Abstract
Neurodevelopmental disorders (NDDs) are disorders with complex genetic etiology that arise during brain development, such as Autism spectrum disorder (ASD), Attention-Deficit/Hyperactivity Disorder (ADHD), and Schizophrenia (SCZ). MicroRNAs (miRNAs), a class of non-coding RNAs, play a crucial role in regulating gene expression, making them key targets for interventions involving neurodevelopmental pathways. This study employs a bioinformatics approach to identify differentially expressed miRNAs and genes using three publicly available GEO datasets (GSE15888, GSE98088, and GSE100670) from which the data were further validated using TargetScan and miRTarBase. High-confidence interactions were identified using miRNet for use in network construction. A network was constructed and analysed, and the most significant genes and miRNAs were identified using Cytoscape. Similarly, the most significant pathways were identified using Reactome. Finally, the disease associations and therapeutic potential of the identified miRNAs and genes were evaluated using the Open Targets platform, miRBase, and Drug-Gene Interaction Database (DGIdb), producing a final list of potential therapeutic targets. The genes CLTC and PTEN were found to be associated with NDDs, and five miRNAs were found to interact with these genes, posing the interactions between both as promising therapeutic targets. These findings offer insights into the regulatory networks involved in neurodevelopment as well as potential biomarkers and therapeutic targets for functional validation and clinical application.
Keywords: miRNA, Genes, Network Biology, Pathway Analysis, Neurodevelopment disorders.
Introduction
Neurodevelopmental disorders are behavioural and cognitive disorders that arise during brain development, and impact intellectual, social, and motor functions1. Among these disorders, ASD and ADHD diagnoses have risen substantially over time, with ASD diagnosis rates increasing by 175% between 2011 and 20222, and ADHD prevalence ranging from a 22% to 54% increase over the last few decades in children and adults in the US3. This general increase in the prevalence of neurodevelopmental disorders highlights the need for research into their etiology as well as potential therapeutic targets. Over the past decade, researchers worldwide have demonstrated that microRNAs play essential regulatory roles in neurodevelopment, influencing neuronal differentiation, synapse formation, and circuit maturation4,5,6.
MicroRNAs (miRNAs) are small, non-coding RNAs that regulate gene expression post-transcriptionally, by binding to the 3’ untranslated region (3’ UTR) of target mRNAs and either degrading or suppressing translation7. Each miRNA can bind to several targets, thus controlling a large number of molecular pathways. miRNAs are essential for neurodevelopment, regulating processes such as neuronal differentiation and migration, axon and dendritic outgrowth and complexity, and synaptic development8, 9. Therefore, they are likely to play an important role in the formation of neurodevelopmental disorders. A previous study found that miR-137 is crucial for neurodevelopment and may contribute to the development of NDDs. They have identified miRNAs associated with disorders such as ASD, Schizophrenia (SCZ), and ADHD through bioinformatic approaches10. In another study, miR-873 was found to regulate ASD risk genes (ARID1B, SHANK3 and NRXN2) in charge of neuronal morphology and cell differentiation, using approaches such as animal testing and qPCR11. A combination of 4 miRNAs was also identified (miR-4433b-5p, miR-15a-5p, miR-335-5p, and miR-1180-3p) to be a biomarker for the early diagnosis of ASD, through pathway analysis and RNA sequencing12. A common bioinformatic technique that has shown promising results is differential analysis, which was used in a study investigating miRNA expression in high myopia and age‑related cataracts, and identified the miRNAs involved13. Despite all the research that has already been done, it is still unclear how miRNAs integrate into broader processes in the brain due to the complexity of neurodevelopment. Most studies in the field focus on single miRNAs, oversimplifying their role by ignoring broader networks. In reality, miRNAs operate in coordinated systems and there is yet to be a comprehensive construction of miRNA-mediated gene regulatory networks involved in neurodevelopment.
A bioinformatics approach in this study eliminates the concerns about cohort sizes and diversity, costs of experimental validation, and reproducibility, that previous studies have faced. Diverse datasets will be combined to identify broader patterns and data will be collated from different sources, leading to more exhaustive research. Tools such as GEO, TargetScan, and Cytoscape will be utilized in identifying the relevant miRNAs and genes, gathering expression datasets, and predicting gene interactions to construct regulatory networks. With the increasing prevalence of neurodevelopmental disorders comes a need for the further exploration of poorly understood targets, such as miRNAs.
This paper aims to identify the role of miRNA-gene interactions in neurodevelopment by integrating publicly available datasets and interaction databases. Specifically, the objectives are to identify key differentially expressed miRNAs and genes associated with relevant NDDs such as ASD and ADHD, using a bioinformatics-based analysis to construct gene regulatory networks and identify therapeutic targets. Furthermore, the study aims to associate the shortlisted targets with specific NDDs. This research can serve as a basis for other studies on miRNA-associated neurodevelopmental disorders by outlining the specific gene regulatory networks at play during the development of the brain. The findings could be used to design therapies and preventive interventions, as well as identify biomarkers at the onset of neurodevelopmental disorders.
Methodology
Data Collection and Processing
Publicly available datasets were retrieved from the Gene Expression Omnibus (GEO) to identify differentially expressed (DE) miRNAs and genes associated with neurodevelopmental disorders. GSE15888 provided miRNA expression data, while GSE98088 and GSE100670 provided gene expression data. Although these datasets are not linked, they were used to check the expression profiles of both miRNAs and genes in NDDs, allowing for a bioinformatic analysis of miRNA-gene regulatory interactions in neurodevelopmental contexts. Although these datasets are derived from neural stem cells (NSC) rather than from patients with NDDs, NSC-based models are widely used in such investigations, as NDDs originate during early embryonic brain development, a stage that cannot be accessed in vivo14. Furthermore, miRNA expression changes during this stage and has been shown to regulate neural stem cell proliferation, differentiation, and lineage commitment, revealing the effects relevant to disease mechanisms15. Similar approaches have been used in several published studies. For example, studies on miR-137 have demonstrated their roles in neural differentiation and NDD-related pathways using stem-cell–based models16. Datasets from miRTarBase and TargetScan containing miRNA and gene interactions were also downloaded and used to identify the interactions between genes and miRNAs. miRTarBase provides experimentally validated miRNA-target interactions, while TargetScan predicts computationally inferred miRNA-gene interactions based on sequence complementarity and evolutionary conservation17. The inclusion of both datasets ensured that both established and potential interactions were integrated.
Differential Expression Analysis
Differential expression analysis was performed using the GEO2R tool, through which samples were grouped into test and control groups. The dataset GSE15888 provided data about miRNA profiles at different stages of human neural development, using NT2 cells treated with Retinoic Acid. The control group was set to untreated cells, and the test group to cells treated with retinoic acid. For GSE98088, which provided data on how a mutation in miR-873-5p affects the regulation of genes critical for neurodevelopment, the control group was set to wild-type (WT) miR-873-5p, and the test group to mutant (Mut) miR-873-5p. GSE100670 investigates how miR-146a influences the differentiation of human neural stem cells into specific neural cell types, with implications for disorders like ASD. The control group was set to miR-146a, and the test group to mutant miR-146a (miR-146a-Mut). Using the data gathered, the gene datasets were merged, and statistically significant nodes were selected using a threshold of p-value < 0.05. To obtain a final list of genes and miRNAs, duplicate entries were then removed. All statistical tests, including pathway enrichment and differential expression, were corrected for multiple hypothesis testing using the Benjamini–Hochberg FDR procedure. Significance was defined as FDR < 0.05.
Network and Pathway Analysis
After obtaining the two lists of DE genes and miRNA, the interactions were identified by filtering for the miRNAs and genes that were in common with the lists from both miRTarBase and TargetScan. Microsoft Excel was used to align each miRNA to its target genes and establish the interactions. Since this resulted in a large list, we filtered only high-confidence interactions for use in constructing a network, for computational feasibility. Only miRNAs and genes with two or more experimental and literature evidence were considered high confidence, and this was done by cross-referencing from the miRNet database. The construction of a network was done using the Cytoscape software, where the miRNAs and genes are represented by nodes and the regulatory interactions are represented by edges. The final list of filtered interactions was used as the input to generate a network and analyse nodes based on various topological parameters. After further processing the data, the top 10 miRNAs and genes involved in neurodevelopmental disorders were identified. Using the analysis tool of Reactome, pathway analysis was carried out. The gene list of the network was used as input to identify the significant pathways. To reduce node-degree bias, we performed degree-preserving network permutations and evaluated enrichment against established neurodevelopmental disorder (NDD) gene sets. Only pathways that remained significant after bias correction and were enriched for NDD-associated genes were retained. A search was conducted for the presence of the top 10 DE genes in the top 25 pathways.
Disease Association and Therapeutic Potential Analysis
The top 10 DE genes and miRNAs identified were examined for previously known associations with NDDs such as ASD, ADHD, SCZ, and intellectual disability (ID). To find the disease associations for the DE genes, the Open Targets Platform was used. A search was conducted with each gene, and the results were filtered to display only ‘disease or phenotype’ results. For DE miRNAs, disease associations were obtained from HMDD v3.2 and miRNet, which curate experimentally supported miRNA–disease relationships. miRBase was used solely for retrieving miRNA sequences and annotation. A similar process was followed, in which a search was conducted, and the results were examined for associations with NDDs. The DE genes and miRNAs that showed associations with NDDs were recorded. Following the disease association analysis, the Drug-Gene Interaction Database (DGIdb) was used to evaluate the therapeutic potential of each of the disease-associated miRNAs and genes. Each of the genes was entered, and the top approved drugs with the highest interaction scores were recorded. The disease-associated miRNAs were examined to see if they interact with the disease-associated genes. This reveals whether the miRNA-gene interactions can be targeted for therapeutic interventions.
Results
Differential Expression Analysis
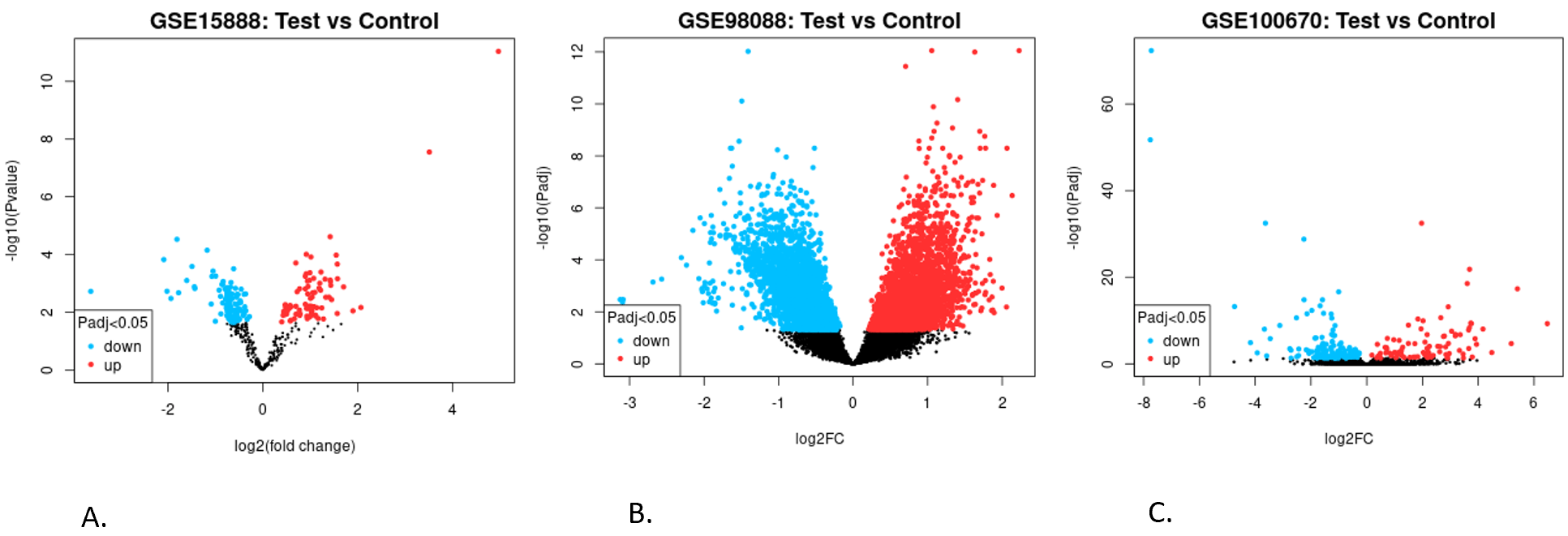
The DE analysis revealed the differentially expressed genes and miRNAs. GSE15888 had 5 control samples and 24 test samples. The DE analysis yielded 283 miRNAs that were significantly DE as they had a p-value<0.05. The GSE98088 had 8 control and 8 test samples, and analysis identified 7953 DE genes. Similarly, GSE100670 had 6 control and 5 test samples, and 229 DE genes were identified. The volcano plots for the differentially expressed genes for each of the studies are shown in Figure 1. The differential expression analysis (Figure 1) replicates the raw dataset processing to enable consistent cross-dataset integration with miRNA regulatory data. While individual DE results are consistent with original publications, here the novelty lies in combining DEGs with high-confidence miRNA targeting relationships to construct a unified miRNA–mRNA regulatory model of neurodevelopmental disorders. Both gene lists were combined, and duplicate entries were removed, resulting in a total of 8095 DE genes.
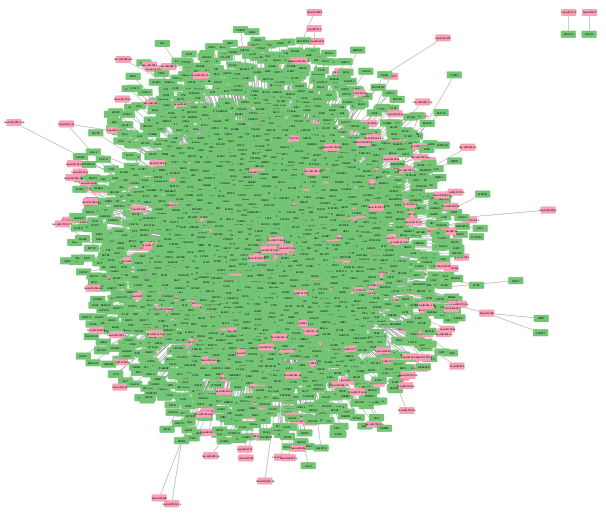
Network and Pathway Analysis
The miRTarBase had 27595 interactions, and TargetScan had 1048575. 148 DE miRNAs and 1913 DE genes from the GSE studies were present in both the databases, from which a list of 318102 total interactions was established. Only 32395 met the criteria for high confidence interactions, which were found using miRNet, which was further visualised in Cytoscape. The resulting Cytoscape network revealed 32395 edges and 4641 nodes, of which 391 were miRNAs and 4250 were genes, as shown in Figure 2. This network was analysed based on the following parameters: AverageShortestPathLength, BetweennessCentrality, ClosenessCentrality, ClusteringCoefficient, Eccentricity, EdgeCount, Indegree, Outdegree, NeighborhoodConnectivity, IsSingleNode, PartnerOfMultiEdgedNodePairs, SelfLoops, and Stress. Out of these, the 4 parameters, BetweennessCentrality, ClosenessCentrality, ClusteringCoefficient, and Degree were selected. These were prioritised as they best estimate the connectivity and influence of a node. High values of Betweenness Centrality indicate nodes that control information flow, as it measures how often a node lies on the shortest path between other nodes. High Closeness Centrality identifies nodes that can interact with others more efficiently, as it measures how close a node is to all the others. Similarly, Clustering Coefficient measures how connected a node’s neighbors are to each other. Lastly, nodes with high Degrees are often key regulatory nodes as it indicates the number of edges a node has. The Degree was sorted from highest to lowest to highlight the most significant nodes. The top 10 significant miRNAs and genes based on the topological parameters are listed in Table 1. The miRNAs and genes in the table do not correspond.
| S. No. | miRNAs | Genes |
| 1 | hsa-miR-34a-5p | CCND1 |
| 2 | hsa-miR-16-5p | CLTC |
| 3 | hsa-miR-7-5p | NUFIP2 |
| 4 | hsa-miR-124-3p | PTEN |
| 5 | hsa-miR-92a-3p | IGF1R |
| 6 | hsa-miR-29b-3p | MYC |
| 7 | hsa-miR-27a-3p | PANK3 |
| 8 | hsa-miR-17-5p | CDK6 |
| 9 | hsa-miR-1-3p | NUCKS1 |
| 10 | hsa-miR-21-5p | HIF1A |
The top 5 pathways identified were Diseases of signal transduction by growth factor receptors and second messengers, Signaling by Receptor Tyrosine Kinases, Signal Transduction, Intracellular signaling by second messengers, and PIP3 activates AKT signaling. Out of the 10 genes, 7 were present in the top 25 pathways – CCND1, CLTC, PTEN, IGF1R, MYC, CDK6, and HIF1A.
Disease Association and Therapeutic Potential Analysis
After performing disease association analysis for DE genes, two out of the ten genes were found to be associated with NDDs, which were CLTC and PTEN. PTEN has well-established associations with macrocephaly, autism spectrum disorder, and disruptions in neuronal growth and synaptic regulation. In contrast, CLTC variants primarily contribute to epilepsy, intellectual disability, and abnormal synaptic vesicle trafficking, but are not associated with macrocephaly-autism phenotypes. Similarly, five out of the 10 miRNAs were found to be associated with NDDs, which were hsa-miR-92a-3p, hsa-miR-29b-3p, hsa-miR-27a-3p, hsa-miR-21-5p, and hsa-miR-7-5p. From the miRNAs, hsa-miR-92a-3p and hsa-miR-29b-3p were associated with SCZ (Ma et al. 2018; Wang et al. 2014). Furthermore, hsa-miR-27a 3p and hsa-miR-21-5p were associated with ASD(Frye et al. 2021). Lastly, hsa-miR-7-5p was associated with both ASD and SCZ (Huang et al. 2021)(Zhang et al. 2015).
Using DGIdb, it was found that CLTC showed therapeutic potential, having the approved drugs Folic Acid and Sulfasalazine already available, each with an interaction score of 0.31. Similarly, PTEN showed therapeutic potential with the approved top medicines being Capivasertib and Capmatinib, each with an interaction score of 0.31 and 0.23, respectively. The reported interaction scores represent the strength of existing evidence supporting the drug-gene interactions, based on the number and quality of data sources. However, these scores do not imply that the drug works effectively for the diseases. Moreover, the approved drugs mentioned interact with CLTC/PTEN in other disease contexts (oncology, inflammation), but there is no evidence linking them to neurodevelopmental disorders. Their relevance is limited to suggesting possible mechanistic pathways, not therapeutic applicability. From the miRNAs, hsa-miR-92a-3p, hsa-miR-29b-3p, hsa-miR-27a-3p, and hsa-miR-21-5p were found to interact with both PTEN and CLTC, while hsa-miR-7-5p was found to interact with only CLTC. This further confirms that the CLTC and PTEN are effective therapeutic targets. Also, the interactions between these two genes and the five miRNAs are good therapeutic targets. By targeting the interactions between the CLTC, PTEN, and miRNAs, the progression of neurodevelopment disorder-related conditions can be prevented.
Discussion
This study aimed to use bioinformatic tools to identify the key miRNAs, genes, and regulatory pathways associated with NDDs, and assess their disease association and therapeutic potential. To achieve this, expression profiles of three GEO studies were retrieved and analysed. The overall findings from the study show that the genes CLTC and PTEN, and the miRNAs hsa-miR-92a-3p, hsa-miR-29b-3p, hsa-miR-27a-3p, hsa-miR-21-5p, and hsa-miR-7-5p, play a significant role in NDDs. Furthermore, the pathways Diseases of signal transduction by growth factor receptors and second messengers, Signaling by Receptor Tyrosine Kinases, Signal Transduction, Intracellular signaling by second messengers, PIP3 activates AKT signaling were found to be the five most heavily involved pathways in NDDs. This information provides insights on potential therapeutic targets.
These results align with existing research suggesting that certain miRNAs and genes (CLTC, PTEN, miRNAs hsa-miR-92a-3p, hsa-miR-29b-3p, hsa-miR-27a-3p, hsa-miR-21-5p, and hsa-miR-7-5p) are found to be differentially expressed in people with NDDs and therefore play a pivotal role in their etiology 18. Several of the identified miRNAs, including miR-125a-5p, miR-338-3p, and miR-873, have previously been implicated in ASD through their roles in synapse maturation, cortical development, and neuronal connectivity. Dysregulation of these miRNAs may contribute to ASD phenotypes by altering the expression of genes such as ARID1B, NRXN2, and SHANK3, all of which are known ASD risk genes. In ADHD, altered dopamine signaling and impaired fronto-striatal circuitry have been linked to differential expression of miRNAs involved in neurotransmitter release and synaptic vesicle cycling19,20. Our identification of CLTC as a central network node aligns with its role in synaptic vesicle recycling, suggesting potential mechanistic overlap with ADHD pathology. Schizophrenia-associated miRNAs (e.g., miR-125b, miR-137) regulate neuronal migration, synaptic pruning, and glutamatergic signaling. Several of the pathways enriched in our analysis, including axon guidance, MAPK signaling, and neurotrophin signaling, are well-established contributors to schizophrenia neurobiology21,22. Collectively, these findings suggest that the miRNA–gene modules identified in our study may serve as promising therapeutic entry points. miRNA mimics or inhibitors, pathway-modulating drugs, or gene-specific interventions may help normalize dysregulated networks in ASD, ADHD, and schizophrenia.
CLTC has been known to play a role in neuronal transmission by facilitating the recycling and release of vesicles at the presynaptic terminals of neurons23. Its association with macrocephaly-autism syndrome and neurodevelopmental delay is in line with previous studies showing its role in neurodevelopment. Similarly, PTEN is crucial in neurogenesis, neuronal migration, and synaptic plasticity24, and its association to autosomal dominant non-syndromic intellectual disability highlights its role in neurodevelopment as per previous studies. While the two genes are already targeted by drugs, the miRNAs show therapeutic potential due to their interactions with PTEN and CLTC. These interactions can be targeted by drugs to better manage the symptoms of and improve clinical outcomes for individuals with NDDs. PTEN and CLTC emerged as central regulatory nodes. PTEN loss disrupts neuronal growth and synaptic stability, making it a candidate for targeted miRNA modulation, particularly in ASD. CLTC variants impair synaptic vesicle cycling, suggesting therapeutic relevance for miRNAs regulating this gene in epilepsy and cognitive disorders. Modulating these miRNA–gene interactions may therefore offer avenues to correct aberrant neurodevelopmental signaling. PTEN is an established therapeutic target in autism-macrocephaly syndromes due to its regulatory role in synaptic development, PI3K–AKT–mTOR signaling, and neuronal growth25,26. CLTC, though not associated with macrocephaly, is implicated in neurodevelopmental epilepsy and synaptic endocytosis deficits, making it a candidate for therapeutic investigation in epilepsy-related NDDs. PTEN mutations cause macrocephaly-autism and disrupt synaptic plasticity via mTOR hyperactivation. Therapeutic strategies targeting PI3K–AKT–mTOR signaling have shown promise in PTEN-related ASD models. CLTC dysfunction leads to impaired synaptic vesicle formation and has been linked to epileptic encephalopathy and cognitive dysfunction, suggesting that modulation of synaptic endocytosis pathways warrants further exploration27,28,29.
A major strength in this study is the integrated use of different bioinformatics tools and techniques. Various validated databases and tools such as TargetScan, DGIdb, and Cytoscape were used to attain results, ensuring robustness. Similarly, techniques such as DE analysis and network analysis provide a holistic and comprehensive view of the role played by miRNAs and genes in NDDs. However, one limitation is the reliance on publicly available datasets which may not represent a global population due to varying experimental conditions and relatively small sample sizes. Moreover, this study lacks functional validation without which the interactions are only predictions, therefore limiting its use in clinical practice. Future research should focus on experimentally validating the top miRNAs and genes identified as therapeutic targets, through techniques such as CRISPR/Cas9 knockdown or RNA sequencing. Additionally, expanding the dataset to include a more diverse population could enhance the generalizability of the findings.
In conclusion, this study demonstrates the power of bioinformatics tools in uncovering key miRNAs and genes involved in NDDs. The identification of the genes CLTC and PTEN, several other miRNAs, and the top regulatory pathways provides valuable insights into the mechanisms underlying NDDs, allowing them to serve as potential therapeutic targets. Overall, the findings of this study contribute to a growing body of research into the genetic etiology of NDDs. Bioinformatics continues to advance, with the potential to improve diagnostics, personalised therapies, and our understanding of complex NDDs.
Acknowledgments
AJ would like to acknowledge Aashna Saraf, Founder of CreatED for providing valuable feedback and guidance throughout the project.
References
- Sherr, Chapter 36 – Neurodevelopmental Disorders, Causes, and Consequences. JournalBook, 587-599 (2016). [↩]
- Grosvenor, Croen, Lynch, Marafino, Maye, Penfold, Simon and Ames, Autism Diagnosis Among US Children and Adults, 2011-2022. JournalBook, e2442218 (2024). [↩]
- Safer, Is ADHD Really Increasing in Youth? JournalBook, 107-115 (2018). [↩]
- Chakrabortty, Patton, Smith and Agarwal, miRNAs: Potential as Biomarkers and Therapeutic Targets for Cancer. JournalBook, (2023). [↩]
- de Souza, Warren Bezerra, de Oliveira, de Morais Gomes, de Melo Rêgo, Pereira, da Rocha Pitta and da Rosa, MicroRNAs in neuroplasticity: a comprehensive review of mechanisms and therapeutic strategies for neurodegenerative diseases. JournalBook, 97-106 (2025). [↩]
- Rashidi, Kalirad, Rafie, Behzad and Dezfouli, The role of microRNAs in neurobiology and pathophysiology of the hippocampus. JournalBook, (2023). [↩]
- Ratti, Lampis, Ghidini, Salati, Mirchev, Valeri and Hahne, MicroRNAs (miRNAs) and Long Non-Coding RNAs (lncRNAs) as New Tools for Cancer Therapy: First Steps from Bench to Bedside. JournalBook, 261-278 (2020). [↩]
- Cao, Li and Chan, MicroRNAs: Key Regulators in the Central Nervous System and Their Implication in Neurological Diseases. JournalBook, (2016). [↩]
- Brum, Paixão-Côrtes, Carvalho, Martins-Silva, Carpena, Ulguim, Luquez, Salatino-Oliveira and Tovo-Rodrigues, Genetic variants in miRNAs differentially expressed during brain development and their relevance to psychiatric disorders susceptibility. JournalBook, 456-467 (2021). [↩]
- Xu, Wang, Mao, Jiao, Li, Zhao, Sun, Feng, Lai, Peng, Fu, Gan, Chen, Zhao, Wei and Cheng, Generation and transcriptomic characterization of MIR137 knockout miniature pig model for neurodevelopmental disorders. JournalBook, 86 (2024). [↩]
- Lu, Zhu, Williams, Watts, Tonta, Coleman, Parkington and Claudianos, Autism-associated miR-873 regulates ARID1B, SHANK3 and NRXN2 involved in neurodevelopment. JournalBook, 418 (2020). [↩]
- Salloum-Asfar, Ltaief, Taha, Nour-Eldine, Abdulla and Al-Shammari, MicroRNA Profiling Identifies Age-Associated MicroRNAs and Potential Biomarkers for Early Diagnosis of Autism. JournalBook, (2025). [↩]
- Huang, Chen, Wu, Zheng, Huang, Wan and Hu, Comprehensive bioinformatics analysis of metabolism‑related microRNAs in high myopia in young and old adults with age‑related cataracts. JournalBook, (2025). [↩]
- Makropoulos, Counsell and Rueckert, A review on automatic fetal and neonatal brain MRI segmentation. JournalBook, 231-248 (2018). [↩]
- Jiang, Jang and Zeng, The Effects of Extrinsic and Intrinsic Factors on Neurogenesis. JournalBook, (2023). [↩]
- Mahmoudi and Cairns, MiR-137: an important player in neural development and neoplastic transformation. JournalBook, 44-55 (2017). [↩]
- Leclercq, Diallo and Blanchette, Prediction of human miRNA target genes using computationally reconstructed ancestral mammalian sequences. JournalBook, 556-566 (2017). [↩]
- Mushtaq, Greig, Anwar, Zamzami, Choudhry, Shaik, Tamargo and Kamal, miRNAs as Circulating Biomarkers for Alzheimer’s Disease and Parkinson’s Disease. JournalBook, 217-225 (2016). [↩]
- Frye, Rose, McCullough, Bennuri, Porter-Gill, Dweep and Gill, MicroRNA Expression Profiles in Autism Spectrum Disorder: Role for miR-181 in Immunomodulation. JournalBook, (2021). [↩]
- Huang, Chen, Guo and Chen, Systematic Review and Bioinformatic Analysis of microRNA Expression in Autism Spectrum Disorder Identifies Pathways Associated With Cancer, Metabolism, Cell Signaling, and Cell Adhesion. JournalBook, (2021). [↩]
- He, Lozano, Stringer, Watanabe, Sakamoto, den Oudsten, Koopmans, Giamberardino, Hammerschlag, Cornelisse, Li, van Weering, Posthuma, Smit, Sullivan and Verhage, MIR137 schizophrenia-associated locus controls synaptic function by regulating synaptogenesis, synapse maturation and synaptic transmission. JournalBook, 1879-1891 (2018). [↩]
- Olde Loohuis, Nadif Kasri, Glennon, van Bokhoven, Hébert, Kaplan, Martens and Aschrafi, The schizophrenia risk gene MIR137 acts as a hippocampal gene network node orchestrating the expression of genes relevant to nervous system development and function. JournalBook, 109-118 (2017). [↩]
- Sveistrup and Myers, Mosaic CLTC pathogenic variant causing focal epilepsy with normal intelligence. JournalBook, 875-878 (2024). [↩]
- Chwin and Kiryk, The integral role of PTEN in brain function: from neurogenesis to synaptic plasticity and social behavior. JournalBook, 309-318 (2025). [↩]
- Skelton, Stan and Luikart, The Role of PTEN in Neurodevelopment. JournalBook, 60-71 (2020). [↩]
- Li, Liu and Ross, PTEN regulation of neural development and CNS stem cells. JournalBook, (2003). [↩]
- Zahedi Abghari, Moradi and Akouchekian, PTEN gene mutations in patients with macrocephaly and classic autism: A systematic review. JournalBook, 10 (2019). [↩]
- Marchese, Conti, Valvo, Moro, Muratori, Tancredi, Santorelli, Guerrini and Sicca, Autism-epilepsy phenotype with macrocephaly suggests PTEN, but not GLIALCAM, genetic screening. JournalBook, 26 (2014). [↩]
- Nabais Sá, Venselaar, Wiel, Trimouille, Lasseaux, Naudion, Lacombe, Piton, Vincent-Delorme, Zweier, Reis, Trollmann, Ruiz, Gabau, Vetro, Guerrini, Bakhtiari, Kruer, Amor, Cooper, Bijlsma, Barakat, van Dooren, van Slegtenhorst, Pfundt, Gilissen, Willemsen, de Vries, de Brouwer and Koolen, De novo CLTC variants are associated with a variable phenotype from mild to severe intellectual disability, microcephaly, hypoplasia of the corpus callosum, and epilepsy. JournalBook, 797-802 (2020). [↩]