaccine, the selected antigen is the spike protein, found on the virus’ surface. This protein was specifically chosen as it plays a crucial role in enabling the virus to bind to host cells and initiate membrane fusion, processes which are essential to viral entry and infection. The spike protein is the primary antigen that triggers immune responses, stimulates neutralizing antibodies, and provides protective immunity against infection. Due to these functions, targeting the spike protein allows the vaccine to simulate a similar reaction to a natural infection of the virus, stimulating the body to produce antibodies to counteract the infection18,25.
On the other hand, most cancers are highly individualized, with tumors carrying unique genetic mutations that differ from patient to patient, even in patients diagnosed with the same type of cancer. However, there are certain cancers which produce a common genetic mutation, such as KRAS and TP53 10. Thus, there are two types of cancer mRNA vaccines: individualized and shared vaccines. However, the main concept follows for both: instead of identifying a common antigen in the pathogen—as is done with infectious disease vaccines—tumor-associated antigens (TAAs) are selected. These are typically neoantigens, which are mutated proteins found only in cancer cells and not in normal human tissue, making them targets for a precise immune response, as they are displayed on the surface of cancer cells via MHCs, which are easily recognized by T cells. This therefore demonstrates a promising approach to cancer mRNA vaccines18.
Although both vaccines rely on an antigen, the approach is fundamentally different. For the COVID-19 vaccine, it is demonstrated that antigen selection is significantly easier than for a cancer vaccine, as antigen selection is not always possible for the case of certain cancers that are not accessible for biopsy. This allows infectious-disease vaccines to be compatible for mass production. Even though it is possible to mass-produce vaccines for cancers that exhibit shared genetic mutations across patients, the majority of individuals diagnosed with cancer do not have the mutation. This is especially limiting for individuals with cancerous cells, which are more difficult to identify due to having fewer genetic mutations in an individual’s cells. Therefore, instead of only considering the coding region, researchers are investigating cryptic antigens, as they are found in a significant number of cancer-associated immunopeptidomes, which can be an alternative to a neoantigen 7. Although this development helps individuals with less identifiable cancerous cells, the process still involves a biopsy, which may not be possible for all patients.
Delivery System
A delivery system is intended to support the mRNA strands to ensure successful deployment. This system is essential, as naked mRNA is inherently unstable and is easily degraded by nucleases in the human body due to the molecule being negatively charged10. This leads to inefficient cellular uptake, making the protection, delivery, and release of mRNA one of the biggest challenges in developing an mRNA vaccine. Additionally, this factor applies to almost all mRNA vaccines. It is not individualized for any mRNA vaccine, as its purpose is to protect the mRNA strands, which does not affect the target application of the mRNA. However, they may have modifications that are better suited to the type of illness for enhanced expression. Therefore, these delivery systems are often researched to improve drug delivery for both infectious and non-infectious diseases.
Following the discovery of mRNA in the 1960s, it was not until 1978 that the liposome-based delivery system was developed for packaging and transporting mRNA, which was then tested in vaccines in 1993. However, liposomes showed potential toxicity during clinical trials, so scientists created ionizable lipid nanoparticles (LNPs), which were developed around the time the COVID-19 pandemic broke out. Therefore, with the readily available information on lipid-based delivery systems, it was clear to use them in the development of the COVID-19 vaccine, as they had shown success in early studies26. Additionally, when comparing this new method to the naked mRNA, studies had shown that naked mRNA strands were approximately 100 times less effective at entering cells compared to mRNA strands protected by liposomes, specifically lipofectin, a liposome containing a cationic lipid27. Due to this success, both Pfizer-BioNTech and Moderna took similar approaches and used LNPs to encapsulate the mRNA strand. These particles are made of cationic lipids, auxiliary lipids, cholesterol, and polyethylene glycol (PEG), creating a lipid bilayer that forms around an aqueous core that carries the mRNA strands. In clinical trials, LNPs have been shown to trigger stronger immune responses compared to naked mRNA, as they protect the mRNA long enough to allow it to produce antigen-specific CD4+ T helper cell responses and generate an effective neutralizing antibody response. Although this delivery system has shown success in protecting the mRNA strands, pre-clinical trials demonstrate that LNPs can have negative after-effects, including inflammation28. Similarly, most cancer vaccines have followed what is known as the standard in mRNA vaccine development and commonly use LNPs as the delivery method due to their success in the COVID-19 vaccine. However, there are various types of lipid-based delivery systems that are also being tested, one of them being the cationic lipid-assisted nanoparticles (CLAN)29. These are made from a cationic lipid combined with a copolymer that is biocompatible and biodegradable, creating a poly(ethylene glycol)-block-poly(lactic-co-glycolic acid) (PEG-b-PLGA). In preclinical trials, studies showed high efficiency with dendritic cell (DC) transfection and were able to stimulate DC cell maturation, which induced a strong CD8+ T cytotoxic cell response. However, there are still gaps in the research on how the components affect the distribution of the nanoparticles in the host cell29. Moreover, lipoplex-type mRNA has also been developed. There are various modifications that can be made to this delivery system, but it is mainly composed of two elements: a lipid and a peptide. In this study, three lipid components were considered: an SS-Cleavable pH-activated lipid-like material combined with vitamin E scaffolds (ssPalmE), myristic acid scaffolds (ssPalmM), and egg phosphatidylcholine (EPC), which are used to increase transfection efficiency and decrease toxicity. The peptide component used was either a cationic peptide “KALA” or an R8 peptide, which helps cellular uptake of mRNA. From these elements, the combination ssPalmE-KALA had significantly higher transfection activity, demonstrating its efficiency in transporting mRNA strands into DCs. Although studies demonstrate it as a possible delivery method, the positively charged lipid, which is crucial for the protection of the mRNA, may interact and stick to other negatively charged molecules, which means that fewer mRNA strands are reaching their intended target cell30,31. Another alternative is the lipid/calcium/phosphate nanoparticles (LCP-NPs), which consist of a calcium phosphate core and an asymmetrical lipid bilayer. It uses mannose, which binds to the mannose receptors found on DC receptors, which promotes the delivery of the antigen and triggers the immune response of CD8+ T cytotoxic cells. In a preclinical study using a murine 4T1 breast cancer model, a mouse model, the vaccine administered demonstrated a strong antigen-specific response. Additionally, when combined with an immunotherapy, anti-CTLA-4 monoclonal antibodies, the anti-tumor immune response is significantly enhanced, compared to either technique used independently. When completing safety examinations, results showed no damage to major organs, and toxicity levels measured in the liver and kidney were in the normal range. However, in the combined method, there was a slight rise in hemoglobin levels. These findings highlight the success with the LCP-NPs, and their use in combined cancer therapy, in protecting mRNA strands in clinical trials; however, more research is needed to ensure that LCP-NPs are safe to use32,18.
Alternatively, there have been developments for chemically based delivery systems such as self-assembled polymeric micelles. This was developed to address inflammation, which was recorded during LNP preclinical trials due to the ionized component of the lipid chain33. The self-assembled polymeric micelles are made up of polyethyleneimine (PEI), a water-soluble cationic polymer, and vitamin E succinate (VE), a hydrophobic molecule. These two components are chemically bonded, creating a copolymer known as PVES. Upon contact with water, the copolymer forms micelles, where the VE clusters form a hydrophobic core and the PEI forms a shell. Between each polymer, there are pockets where mRNA strands are placed for protection when transporting them into host cells. These self-assembled polymeric micelles are only 10 to 100 nanometers wide, which gives them an advantage compared to other lipid nanoparticles, as it can easily access tumors. Additionally, they can be easily modified to add active targeting ligands to increase cell uptake, making the process more efficient. They also showed no inflammation during the pre-clinical trials, which was a concern from using LNPs, demonstrating this to be an improvement from the standard delivery system currently used. However, some challenges still need to be addressed, one of them being loading drugs in the micellar core, as its stability is based upon the bond of the polymer and hydrophobic molecule, so suitable amphiphilic polymers are still being tested to ensure stability. Additionally, due to it being self-assembled, it is less stable as it is reactive to changing pH levels. This can affect the kinetic stability of the micelle in vivo and also during drug release, and research in this area is still insufficient. Further studies are also needed to evaluate the toxicity and degradation of the components used34,35.
Finally, there are also DNA-based delivery strategies, such as nanoswitches, a pH-responsive i-motif cross-linked scaffold. There are variations of this method, but the main difference is their structure: triplex or quadruplex. The main concept is that the nanoswitches use gold nanoparticles and two types of DNA strands: one is an anti-MUC1 aptamer, which acts as a targeting device, binding to a tumor-associated membrane protein MUC1, and the other is a switchable DNA strand designed to change conformation upon contact with different levels of pH. At a neutral pH, the nanoswitch takes a linear conformation. In acidic pH, when the nanoswitch comes into contact with the cell during endocytosis, it changes into either the triplex or quadruplex conformation to release the mRNA strands. Additionally, not only does it respond to pH environments to change conformation, but it is also reversible. This means that the nanoswitch is not one-use, and it can control when mRNA strands are released, as it is pH reactive. This function would potentially allow for partial release of the mRNA strand, which would control the dose of the vaccine. However, the folding of the i-motif is very complex, as it may fold into multiple i-motifs, so the mechanisms of its structure are still being researched36,37.
To reiterate, delivery systems have been crucial for developing mRNA vaccines to ensure stability and efficiency when transferring mRNA strands to host cells. For the most part, delivery systems can be used for various illnesses. However, there are fewer studies focusing on the COVID-19 mRNA vaccine delivery strategy as it has already been developed and shown to be successful. Although the LNPs, the standard for mRNA delivery systems, demonstrate effective transfection and immune response, there are always limitations to the method, which is why there are so many drug delivery strategies in development to treat other illnesses, such as cancer.
| Key Factors | COVID-19 Vaccine | Cancer Vaccine |
| Vaccine Design | ORF encoded with the spike protein with various mutations to ensure stability and improve expression | ORF encoded with tumor-specific neoantigens, often unique to one patient |
| Antigen Selection | Spike protein is the single, universal antigen that is common amongst COVID-19 cases | Neoantigens are identified through biopsy and sequencing and vary across cancers and individuals |
| Delivery System | LNPs are the main delivery system in use right now, such as in the existing COVID-19 mRNA vaccine. Other delivery systems are being explored for mRNA use in general, such as lipoplexes, CLAN, and nanoswitches. | |
Limitations and Discussion
mRNA vaccines are still relatively new in the medical field, meaning that their development and production stages have limitations. Although the field has had various successes with its development, in particular the COVID-19 vaccines, there are still challenges and concerns which need to be addressed, not only for the COVID-19 mRNA vaccines, but also for the therapeutic ones which treat cancers.
Regarding the COVID-19 vaccine, due to SARS-CoV-2 virus’ ability to rapidly mutate, it remains a challenge to ensure sustained protection with the emergence of various variants such as B.1.1.7 (alpha), B.1.351 (beta), B.1.617.2 (delta), and P.1 (gamma). Vaccines need to be updated frequently to better target immune responses with the specific variant38. Additionally, vaccines lose neutralization potency over time, demonstrating the short protection period, as studies have shown a reduction in neutralization titers, the concentration of neutralizing antibodies39. Although there are booster shots that individuals are provided with, these vaccines still demonstrate a significant decrease in neutralization against the new variants of the SARS-CoV-2 virus. For instance, the B.1.1.529 (omicron) had 32 different mutations in the spike protein, which demonstrated a tenfold decrease in neutralization titers40. Even with multiple booster shots, neutralization ability was significantly lower with the variants. Therefore, new strategies must be considered to overcome the spread of variants.
Moreover, some studies have shown relatively high cases of adverse side effects when being administered the Pfizer/BioNTech and Moderna vaccines. It is usual for vaccines to cause fever, chills, and muscle aches, as this is essential to activating the immune response41. However, there were reports associating the mRNA vaccine with anaphylaxis, an allergic reaction that can be life-threatening. These reports concluded that it may result from the component polyethylene glycol (PEG), which helps stabilize and prolong its half-life. This allergy is estimated to affect up to 72% of individuals, demonstrating its possible role in triggering certain reactions, but this has not been proved to be the direct cause42,43.
Another limitation of the COVID-19 mRNA vaccines is their shelf life stability. For one, they are highly sensitive to temperature, meaning that they need to be stored in cooler temperatures to ensure stability during transportation and to ensure their efficacy44. This is referred to as the cold chain, and specifically for mRNA vaccines, they must be kept in temperatures of at least -20oC. For Moderna’s mRNA-1273, however, it must be stored between -60oC and -80oC. It is suggested that this is due to mRNA’s instability, and these temperatures are vital for maintaining the stability of the LNP-mRNA complexes. Therefore, this remains a challenge for mRNA vaccines, as there have not been many studies on developing methods to overcome this limitation. Another point to consider is that the COVID-19 mRNA vaccine shelf life is only six months if kept at the standard temperatures mentioned above. This shelf life reduces to only 30 days after being thawed at temperatures between 2oC and 8oC, which would affect its distribution rate, specifically in resource-poor countries45. Recent studies have considered freeze-drying the mRNA vaccines for conservation to reduce storage limitations, possibly extending the shelf life of the vaccine to 21 months when stored at 4oC46. This limitation can also apply to the cancer mRNA vaccine; however, due to its ongoing development and individual-specific nature, it is not currently as pressing of an issue compared with mRNA vaccines for infectious diseases.
Moreover, cancer vaccines have a similar limitation with their antigen selection. The new discovery of neoantigens to be encoded into mRNA has been a breakthrough, but these neoantigens can also further mutate and no longer be compatible. A way to address this is to target cancerous cells that are crucial for the survival of the cancer, but this topic is still a major concern for the development of cancer mRNA vaccines47. Furthermore, there is also tumor heterogeneity. There are two types: temporal heterogeneity, which refers to the genomes of the tumor changing over time, and spatial heterogeneity, which refers to tumors that have multiple genetically distinct cells. As mRNA strands for cancer vaccines are determined through biopsy, which cannot comprehensively determine the gene profile since only a small portion of the tumor is sampled, this limitation increases the risk of overlooking all mutations that could affect the efficacy of the vaccine. To combat the challenge, researchers have looked into multipoint sampling, which is when multiple samples are taken from an individual’s tumor to have a holistic view of the tumor. However, this is still a limitation for individuals who have tumors in places that cannot be operated on. Therefore, heterogeneity and tumor mutations can limit the success of developing an mRNA vaccine48,49,50.
In addition to the antigen selection limitation, another concern is the sustained growth of a tumor, which is strongly influenced by the tumor microenvironment (TME). The region around the tumor is immunosuppressive, meaning that it suppresses the body’s immune system to respond to foreign species through inhibiting the activation and function of T cells, which are critical for eliminating cancerous cells. The TME not only facilitates the evasion of tumors from the immune system, but it also reduces the effectiveness of immunotherapy51,52. For example, T regulatory cells (Tregs) or myeloid-derived suppressor cells (MDSC) expressing Programed Death-Ligand 1 (PD-L1) accumulate in the TME, and suppress vaccine primed T-cell activation. These mechanisms induce T-cell exhaustion which diminishes cancer vaccine efficacy53. To address this, scientists have investigated combined therapy, which refers to when the vaccines are used alongside other cancer treatments. For example, one current line of inquiry is using adoptive T-cell therapy to enhance the immune response induced by the mRNA vaccine by modifying the cell itself to better respond 54.
Due to cancer mRNA’s individualized nature, it is also costly to develop. For instance, an individualized cancer mRNA vaccine to treat brain cancer is expected to cost approximately 350,000 USD per patient. The details of this value are not fully disclosed, but such complex manufacturing processes and individualization demonstrate a significant economic barrier for most individuals55.
Despite these barriers, there have been promising clinical trials for cancer mRNA vaccines. An example is the mRNA-4157. Developed by Moderna in collaboration with Merck, mRNA-4157 is a personalized neoantigen vaccine which is currently in Phases II/III of testing. In a phase 2b trial with patients who had high risk melanoma, mRNA-4157/V940 was combined with pembrolizumab, a drug that helps the immune system recognize and attack cancerous cells. The combination treatment helped patients stay cancer free for longer, with 79% of patients staying disease free, compared to 62% in the pembrolizumab only group56. This vaccine suggests that personalized mRNA cancer vaccines like the mRNA-4157 could significantly improve outcomes for high-risk cancer patients.
In parallel, the COVID-19 mRNA vaccine has shown real-world effectiveness in the booster shots, for both Pfizer and Moderna. Studies in the UK found that people who received a booster were 70% lower risk of hospitalization and 89% lower risk of death from COVID-19 compared to those who did not get the booster. However, the effects waned over time, being the strongest during the first three months and by the sixth-month mark, only had 25% protection against testing positive57. These findings highlight the importance of boosters in maintaining protection from COVID-19.
Taken together, the COVID-19 mRNA vaccine and cancer vaccines are structurally very similar, with minor differences that tailor the vaccine to the specific disease. These differences are prevalent in the ORF of the vaccine design and the antigen selection, as these components are designed and chosen to target their respective illnesses. However, regarding the delivery systems, these are not as individualized to a specific mRNA vaccine. Their primary role is to protect the mRNA strands, but these systems are still developing to find a solution that optimizes antigen expression and mRNA stability.
Additionally, it is clear that due to the individualized nature of the cancer mRNA vaccine and the larger sense of urgency during the COVID-19 pandemic to create the vaccine, the trajectory of the development was considerably different. These factors demonstrate why the developmental period for the COVID-19 vaccine was significantly shorter than that of cancer vaccines, which are still ongoing.
The significance of using mRNA vaccines for prophylactic use and therapeutic use demonstrates the versatility of this technology. While the antigen selection highlights the distinct challenges between the two vaccines, it is clear that the technology is still highly applicable in both situations, with the COVID-19 vaccine serving as a proof of concept for mass production of mRNA vaccines for infectious diseases.
| Factor | COVID-19 Vaccine | Cancer Vaccine |
| Purpose | Prophylactic: aims to induce immunity before viral exposure | Therapeutic: aims to stimulate and target existing cancerous cells to eliminate them |
| Patient Specificity | Universal vaccine where the same design can be used for all individuals. | Primarily individualized to an individual, however, there are certain mutations (KRAS or TP53) that are common amongst cancerous cells which make a universal vaccine possible. |
| Antigen Mutations | SARS-CoV-2 mutates rapidly, creating variants, which requires vaccine updates and boosters. | Neoantigens can mutate becoming incompatible with the vaccine. Thorough initial biopsy is crucial for antigen identification. |
| Safety | Side effects include fever, chills, and muscle aches | Safety concerns would relate to biopsy complexity. No safety concerns of distribution currently exist. |
| Storage requirements | Cold chain is required roughly between -20oC to -80oC to ensure stability. | Not applicable as no cancer vaccine is available for distribution. Similar barriers may apply as the COVID-19 vaccine. |
Conclusion
This study highlights that, while the COVID-19 and cancer mRNA vaccines share a common framework, their development has been different. The COVID-19 vaccine was enabled by a clear, universal target, while the cancer vaccine encounters more challenges, as it is a therapeutic vaccine that focuses on the population affected by cancer. However, this also demonstrates the potential of mRNA vaccines for broader applications, meaning that they can be applied to other diseases beyond the treatment of COVID-19 and cancer, signifying their contributions to the future of medicine. Nevertheless, many limitations remain in the development of both vaccines, all of which scientists are finding ways to overcome. Further studies need to be carried out on the stability and delivery of mRNA, but mRNA vaccines remain a promising technology in modern medicine.
References
- Cobb, M. (2015). Who discovered messenger RNA? Current Biology, 25(13), R526–R532. https://doi.org/10.1016/j.cub.2015.05.032 [↩]
- Anand, P., & Stahel, V. P. (2021). The safety of Covid-19 mRNA vaccines: a review. Patient Safety in Surgery, 15(1). https://doi.org/10.1186/s13037-021-00291-9 [↩]
- Gote, V., Bolla, P. K., Kommineni, N., Butreddy, A., Nukala, P. K., Palakurthi, S. S., & Khan, W. (2023). A Comprehensive Review of mRNA Vaccines. International Journal of Molecular Sciences, 24(3), 2700. https://doi.org/10.3390/ijms24032700 [↩]
- Chandra, S., Wilson, J. C., Good, D., & Wei, M. Q. (2024). mRNA vaccines: a new era in vaccine development. Oncology Research, 32(10), 1543–1564. https://doi.org/10.32604/or.2024.043987 [↩]
- Park, J.W., Lagniton, P.N.P., Liu, Y., & Xu, R.-H. (2021). mRNA vaccines for COVID-19: what, why and how. https://www.ijbs.com/ v17p1446.htm. [↩]
- Mayo Clinic (2025). How do different types of COVID-19 vaccines work? https://www.mayoclinic.org/diseases-conditions/coronavirus/in-depth/different-types-of-covid-19-vaccines/ art-20506465. [↩]
- Hao, Q., Long, Y., Yang, Y., Deng, Y., Ding, Z., Yang, L., Shu, Y., & Xu, H. (2024). Development and Clinical Applications of Therapeutic Cancer Vaccines with Individualized and Shared Neoantigens. Vaccines, 12(7), 717–717. https://doi.org/10.3390/vaccines12070717 [↩] [↩] [↩]
- Cui, C., Wang, J., Fagerberg, E., Chen, P.-M., Connolly, K. A., Damo, M., Cheung, J. F., Mao, T., Askari, A. S., Chen, S., Fitzgerald, B., Foster, G. G., Eisenbarth, S. C., Zhao, H., Craft, J., & Joshi, N. S. (2021). Neoantigen-driven B cell and CD4 T follicular helper cell [↩]
- Matheoud, D., Perié, L., Hoeffel, G., Vimeux, L., Parent, I., Marañón, C., Bourdoncle, P., Renia, L., Prevost-Blondel, A., Lucas, B., Feuillet, V., & Hosmalin, A. (2010). Cross-presentation by dendritic cells from live cells induces protective immune responses in vivo. Blood, 115(22), 4412–4420. https://doi.org/10.1182/blood-2009-11-255935 [↩]
- Li, X., You, J., Hong, L., Liu, W., Guo, P., & Hao, X. (2023). Neoantigen cancer vaccines: a new star on the horizon. Cancer Biology & Medicine, 1–38. https://doi.org/10.20892/j.issn.2095-3941.2023.0395 [↩] [↩] [↩]
- Stebbing, J. (2021). An update on mRNA cancer vaccines. https://www.rcpath.org/resource-report/an-update-on-mrna-cancer-vaccines.html. [↩]
- Sikorski, P. J., Warminski, M., Kubacka, D., Ratajczak, T., Nowis, D., Kowalska, J., & Jemielity, J. (2020). The identity and methylation status of the first transcribed nucleotide in eukaryotic mRNA 5′ cap modulates protein expression in living cells. Nucleic Acids Research, 48(4), 1607–1626. https://doi.org/10.1093/nar/gkaa032 [↩]
- Tanguay, R. L., & Gallie, D. R. (1996). Translational efficiency is regulated by the length of the 3’ untranslated region. Molecular and Cellular Biology, 16(1), 146–156. https://doi.org/10.1128/mcb.16.1.146 [↩]
- Pontrelli, L., Sidiropoulos, K. G., & Adeli, K. (2004). Translational Control of Apolipoprotein B mRNA: Regulation via Cis Elements in the 5‘ and 3‘ Untranslated Regions†. Biochemistry, 43(21), 6734–6744. https://doi.org/10.1021/bi049887s [↩]
- Fu, Q., Zhao, X., Hu, J., Jiao, Y., Yan, Y., Pan, X., Wang, X., & Jiao, F. (2025). mRNA vaccines in the context of cancer treatment: from concept to application. Journal of Translational Medicine, 23(1). https://doi.org/10.1186/s12967-024-06033-6 [↩]
- Gallie, D. R. (1991). The cap and poly(A) tail function synergistically to regulate mRNA translational efficiency. Genes & Development, 5(11), 2108–2116. https://doi.org/10.1101/gad.5.11.2108 [↩]
- Jackson, N.A.C., Kester, K.E., Casimiro, D., Gurunathan, S., & DeRosa, F. (2020). The promise of mRNA vaccines: a biotech and industrial perspective. Npj Vaccines, Vol. 5, No. 1, pp. 1–6. https://doi.org/10.1038/s41541-020-0159-8 [↩]
- Deng, Z., Tian, Y., Song, J., An, G., & Yang, P. (2022). mRNA Vaccines: The Dawn of a New Era of Cancer Immunotherapy. Frontiers in Immunology, 13. https://doi.org/10.3389/fimmu.2022.887125 [↩] [↩] [↩] [↩] [↩]
- Tan, T. J. C., Mou, Z., Lei, R., Ouyang, W. O., Yuan, M., Song, G., Andrabi, R., Wilson, I. A., Kieffer, C., Dai, X., Matreyek, K. A., & Wu, N. C. (2023). High-throughput identification of prefusion-stabilizing mutations in SARS-CoV-2 spike. Nature Communications, 14(1). https://doi.org/10.1038/s41467-023-37786-1 [↩]
- Xia, X. (2021). Domains and Functions of Spike Protein in SARS-Cov-2 in the Context of Vaccine Design. Viruses, 13(1), 109. https://doi.org/10.3390/v13010109 [↩] [↩]
- Anderson, B. R., Muramatsu, H., Jha, B. K., Silverman, R. H., Weissman, D., & Kariko, K. (2011). Nucleoside modifications in RNA limit activation of 2’-5’-oligoadenylate synthetase and increase resistance to cleavage by RNase L. Nucleic Acids Research, 39(21), 9329–9338. https://doi.org/10.1093/nar/gkr586 [↩]
- Braun, D. A., Moranzoni, G., Chea, V., McGregor, B. A., Blass, E., Tu, C. R., Vanasse, A. P., Forman, C., Forman, J., Afeyan, A. B., Schindler, N. R., Liu, Y., Li, S., Southard, J., Chang, S. L., Hirsch, M. S., LeBoeuf, N. R., Olive, O., Mehndiratta, A., & Greenslade, H. (2025). A neoantigen vaccine generates antitumour immunity in renal cell carcinoma [↩]
- Sheridan, C. (2025). Individualized mRNA cancer vaccines make strides. Nature Biotechnology, 43(6), 833–836. https://doi.org/10.1038/s41587-025-02708-7 [↩]
- Ni, L. (2023). Advances in mRNA-Based Cancer Vaccines. Vaccines, Vol. 11, No. 10, 1599. https://doi.org/10.3390/vaccines11101599. [↩]
- Bettini, E., & Locci, M. (2021). SARS-CoV-2 mRNA Vaccines: Immunological Mechanism and Beyond. Vaccines, 9(2), 147. https://doi.org/10.3390/vaccines9020147 [↩]
- Szabó, G. T., Mahiny, A. J., & Vlatkovic, I. (2022). COVID-19 mRNA vaccines: Platforms and current developments. Molecular Therapy, 30(5). https://doi.org/10.1016/j.ymthe.2022.02.016 [↩]
- Malone, R. W., Felgner, P. L., & Verma, I. M. (1989). Cationic liposome-mediated RNA transfection. Proceedings of the National Academy of Sciences of the United States of America, 86(16), 6077–6081. https://doi.org/10.1073/pnas.86.16.6077 [↩]
- Pollard, C., Rejman, J., De Haes, W., Verrier, B., Van Gulck, E., Naessens, T., De Smedt, S., Bogaert, P., Grooten, J., Vanham, G., & De Koker, S. (2013). Type I IFN Counteracts the Induction of Antigen-Specific Immune Responses by Lipid-Based Delivery of mRNA Vaccines. Molecular Therapy, 21(1), 251–259. https://doi.org/10.1038/mt.2012.202 [↩]
- Fan, Y.-N., Li, M., Luo, Y.-L., Chen, Q., Wang, L., Zhang, H.-B., Shen, S., Gu, Z., & Wang, J. (2018). Cationic lipid-assisted nanoparticles for delivery of mRNA cancer vaccine. Biomaterials Science, 6(11), 3009–3018. https://doi.org/10.1039/c8bm00908b [↩] [↩]
- Liu, J., Feng, M., Liang, D., Yang, J., & Tang, X. (2016). Vitamin E-Labeled Polyethylenimine for in vitro and in vivo Gene Delivery. Biomacromolecules, 17(10), 3153–3161. https://doi.org/10.1021/acs.biomac.6b00776 [↩]
- Tateshita, N., Miura, N., Tanaka, H., Masuda, T., Ohtsuki, S., Tange, K., Nakai, Y., Yoshioka, H., & Akita, H. (2019). Development of a lipoplex-type mRNA carrier composed of an ionizable lipid with a vitamin E scaffold and the KALA peptide for use as an ex vivo dendritic cell-based cancer vaccine. Journal of Controlled Release, 310, 36–46. https://doi.org/10.1016/j.jconrel.2019.08.002 [↩]
- Liu, L., Wang, Y., Miao, L., Liu, Q., Musetti, S., Li, J., & Huang, L. (2018). Combination Immunotherapy of MUC1 mRNA Nano-vaccine and CTLA-4 Blockade Effectively Inhibits Growth of Triple Negative Breast Cancer. Molecular Therapy, 26(1), 45–55. https://doi.org/10.1016/j.ymthe.2017.10.020 [↩]
- Ndeupen, S., Qin, Z., Jacobsen, S., Bouteau, A., Estanbouli, H., & Igyártó, B. Z. (2021). The mRNA-LNP platform’s lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. IScience, 24(12), 103479. https://doi.org/10.1016/j.isci.2021.103479 [↩]
- Ren, J., Cao, Y., Li, L., Wang, X., Lu, H., Yang, J., & Wang, S. (2021). Self-assembled polymeric micelle as a novel mRNA delivery carrier. Journal of Controlled Release, 338, 537–547. https://doi.org/10.1016/j.jconrel.2021.08.061 [↩]
- Ghosh, B., & Biswas, S. (2021). Polymeric micelles in cancer therapy: State of the art. Journal of Controlled Release, 332, 127–147. https://doi.org/10.1016/j.jconrel.2021.02.016 [↩]
- Chen, C., Pu, F., Huang, Z., Liu, Z., Ren, J., & Qu, X. (2010). Stimuli-responsive controlled-release system using quadruplex DNA-capped silica nanocontainers. Nucleic Acids Research, 39(4), 1638–1644. https://doi.org/10.1093/nar/gkq893 [↩]
- Chen, X., Chen, T., Ren, L., Chen, G., Gao, X., Li, G., & Zhu, X. (2019). Triplex DNA Nanoswitch for pH-Sensitive Release of Multiple Cancer Drugs. ACS Nano, 13(6), 7333–7344. https://doi.org/10.1021/acsnano.9b03846 [↩]
- Garcia-Beltran, W. F., Lam, E. C., Denis, K. S., Nitido, A. D., Garcia, Z. H., Hauser, B. M., Feldman, J., Pavlovic, M. N., Gregory, D. J., Poznansky, M. C., Sigal, A., Schmidt, A. G., Iafrate, A. J., Naranbhai, V., & Balazs, A. B. (2021). Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell, 184(9), 2372-2383.e9. https://doi.org/10.1016/j.cell.2021.03.013 [↩]
- Lustig, Y., Nemet, I., Kliker, L., Zuckerman, N., Yishai, R., Alroy-Preis, S., Mendelson, E., & Mandelboim, M. (2021). Neutralizing Response against Variants after SARS-CoV-2 Infection and One Dose of BNT162b2. New England Journal of Medicine. https://doi.org/10.1056/nejmc2104036 [↩]
- Nemet, I., Kliker, L., Lustig, Y., Zuckerman, N., Erster, O., Cohen, C., Kreiss, Y., Alroy-Preis, S., Regev-Yochay, G., Mendelson, E., & Mandelboim, M. (2021). Third BNT162b2 Vaccination Neutralization of SARS-CoV-2 Omicron Infection. New England Journal of Medicine. https://doi.org/10.1056/nejmc2119358 [↩]
- Brüssow, H. (2021). COVID-19: vaccination problems. Environmental Microbiology, 23(6), 2878–2890. https://doi.org/10.1111/1462-2920.15549 [↩]
- Yang, Q., Jacobs, T. M., McCallen, J. D., Moore, D. T., Huckaby, J. T., Edelstein, J. N., & Lai, S. K. (2016). Analysis of Pre-existing IgG and IgM Antibodies against Polyethylene Glycol (PEG) in the General Population. Analytical Chemistry, 88(23), 11804–11812. https://doi.org/10.1021/acs.analchem.6b03437 [↩]
- Shimabukuro, T. T., Cole, M., & Su, J. R. (2021). Reports of Anaphylaxis After Receipt of mRNA COVID-19 Vaccines in the US—December 14, 2020-January 18, 2021. JAMA, 325(11). https://doi.org/10.1001/jama.2021.1967 [↩]
- Lin, Q., Zhao, Q., & Lev, B. (2020). Cold chain transportation decision in the vaccine supply chain. European Journal of Operational Research, 283(1), 182–195. https://doi.org/10.1016/j.ejor.2019.11.005 [↩]
- Uddin, M. N., & Roni, M. A. (2021). Challenges of Storage and Stability of mRNA-Based COVID-19 Vaccines. Vaccines, 9(9), 1033. https://doi.org/10.3390/vaccines9091033 [↩]
- Zhao, P., Hou, X., Yan, J., Du, S., Xue, Y., Li, W., Xiang, G., & Dong, Y. (2020). Long-term storage of lipid-like nanoparticles for mRNA delivery. Bioactive Materials, 5(2), 358–363. https://doi.org/10.1016/j.bioactmat.2020.03.001 [↩]
- Rosenthal, R., Cadieux, E. L., Salgado, R., Bakir, M. A., Moore, D. A., Hiley, C. T., Lund, T., Tanić, M., Reading, J. L., Joshi, K., Henry, J. Y., Ghorani, E., Wilson, G. A., Birkbak, N. J., Jamal-Hanjani, M., Veeriah, S., Szallasi, Z., Loi, S., Hellmann, M. D., & Feber, A. (2019). Neoantigen-directed immune escape in lung cancer evolution. Nature, 567(7749), 479–485. https://doi.org/10.1038/s41586-019-1032-7 [↩]
- Anderson, K., Lutz, C., van Delft, F. W., Bateman, C. M., Guo, Y., Colman, S. M., Kempski, H., Moorman, A. V., Titley, I., Swansbury, J., Kearney, L., Enver, T., & Greaves, M. (2011). Genetic variegation of clonal architecture and propagating cells in leukaemia. Nature, 469(7330), 356–361. https://doi.org/10.1038/nature09650 [↩]
- Lohr, Jens G., Stojanov, P., Carter, Scott L., Cruz-Gordillo, P., Lawrence, Michael S., Auclair, D., Sougnez, C., Knoechel, B., Gould, J., Saksena, G., Cibulskis, K., McKenna, A., Chapman, Michael A., Straussman, R., Levy, J., Perkins, Louise M., Keats, Jonathan J., Schumacher, Steven E., Rosenberg, M., & Getz, G. (2014). Widespread Genetic Heterogeneity in Multiple Myeloma: Implications for Targeted Therapy. Cancer Cell, 25(1), 91–101. https://doi.org/10.1016/j.ccr.2013.12.015 [↩]
- Bolli, N., Avet-Loiseau, H., Wedge, D. C., Van Loo, P., Alexandrov, L. B., Martincorena, I., Dawson, K. J., Iorio, F., Nik-Zainal, S., Bignell, G. R., Hinton, J. W., Li, Y., Tubio, J. M. C., McLaren, S., O’ Meara, S., Butler, A. P., Teague, J. W., Mudie, L., Anderson, E., & Rashid, N. (2014). Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nature Communications, 5(1). https://doi.org/10.1038/ncomms3997 [↩]
- Rupaimoole, R., Calin, G. A., Lopez-Berestein, G., & Sood, A. K. (2016). miRNA Deregulation in Cancer Cells and the Tumor Microenvironment. Cancer Discovery, 6(3), 235–246. https://doi.org/10.1158/2159-8290.CD-15-0893 [↩]
- Hinshaw, D. C., & Shevde, L. A. (2019). The Tumor Microenvironment Innately Modulates Cancer Progression. Cancer Research, 79(18), 4557–4566. https://doi.org/10.1158/0008-5472.can-18-3962 [↩]
- Fan, T., Zhang, M., Yang, J., Zhu, Z., Cao, W., & Dong, C. (2023). Therapeutic cancer vaccines: advancements, challenges, and prospects. Signal Transduction and Targeted Therapy, 8(1). https://doi.org/10.1038/s41392-023-01674-3 [↩]
- Matthias Birtel, Voss, R.-H., Reinhard, K., Rengstl, B., Yasmina Ouchan, Michel, K., Hayduk, N., Tillmann, B., Becker, R., Suchan, M., Theobald, M., Oehm, P., Oezlem Tuereci, & Sahin, U. (2022). A TCR-like CAR Promotes Sensitive Antigen Recognition and Controlled T-cell Expansion Upon mRNA Vaccination. Cancer Research Communications, 2(8), 827–841. https://doi.org/10.1158/2767-9764.crc-21-0154 [↩]
- Lin, F., Lin, E., Misa Anekoji, Ichim, T. E., Hu, J., Marincola, F. M., Jones, L., Santosh Kesari, & Shashaanka Ashili. (2023). Advancing personalized medicine in brain cancer: exploring the role of mRNA vaccines. Journal of Translational Medicine, 21(1). https://doi.org/10.1186/s12967-023-04724-0 [↩]
- Weber, J. S., Carlino, M. S., Khattak, A., Tarek Meniawy, Ansstas, G., Taylor, M. H., Kim, K. B., McKean, M., Long, G. V., Sullivan, R. J., Faries, M., Tran, T. T., C Lance Cowey, Pecora, A., Shaheen, M., Segar, J., Medina, T., Atkinson, V., Gibney, G. T., & Luke, J. J. (2024). Individualised neoantigen therapy mRNA-4157 (V940) plus pembrolizumab versus pembrolizumab monotherapy in resected melanoma (KEYNOTE-942): a randomised, phase 2b study. The Lancet, 403(10427). https://doi.org/10.1016/s0140-6736(23)02268-7 [↩]
- Horne, E. M. F., Hulme, W. J., Parker, E. P. K., Keogh, R. H., Williamson, E. J., Walker, V. M., Palmer, T. M., Denholm, R., Knight, R., Curtis, H. J., Walker, A. J., Andrews, C. D., Mehrkar, A., Morley, J., MacKenna, B., Bacon, S. C. J., Goldacre, B., Hernán, M. A., & Sterne, J. A. C. (2024). Effectiveness of mRNA COVID-19 Vaccines as First Booster Doses in England: An Observational Study in OpenSAFELY-TPP. Epidemiology, 35(4), 568–578. https://doi.org/10.1097/ede.0000000000001747 [↩]






