Abstract
Over past decades, solutions offered by synthetic biology have solved numerous pressing issues in multiple fields. For instance, it has offered solutions to medical problems via CRISPR-derived therapies and agricultural problems via genetic modifications of crops. In recent years, it has shifted from developing single-purpose, isolated tools to building integrated, systemized mechanisms that can be applied to real-world scenarios. A literature search was conducted for peer-reviewed articles published from January 2010 to October 2025. This search covered major academic databases such as PubMed and Google Scholar. Eventually, 45 records were selected and 12 studies were included for the final review. This study examines the development, current status, and future prospects of synthetic biology in medicine and agriculture. It gathers evidence from primary research and scientific literature to compare the key areas in which synthetic biology can be applied, identify major limitations in such technology, and analyze potential. The literature suggests that synthetic biology is progressing from single proof-of-concept experiments and aiming toward clinically and agriculturally applicable systems, based upon gene editing and engineered microbial communities. However, potential advances could be limited by an incomplete understanding of the intricacy of natural environments, variability, and ethical issues. In short, synthetic biology is likely to play a major role in future healthcare and agriculture, but its long-term impact relies on responsible regulation and improved control over engineered systems.
Keywords: synthetic biology; CRISPR; gene editing; immunotherapy; CAR-T; engineered microbes; biotechnology
Introduction
Rare genetic diseases affect approximately 300 million people worldwide1. And yet, many of these individuals have limited access to effective treatments. This is where synthetic biology can take a role; it is an interdisciplinary field that combines engineering principles with biology to design and construct new biological systems. With developments such as engineered microbial factories and programmable gene circuits, synthetic biology has already shown its ability to address biomedical problems. As the world faces challenges such as genetic disorders, malnutrition, and insufficient food security, synthetic biology has been applied to develop new approaches. It may significantly influence human medicine and agriculture, solving those challenges in the meantime. For instance, according to the FAO et al. (2023), approximately 735–783 million people worldwide were affected by undernourishment in 20222.
This highlights the urgent need for innovative agricultural developments that enhance crop resilience to environmental changes.
Despite increasing research and investment in the field, several limitations remain. As of the status quo, current reviews focus on isolated applications in one particular field, either in medicine or agriculture; few provide systematic frameworks that evaluate how lessons learned in one field can assist in the development of the other. Consequently, a more comprehensive analysis is required to assess the history, current state, and potential of synthetic biology.
This study provides such an analysis of synthetic biology’s evolution, current status, and future prospects in both pharmaceutical/medical and agricultural fields, in an attempt to address the aforementioned critical gap. The paper consists of three parts. First, it outlines the field’s historical development, such as the invention of recombinant DNA techniques and CRISPR. Subsequently, it analyzes the status quo, noting current advances, successes, and technical challenges. Finally, it speculates on future advancements, investigating how novel therapeutics can contribute to medicine and how new crop engineering strategies can transform agriculture.
Literature Review
Definition of Synthetic Biology
Synthetic biology is commonly described as a blend between engineering and biology. The National Human Genome Research Institute defines it as “a field of science that involves redesigning organisms for useful purposes by engineering them to have new abilities”3. In synthetic biology, genetic components can be treated as standardized parts in a “genetic circuit”; by combining this with computational modeling, the field facilitates the construction of new metabolic pathways. These new pathways pave the way for breakthroughs in targeted therapies and climate-resilient crops.
In the 1960s, Jacques Monod and François Jacob’s operon model was the first to mention that gene expression could be regulated4. Building on this basic foundation, recombinant DNA techniques were developed; a 1973 paper by Cohen et al. showed that bacterial plasmids could be cut, reassembled with foreign DNA, and transformed back into cells5. This is the basis for modern synthetic biology. By 1978, humans were engineering E. coli bacteria to produce synthetic human insulin at scale, using the bacteria as protein factories6.
Another milestone in the development of synthetic biology was the synthesis of a complete bacterial genome. Gibson et al. (2010) reported the creation of a bacterial cell controlled by an artificially synthesized genome, serving as a demonstration that entire genomes could be designed and assembled in the laboratory using artificial means7. This work significantly advanced synthetic biology from manipulating individual genes to engineering whole genomes.
By the turn of the millennium, synthetic biology advanced from relatively-simple gene cloning to the designing of entire genetic circuits using fundamental genetic components. In 2000, Gardner et al. built the first “toggle switch” in E. coli, and Elowitz & Leibler constructed a synthetic oscillator, showing that cells could be reprogrammed to toggle between different states or express different genes8,9. These studies introduced the concept of “parts” that can be utilized to build a genetic circuit, leading to the creation of the BioBrick assembly method and competitions like iGEM, spurring innovation in the field of synthetic biology. The conceptual framework of synthetic biology as an engineering discipline was further articulated by Endy (2005), who proposed that biological systems could be designed using standardized components, abstraction hierarchies, and modular design principles. This perspective helped establish synthetic biology as an engineering-oriented field rather than solely an extension of molecular biology10.
Over the past decade, the field has developed into a highly-automated and engineering-oriented field. With increasingly automated assembly methods and computer software, Design-Build-Test-Learn cycles are compressed significantly; from numerous months to just a few weeks. The introduction of CRISPR-Cas9 in 2012 presented a method capable of editing genes extremely precisely, while displaying high versatility11. This allows for sophisticated genome modifications across organisms; in addition, CRISPR-Cas9 significantly lowered the cost of genetic modification, making the technology available to a wide range of individuals. Today, researchers enjoy an expansive library of standardized genetic “parts”, using them to assemble genetic circuits that serve all sorts of functions. As of the status quo, synthetic biology assists in the development of novel pharmaceuticals and engineering crops with greater resilience and/or increased nutritional value. More recently, genome editing technologies have evolved beyond the original CRISPR-Cas9 system and facilitated the creation of new engineering mechanisms. For example, Anzalone et al. (2019) introduced prime editing, which is a technique that allows for precise insertion, deletion, and base substitution to occur without double-strand DNA breaks12. This development serves as testament to the development of genome editing tools with improved precision and reduced off-target risks. However, most current clinical applications still remain limited to early-stage trials or had tests conducted under extremely strict laboratory experiments, implicating that these examples should be interpreted as proof-of-concept demonstrations rather than solid evidence of the technology’s clinical maturity and applicability in real-world scenarios.
Applications of Synthetic Biology
One of the most promising applications of synthetic biology is in the field of medicinal research and development. Developments of genetic therapies for long-standing genetic diseases include those for cystic fibrosis and sickle cell disease (SCD). Cystic fibrosis, an autosomal monogenic disease, is caused by a CFTR mutation, resulting in insufficient production of the CFTR protein. Experimental gene-editing approaches using CRISPR-Cas9 have been explored for correcting CFTR mutations in cystic fibrosis models13. Furthermore, the monogenic autosomal blood disorder SCD results from a single mutation of the β-globin gene, causing an alteration of the sixth residue’s glutamic acid to a valine14. The recently FDA-approved gene therapy Casgevy is based on CRISPR-Cas9, and it increases production of fetal hemoglobin, of which increased amounts in the bloodstream could prevent the sickling of red blood cells15.
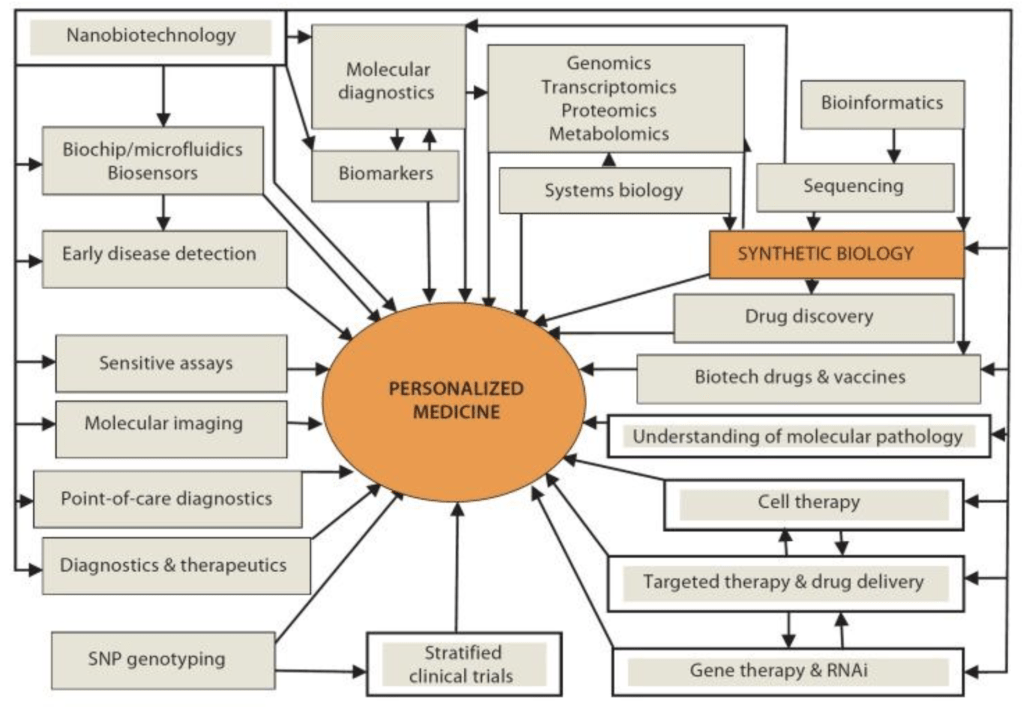
In addition, synthetic biology can offer patients personalized treatments and medicine, rather than the regular one-size-fits-all pill. For instance, tools developed for synthetic biology are versatile and can lead to a more productive research and development cycle for new pharmaceuticals; furthermore, its vast databases can assist in the construction of microbes with new metabolic pathways, enabling better quantitative control over development and engineering of new drugs16. Figure 1 shows the role of synthetic biology in developing personalized medicine, as well as its connection to other fields and research and development strategies.
Furthermore, the utilization of synthetic biology in the field of agriculture also has the potential to yield positive results that could address the issues of food insecurity, malnutrition, and malnourishment, and increase the productivity and sustainability of crop production. In particular, Bt corn and Golden Rice are widely cited examples of the application of genetic engineering in agriculture and have been proposed as strategies to improve crop productivity and nutritional outcomes in certain regions.
Bt maize is genetically engineered to express insecticidal proteins derived from the bacterium Bacillus thuringiensis, providing them with resistance to certain insect pests17. Its primary purpose was to aim for immunity from insect pests. The crop prevention is crucial for regions that suffer from unpredictable pest infestation; Bt corn can ensure yield stability and provide support to developing nations that have maize as a staple food. Additionally, the usage of pesticides recommended for European corn borer (ECB) control dropped from coverage of 6 million acres to 4 million in 199918. Such results display the effectiveness of Bt corn in reducing pesticide use while maintaining the equivalent agricultural output. The reduced pesticide application also benefits the health and safety of farm workers, who are often exposed to hazardous chemicals during traditional pest control methods. Golden Rice, a genetically modified version of rice bioengineered to contain β-carotene, aimed to address vitamin A deficiency in areas where rice is a staple food. The genetic modification was made via the addition of two genes, phytoene synthase (Zmpsy1) and carotene desaturase (crtI) into Kaybonnet, an American rice variety. Introgression of event GR2E in the bio-engineered Kaybonnet into widely-grown rice varieties in the affected regions was successful, without the occurrence of any abnormal phenotypes during the process19. An estimated 250,000 to 500,000 children with Vitamin A Deficiency (VAD) become blind every year; in addition, half of the children die within 12 months of suffering from VAD-induced blindness20. Golden Rice has been proposed as a solution to address the vitamin A deficiency problem in regions where rice is the primary source of food, and it has been continuing its fight with VAD in less economically developed and developing countries ever since. Figure 2 illustrates the biosynthetic pathway used in golden rice to achieve the production of β-carotene.
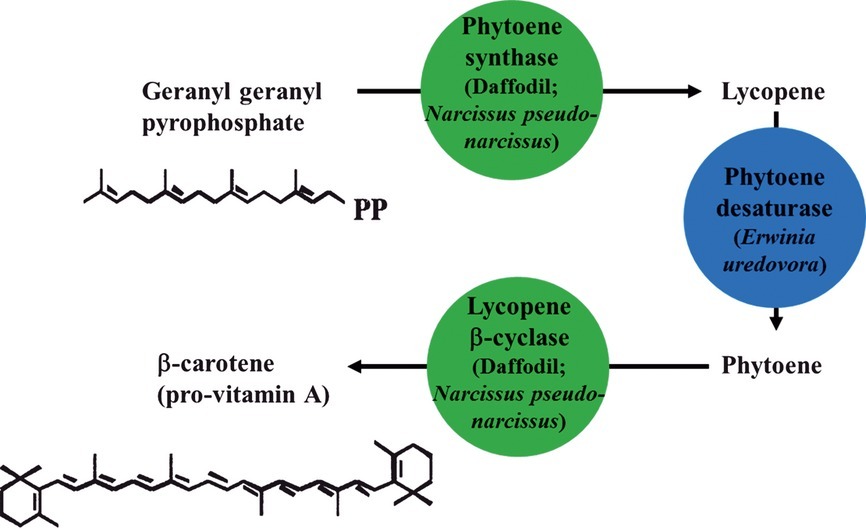
It should be noted that many of these studies are conducted in specific crop systems or regional agricultural conditions, and therefore the results should not be interpreted as universally representative of all farming contexts.
However, advances in synthetic biology regarding medicine and agriculture also raise considerable ethical issues and concerns. Among these, concerns on modification of the human germline are some of the most debated. Synthetic biology allows for more convenient modification of organisms, but misuse of the associated technology can be critically harmful for humans or even the entire ecosystem. For instance, once introduced, human germline modification is heritable and also irreversible; this technology raises ethical dilemmas such as consent of future generations and its potential to widen societal gaps22. Moreover, in the field of agriculture, the release of genetically modified crops may disrupt local ecosystems by introducing new strands that intermix with local or native ones, leading to ecological imbalance. Such gene flow, including the risk of horizontal gene transfer from GM plants, makes the implementation of strict regulations necessary to prevent unintended consequences23. Such challenges demonstrate the need for international regulatory guidelines to be implemented to ensure that the potential and power of synthetic biology are harnessed responsibly.
Apart from concerns regarding gene editing and gene flow, the field of synthetic biology is facing increasing scrutiny from broader governance issues. Such issues include: the need for effective clinical supervision in emerging gene editing therapies; biosafety regulations for engineered organisms used in agriculture and environmental applications; the distribution of benefits from technological innovations to the public; and the management of dual-use risks (i.e., the situation where technologies originally intended for beneficial purposes may be misused). Considering these governance challenges helps to place the technological developments discussed in this article within a broader social and regulatory context.
Research Gap
Synthetic biology, as of the status quo, is a novel field, meaning that it is still in a relatively early and exploratory stage of development, with many core tools and methodologies still being developed. While the potential of synthetic biology is widely recognized, with valuable tools such as CRISPR-Cas9 and programmable gene circuits being significant examples24, widespread applications are rare. As Cameron et al. (2014) observed, much of the current research in synthetic biology is aimed at building the foundational tools and conducting proof-of-concept experiments, rather than fully-developed public commercial applications25.
In reviewing recent literature, it becomes clear that most studies investigate synthetic biology in distinct domains, either focusing on its applications in medicine or in agriculture. While both of these fields have promising procedures, including gene-editing therapies for hereditary disorders and genetically engineered crops that have increased yield, they are regularly treated as two separate areas of innovation. Thus, this study will explore both application fields in parallel, analyzing developments in medicine and agriculture side by side, with the aim of providing a more holistic analysis of how synthetic biology is currently being applied and where it will head in the future.
Methodology
This chapter focuses on the method in which academic literature was surveyed to investigate synthetic biology’s applications in medicine and agriculture.
Literature Collection
Academic articles were gathered from major scholarly databases (such as PubMed and Google Scholar) to ensure comprehensive coverage of peer-reviewed work. Search terms used include “synthetic biology” AND “medicine”, “synthetic biology” AND “agriculture”, “CRISPR”, “metabolic engineering”, and “agricultural biotechnology.” Although the main focus of this review was on peer-reviewed academic literature, when reporting global statistics or policy background, a few grey literature sources (such as regulatory agency reports or statistical databases) were also referred to. These materials are only used to provide descriptive background information and are not regarded as original scientific evidence when evaluating the effectiveness of synthetic biology technologies and synthesizing the analysis. Where relevant, the level of evidence (e.g., preclinical experiment, field trial, early clinical study, or animal experiment) was noted to provide context for interpreting the maturity of the findings.
Article Screening
Regarding the results and discussion sections, the search scope was limited to English peer-reviewed articles published between January 2010 and October 2025. Some early landmark studies were also cited, and the background section described the historical development of the field.
The titles and abstracts of results from this process were noted for initial screening. A screening process was applied. First, titles and abstracts were reviewed to exclude irrelevant papers; for instance, ones without examples of concrete application were left out. Subsequently, full texts of the remaining articles were reviewed to confirm that each had potential case-study or experimental applications in medicine or agriculture. Articles that are non-data opinion pieces were also excluded.
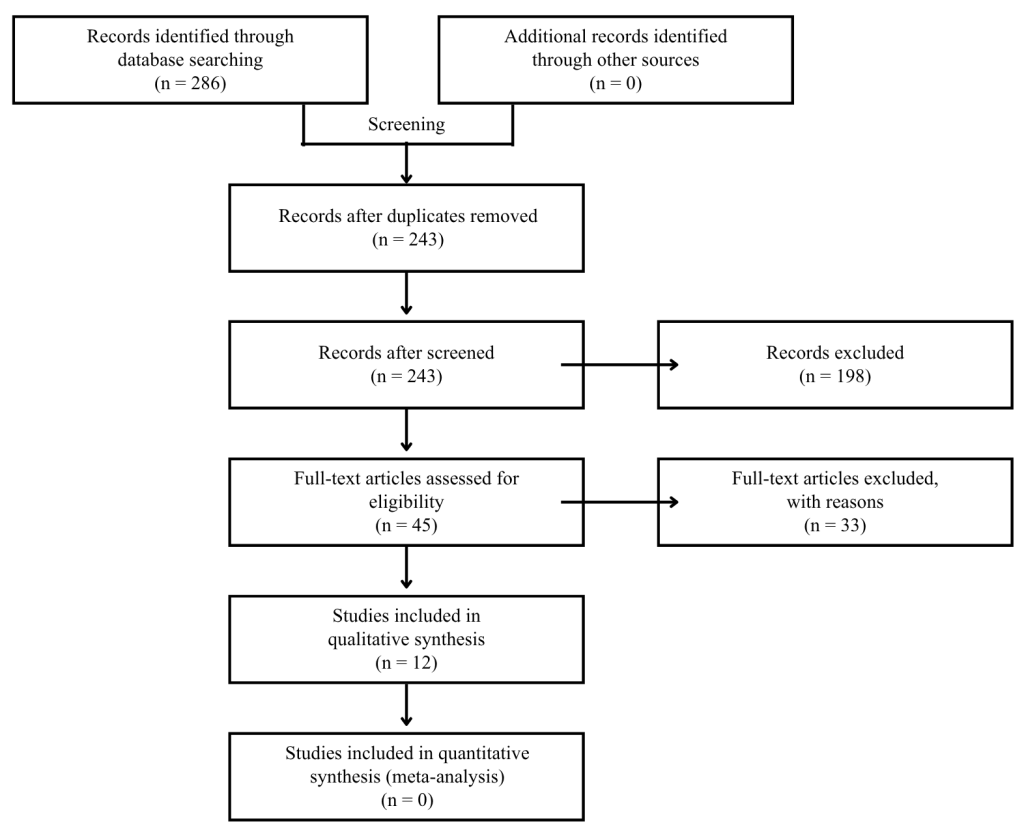
The database search initially identified 286 records. After removing duplicates, there were still 243 records available for title and abstract screening. Among them, 198 articles were excluded because they did not meet the inclusion criteria. Subsequently, the full texts of 45 articles were evaluated to determine if they met the inclusion criteria, and finally 12 studies were included in the final review (Figure 3).
To ensure methodological rigor, all included studies underwent a formal quality assessment based on adapted checklists (e.g., CASP, Joanna Briggs Institute), considering: clarity of objectives, suitability of methods, data reliability, peer-review status, and transparency of reporting. Each study was scored (1 = low, 2 = medium, 3 = high), and those below the threshold were excluded, ensuring that the synthesized evidence is reliable.
Data Extraction/Compilation
For each included article, key information from the main text was extracted and compiled into a structured document. The goal of this process was to organize their findings in a way that directly supports the objective. The extracted information was then sorted and categorized, according to its function, into three main categories: pharmaceutical applications, agricultural applications, and cross-domain applications.
Empirical results, case studies, and past analyses from existing academic studies were identified as material relevant to the Results & Discussion chapter. Conceptual and background information, including definitions, historical context, and overviews of major technologies, were categorized as material for the Literature Review.
The articles for the Results and Discussion section were further classified and placed in three main dimensions: A: Strengths, B: Limitations, and C: Potential. Classifying each study according to these dimensions allowed the research findings to be compared across medicine and agriculture, establishing an evidence-based foundation for the subsequent evaluation and discussion of synthetic biology.
This process ensures reliability and allows the literature selection to be replicated by future researchers. Since this study evaluates the development, applications, and future prospects of an emerging field, the literature-based approach taken in this study is methodologically appropriate, as it allows for broad, cross-disciplinary comparisons.
Results & Discussion
| Study | Study type | Population / model | Intervention | Outcomes measured | Main findings |
| Frangoul et al., 2021 | Early clinical trial | Patients with β-thalassemia and sickle cell disease | CRISPR-Cas9 editing of BCL11A enhancer | Fetal hemoglobin levels | Editing increased fetal hemoglobin; demonstrates feasibility of regulatory-level gene editing in humans |
| Stadtmauer et al., 2020 | Human clinical trial | Cancer patients | CRISPR-edited T cells with PD-1 knockout and synthetic TCR insertion | Engraftment and safety | Engineered T cells showed durable engraftment and multiplex editing feasibility |
| Sterner & Sterner, 2021 | Review of clinical therapies | Cancer patients | CAR-T engineered immune cells | Therapeutic response | CAR-T therapy shows strong efficacy in hematological malignancies |
| Gillmore et al., 2021 | Clinical trial | Patients with transthyretin amyloidosis | In vivo CRISPR therapy (NTLA-2001) | Serum TTR level | TTR protein decreased by up to 87%, demonstrating potential of in vivo editing |
| Böck et al., 2022 | Animal experiment | Mouse model of phenylketonuria | Prime editing in liver cells | Editing efficiency and phenylalanine levels | Editing efficiency up to 58%; metabolic correction observed |
| Qi et al., 2022 | Clinical study | Patients with gastrointestinal cancer | Claudin18.2-targeted CAR-T therapy | Response rate and safety | Response rate 48.6%, demonstrating progress of CAR-T in solid tumors |
| Study | Study type | Organism / system | Intervention | Outcomes measured | Main findings |
| Wei et al., 2022 | Field trial | Rice | Overexpression of transcription factor OsDREB1C | Yield, nitrogen use efficiency | Yield increased 41–68% under field conditions |
| Martinez-Feria et al., 2024 | Field and laboratory experiments | Root-associated bacteria | Gene-edited nitrogen-fixing microbes | Nitrogen fixation and ammonium excretion | Engineered strains enhanced biological nitrogen fixation |
| Kabir et al., 2024 | Review | Agricultural microbiomes | Microbiome engineering approaches | Ecological stability and scalability | Real-world applications limited by complex host–microbiome interactions |
| Toju et al., 2018 | Review | Agroecosystem microbiomes | Microbial community management | Stability of microbial communities | Maintaining engineered microbiomes remains difficult |
| Wen et al., 2021 | Field trials | Corn | Synthetic-biology-optimized diazotroph | Nitrogen fixation and crop yield | Nitrogen fixation increased up to 122-fold under experimental conditions |
| Cook et al., 2025 | Field experiment | Wheat | Transfer of CNGC15 gene affecting symbiosis | Nutrient uptake and AM colonization | Improved nutrient acquisition and symbiotic efficiency |
Current State (Strengths)
Medicine
Across surveyed literature, current strengths in pharmaceutical synthetic biology are centered around (i) regulatory-level genome editing and (ii) increasingly programmable cell-based therapies. A prevalent shift that can be identified is that many approaches are focusing less on rewriting genetic sequences and more on reprogramming gene regulation. Changing genetic regulatory control can reduce the reliance of current therapies on precisely correcting every disease-causing mutation.
Frangoul et al. (2021) provided preliminary evidence from an early-stage human clinical trial, which aimed to study the use of CRISPR-Cas9 technology for gene editing in patients with beta-thalassemia and sickle cell disease who rely on blood transfusions. The study targeted the erythrocyte-specific enhancer of the BCL11A gene, which inhibits the expression of fetal hemoglobin. After editing this enhancer, the level of fetal hemoglobin in the treated patients’ bodies was observed to have increased26.
As clinical evidence, this study demonstrates that gene editing at the regulatory level in human therapeutic contexts is feasible. However, as an early clinical study with a limited sample size, its results mainly support the feasibility of CRISPR-based treatment strategies and is a proof-of-concept rather than a total proof for its general clinical effectiveness in treating single-gene diseases.
Beyond genome editing, the field of synthetic biology is also advancing toward programmable therapeutic cells, where engineered cells can be designed to perform specific therapeutic functions. Human clinical trials confirm this advancement; for instance, Stadtmauer et al. (2020) provides such a trial for programmable cell-based therapies. In small early-phase clinical study involving three patients with refractory cancer, the researchers tested the safety of CRISPR-Cas9-based T cell therapy in three patients with refractory cancer. This process involved the deletion of endogenous T cell receptor (TCR) genes to reduce mispairing and enhance the expression of a synthetic TCR transgene as well as the knocking out of the gene coding for PD-1, which improve anti-tumor immunity. Following their adoptive transfers into patients, they exhibited durable engraftment for up to 9 months; chromosomal translocations also decreased over time27. This study is impactful, as it demonstrates that CRISPR can be used to “program” therapeutic immune cells, which coordinate multiple edits in human cells; as such, it supports the feasibility of multiplex gene editing for cancer immunotherapy.
CAR-T represents the most clinically established “engineered cell” therapy enabled by synthetic biology. In CAR-T therapy, T cells are reprogrammed to express a chimeric antigen receptor (CAR) that couples an extracellular antigen recognition domain with an intracellular signal transduction pathway and a co-stimulatory domain. Importantly, CARs are MHC-independent, enabling T-cell activation even when cancerous cells reduce MHC molecule production to evade conventional T-cell receptor recognition. Clinically, CAR-T therapy’s impact is the most noticeable in the treatment of hematological malignancies, such as the utilization of second-generation CAR-T cells that have produced strong therapeutic responses28.
Agriculture
The survey of scientific literature indicates that the strengths of synthetic biology in agriculture include translation into field-level gains in productivity and resource-use efficiency. One approach is targeting transcriptional regulators that coordinate large gene networks, allowing one change to contribute to downstream and higher-impact phenotypic changes.
Its impact upon the field is demonstrated through the genetic regulatory engineering of rice to increase crop yield while maintaining agronomic performance. In a screening of photosynthesis-related transcription factors, Wei et al. (2022) identified OsDREB1C, whose expression drives transcriptional programs like photosynthetic capacity, nitrogen use efficiency, and flowering time. In controlled field trials conducted in experimental plots, the study found that OsDREB1C-overexpressing rice increases yield by 41.3%–68.3% under the specific experimental conditions reported in the study, alongside shorter growth duration and better nitrogen use efficiency29. These results indicate that regulatory-level engineering can translate into practical gains in agricultural productivity and resource-use efficiency. Because these results were obtained under field trial conditions rather than laboratory experiments, they provide stronger support for the potential agricultural productivity benefits of regulatory-level genetic engineering.
Beyond internal plant regulatory rewiring, agricultural synthetic biology is also participating in the engineering of root-associated microbes to improve nutrient supply. Martinez-Feria et al. (2024) reports gene-edited strains of Klebsiella variicola and Kosakonia sacchari that have been optimized for root-associated biological nitrogen fixation (BNF) and ammonium excretion even under nitrogen-rich rhizosphere conditions. Evidence from laboratory experiments, greenhouse plant tests, and limited field trials indicate that the study’s genetic remodeling strategy successfully derepresses BNF activity and increases ammonium excretion relative to wild-type strains30. These results are impactful because they show that engineered microbiome intervention can lead to noticeable nitrogen contributions under real agricultural conditions; such modified environments can support crop nitrogen status and reduce the risk of losing crop yield when synthetic fertilizer inputs are reduced.
However, despite these advances, the same scientific literature also emphasizes that the clinical and agricultural application of synthetic biology may face technical and ethical barriers. The subsequent section evaluates the current challenges reported across studies and reviews.
Limitations (Challenges)
Medicine
Although regulatory-level genome editing introduces powerful therapeutic capabilities, its precision also introduces technical challenges. Guo et al. (2023) outlines the risk of CRISPR-Cas9 genetic modification technology inducing off-target effects. Despite the many benefits it has brought and its success in both in vitro and clinical experiments, the system is still prone to potential off-target effects that may cause harmful biological consequences to patients31. More importantly, the review emphasizes that a major bottleneck is both in reducing off-target activity and reliably detecting such unintended edits across cell types; consequently, off-target risk remains a major roadblock to CRISPR-Cas9 gene editing’s application in clinical therapies.
In addition, the process of translation itself is also limited by logistics and the engineering of cells at a consistent pharmaceutical-grade quality. Kohn, Chen & Spencer (2023) notes that despite the efficacy of synthetic biology therapies in vivo, practical barriers block their translation into real-world, widely-available therapies. The article notes ongoing challenges, including the high one-time costs of production, the need for pharmaceutical consistency, and potential genotoxicity risks (for example, unintended genomic disruption as a result of viral vector integration) as examples of such barriers32. Problems such as these increase regulatory burden and necessitate the need for long-term monitoring to ensure safety.
Adoptive cell therapies developed by synthetic biology, namely CAR-T therapy, do have their limitations. For instance, according to the same article Sterner & Sterner (2021), CAR-T therapies exhibit limited efficacy in reducing solid tumors; while it has been successful in treating hematological malignancies (such as leukemia and lymphomas), solid tumors inhibit CAR-T activity. Solid tumors can produce immunosuppressive microenvironments that make them more robust to external regulation, and their nature reduces CAR-T cells’ access to the tumor site. The review also marks antigen escape as a major drawback to CAR-T therapy. Evolutionary pressure exerted by CAR-T cells encourages tumor cells to downregulate or eliminate antigen expression, leading to potential recurrence and contributing to relapses in both hematologic and solid tumors28.
Agriculture
While synthetic biology approaches show strong productivity potential in controlled experiments, translating these strategies to applications in complex real-world agricultural ecosystems reveal significant challenges. For instance, Kabir et al. (2024) notes that progress is impeded by the complex nature of microbiome-host interactions in real environmental settings. Current technological tools and understanding of ecology are still insufficient for determining the real-world impact of synthetically engineered microbial interventions33. Despite these approaches showing great promise in controlled lab environments, translation to scalable, real-world agriculture requires them to remain stable across diverse climates and environments. Toju et al. (2018) also notes that a major limitation to microbiome-based agricultural synthetic biology is the difficulty of maintenance. The proposed solution must reliably manage complex, species-rich microbial communities; community dynamics can be difficult to track, especially in open agroecosystems34. While it recognizes the necessity of maximizing microbial function in the face of climate change, it emphasizes the importance of management and improving controllability, in the absence of which microbial solutions may fail to perform across different environments.
Potential (Opportunities)
Medicine
Because these technical and translational limitations remain unresolved, current research is increasingly focused on developing safer and more controllable therapeutic systems. Looking forward, synthetic biology can be a significant catalyst for pharmaceutical development once the safety and controllability of therapies improve and the range of treatable diseases increases.
The potential possibilities of future therapeutic applications are based on evidence from two different experimental levels. Firstly, human clinical research evidence was provided by Gillmore et al. (2021), who conducted tests on patients with transthyretin amyloidosis using a CRISPR-based gene editing therapy (NTLA-2001). The research report stated that the serum TTR level decreased by up to 87% after treatment, indicating that in the early trials, in vivo gene editing was capable of being safely implemented in humans35. Secondly, animal experiments further explored the new editing technology. Böck et al. (2022) demonstrated the feasibility of in situ editing technology in a phenylketonuria mouse model. A 58% editing efficiency was observed in mouse liver cells. Although these findings provided important mechanistic insights, the animal experiments mainly showed its feasibility in biological organisms in a broader sense rather than its clinical applicability in humans. In mice, engineered prime editors achieved an editing efficiency of up to 58%; more importantly, in a phenylketonuria (PKU) mouse model, the study achieved a correction efficiency of up to 17.4%36. The subsequent reduction in blood phenylalanine suggests that this procedure can be developed into a curative therapy for PKU and other genetic liver diseases. Taken together, such studies provide proof-of-concept demonstrations derived from synthetic biology, supporting the idea that next-generation therapies can extend ex vivo lab advances into in vivo interventions. This outlines the trajectory of synthetic biology therapies toward creating cures for monogenic disorders, particularly those in the liver.
In CAR-T therapy, its future opportunity is essentially defined by its transition from successfully tackling hematologic malignancies to solid tumors, where efficacy has been historically limited (as discussed in the 4.2.1 Limitations section). However, trials have shown that improved engineering techniques and better targeting can lead to meaningful responses in solid tumors. For instance, Qi et al. (2022) reported positive results in a Phase 1 clinical trial using Claudin18.2-redirected CAR-T cells in gastrointestinal cancers. Despite grade 1–2 cytokine release syndrome (CRS) occurring in 94.6% of participating patients, the treatment did not lead to any neurotoxicities, treatment-related deaths, or grade ≥3 CRS. Furthermore, the overall response rate and disease control rate reached 48.6% and 73.0%, respectively37. This study provides concrete evidence that CAR-T therapy is being developed to overcome the barriers of solid tumors, indicating an expansion from current focuses on its utilization in hematologic malignancies.
Agriculture
To address these ecological and scalability challenges, current research increasingly focuses on developing more stable microbial systems and plant–microbe symbioses. The application of artificially engineered, root-associated nitrogen-fixing microbes in the agricultural field could partially replace the need for synthetic nitrogen fertilizer while maintaining or even improving yield. Wen et al. (2021) outlines the commercialization of a synthetic-biology-optimized diazotroph for corn in fertilized fields. The paper notes that the product increased nitrogen fixation 122-fold in nitrogen-rich conditions when conducting experimental trials, and produced higher yields across multi-year U.S. Corn Belt field trials38.
Beyond microbial products oriented around biological nitrogen fixation (BNF), synthetic biology can also help in reducing fertilizer dependence by developing plant-microbe symbioses that improve nutrient uptake. Cook et al. (2025) reports that cyclic nucleotide-gated channel 15 (CNGC15) enhances endosymbiosis and plant nutrient acquisition. Importantly, when this trait was transferred to wheat, the resulting field-grown wheat exhibited stronger nutrient acquisition abilities and increased arbuscular mycorrhizal (AM) colonization39. This study demonstrates that engineered symbiosis frameworks are a viable way of sustaining plant growth and can be considered as an alternative to inorganic fertilizers.
Cross-domain insights from synthetic biology applications
Scientific reviews often assess the applications of synthetic biology in medicine and agriculture separately and treat them as two distinct branches, creating a research gap in this area. However, numerous commonalities can be identified between the two fields.
Both fields use similar fundamental tools to facilitate and refine technological advancements. Examples of such tools include CRISPR and artificial engineering of genetic circuits. Furthermore, synthetic biology solutions all involve the utilization of engineered microorganisms or cells, regardless of its use; they can be involved in the production of therapeutic molecules or used to enhance the nutritional content of crops. This commonality between the two fields shows that the same basic rules and principles can be applied to different biological domains regardless of the research subject.
Despite this, the transformation pathway from highly theoretical knowledge into real-world applications is different in both fields. On one hand, medical applications of theory typically require a larger number of clinical trials that are contained within stages; in addition, they usually need to obtain regulatory approval before public release into the market. On the other hand, agricultural applications often need to consider factors pertaining to society, the overall economy, and environmental policies. Understanding these commonalities between the two fields and where they diverge may help guide interdisciplinary research in synthetic biology in the future.
Synthetic biology principles have contributed to technological advancements in multiple fields; however, its real-world applications are still affected by socioeconomic factors. Among these factors are affordability, accessibility, and medical equity. As of the status quo, many advanced therapies developed with synthetic biology are expensive and require specialized clinical infrastructure, hence limiting their public accessibility. This problem raises concern over who actually benefits from the aforementioned medical innovations created by synthetic biology and whether they will widen the access disparity between the rich and the poor to advanced treatments. Similar problems emerge when considering the agricultural field. Although technologies such as genetically engineered crops and microbiome-modifying fertilizers are developed to boost agricultural productivity, their implementation may be impeded by numerous factors; among these, regulatory frameworks, public acceptance (or the lack thereof), intellectual property structures, and the economic affluence of farmers are major factors that affect their implementation.
In short, in both medicine and agriculture, the impact that synthetic biology brings depends on the technology’s technical feasibility, the socioeconomic conditions of consumers, and current regulations regarding research and release to the consumer market. The ability to program biological systems facilitates development of new technologies regardless of the subject; however, its applications are constrained by technology, regulations, and ecological concerns. Resolving issues and approving targeted regulations regarding these impediments may motivate the subsequent generation of novel research directions in synthetic biology, ultimately creating a positive feedback loop between innovation and regulatory adaptation.
Conclusion
In aggregate, this study evaluated the historical development, current status, and future prospects of synthetic biology in medicine and agriculture, in an attempt to address the research gap between review papers that treat the two as distinct fields. From surveying peer-reviewed literature from 2010 – 2025 according to the methodology and classifying those sources into strengths, limitations, and opportunities, it can be noticed that synthetic biology has recently progressed from developing individual tools to engineering increasingly programmable systems that are capable of wider applications.
In medicine, current strengths of synthetic biology include regulatory-level genome editing and engineered cell therapies. Clinical studies have shown that treating diseases does not require precise correction of every mutation; instead, editing gene regulation can treat disease mechanisms. Adoptive cell therapies, including CAR-T therapy, have been shown to be capable of treating blood cancers and potentially developing complete cures for them. However, scientific literature also notes the limitations of these methods. Specifically, CRISPR-based therapies are prone to off-target effects and regulatory blindspots; in addition, the translation of in vitro experiments into in vivo applications requires both consistency in manufacturing and significant capital. Furthermore, CAR-T cells may cause antigen escapes and have been shown to be less effective toward solid tumors during treatment.
In agriculture, real-world applications of synthetic biology have already shown that modifications of regulatory mechanisms in plants can increase crop yield and resource-use efficiency. The development of engineered microbes capable of biological nitrogen fixation (BNF) have contributed to major rises in plant nitrogen availability. However, despite these successes, synthetic biology’s agricultural translation into real-world solutions remains limited by the complex and unstable nature of microbiomes. Reviews of scientific literature reveal how the complex interactive webs between organisms in ecosystems are difficult to regulate and may be of impediment to application of synthetic tools due to the disruption of the interconnected webs, hence triggering unintended consequences.
Considering future prospects, synthetic biology’s developments in medicine and agriculture may develop novel therapies after current regulatory and safety problems have been resolved. Once such problems are resolved, these applications can contribute to real advancement in society. In medicine, in vivo trials and prime editing demonstrate synthetic biology’s shift from facilitating highly-theoretical, proof-of-concept experiments to creating actual curative therapies. In particular, such in vivo experiments and proof-of-concepts indicate the future development of therapies for monogenic disorders, particularly those in the liver. In agriculture, products involving engineered microbes capable of biological nitrogen fixation (BNF) and engineered symbioses suggest dependence on synthetic, inorganic fertilizers can be reduced while ensuring yield. Such methods help maintain a healthier natural ecosystem. Overall, the analysis of scientific literature has revealed that synthetic biology is shifting from creating fundamental tools to building entire application-based systems; however, dilemmas on safety and controllability have to be overcome in order to achieve this goal.
References
- Lee, C.E., Singleton, K. s, Wallin, M. and Faundez, V. (2020). Rare Genetic Diseases: Nature’s Experiments on Human Development. iScience, [online] 23(5), p.101123. doi:https://doi.org/10.1016/j.isci.2020.101123. [↩]
- Fao.org. (2026). FAO Knowledge Repository. [online] Available at: https://openknowledge.fao.org/items/445c9d27-b396-4126-96c9-50b335364d01. [↩]
- National Human Genome Research Institute. (2019, August 14). Synthetic Biology. Genome.gov; National Human Genome Research Institute. https://www.genome.gov/about-genomics/policy-issues/Synthetic-Biology. [↩]
- Jacob, F., & Monod, J. (1961). Genetic regulatory mechanisms in the synthesis of proteins. Journal of Molecular Biology, 3(3), 318–356. https://doi.org/10.1016/s0022-2836(61)80072-7. [↩]
- Cohen, S. N., Chang, A. C. Y., Boyer, H. W., & Helling, R. B. (1973). Construction of Biologically Functional Bacterial Plasmids In Vitro. Proceedings of the National Academy of Sciences, 70(11), 3240–3244. https://doi.org/10.1073/pnas.70.11.3240. [↩]
- Quianzon, C. C., & Cheikh, I. (2012). History of insulin. Journal of Community Hospital Internal Medicine Perspectives, 2(2), 18701. https://doi.org/10.3402/jchimp.v2i2.18701. [↩]
- Gibson, D.G., Glass, J.I., Lartigue, C., et al. (2010).Creation of a bacterial cell controlled by a chemically synthesized genome. Science, 329(5987), pp.52–56. Available from: https://doi.org/10.1126/science.1190719. [↩]
- Gardner, T. S., Cantor, C. R., & Collins, J. J. (2000). Construction of a genetic toggle switch in Escherichia coli. Nature, 403(6767), 339–342. https://doi.org/10.1038/35002131. [↩]
- Elowitz, M. B., & Leibler, S. (2000). A synthetic oscillatory network of transcriptional regulators. Nature, 403(6767), 335–338. https://doi.org/10.1038/35002125. [↩]
- Endy, D. (2005). Foundations for engineering biology. Nature, 438(7067), 449–453. https://doi.org/10.1038/nature04342. [↩]
- Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). A Programmable Dual-RNA-Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science, 337(6096), 816–821. https://doi.org/10.1126/science.1225829. [↩]
- Anzalone, A. V., Randolph, P. B., Davis, J. R., Sousa, A. A., Koblan, L. W., Levy, J. M., Chen, P. J., Wilson, C., Newby, G. A., Raguram, A., & Liu, D. R. (2019). Search-and-replace Genome Editing without double-strand Breaks or Donor DNA. Nature, 576(576). https://doi.org/10.1038/s41586-019-1711-4. [↩]
- Schwank, G., Koo, B.-K., Sasselli, V., Dekkers, Johanna F., Heo, I., Demircan, T., Sasaki, N., Boymans, S., Cuppen, E., van der Ent, Cornelis K., Nieuwenhuis, Edward E. S., Beekman, Jeffrey M., & Clevers, H. (2013). Functional Repair of CFTR by CRISPR/Cas9 in Intestinal Stem Cell Organoids of Cystic Fibrosis Patients. Cell Stem Cell, 13(6), 653–658. https://doi.org/10.1016/j.stem.2013.11.002. [↩]
- Park, S.H. and Bao, G. (2021). CRISPR/Cas9 gene editing for curing sickle cell disease. Transfusion and Apheresis Science, [online] 60(1), p.103060. doi:https://doi.org/10.1016/j.transci.2021.103060. [↩]
- FDA (2023). FDA Approves First Gene Therapies to Treat Patients with Sickle Cell Disease. [online] FDA. Available at: https://www.fda.gov/news-events/press-announcements/fda-approves-first-gene-therapies-treat-patients-sickle-cell-disease. [↩]
- Jain, K.K. (2013). Synthetic Biology and Personalized Medicine. Medical Principles and Practice, [online] 22(3), pp.209–219. doi:https://doi.org/10.1159/000341794. [↩] [↩]
- Shelton, A. M., Zhao, J. Z., & Roush, R. T. (2002). Economic, ecological, food safety, and social consequences of the deployment of Bt transgenic plants. Annual review of entomology, 47(1), 845-881. [↩]
- Bt Plant-Pesticides Biopesticides Registration Action Document IIE1 E. Benefits Assessment. (n.d.). Available at: https://archive.epa.gov/scipoly/sap/meetings/web/pdf/brad5_benefits_corn.pdf. [↩]
- Mallikarjuna Swamy, B.P., Marundan, S., Samia, M., Ordonio, R.L., Rebong, D.B., Miranda, R., Alibuyog, A., Rebong, A.T., Tabil, M.A., Suralta, R.R., Alfonso, A.A., Biswas, P.S., Kader, M.A., Reinke, R.F., Boncodin, R. and MacKenzie, D.J. (2021). Development and characterization of GR2E Golden rice introgression lines. Scientific Reports, [online] 11(1), p.2496. doi:https://doi.org/10.1038/s41598-021-82001-0. [↩]
- Xu, Y., Shan, Y., Lin, X., Miao, Q., Lou, L., Wang, Y. and Ye, J. (2021). Global patterns in vision loss burden due to vitamin A deficiency from 1990 to 2017. Public Health Nutrition, 24(17), pp.5786–5794. doi:https://doi.org/10.1017/s1368980021001324. [↩]
- Halford, N.G. (2012). Toward two decades of plant biotechnology: successes, failures, and prospects. Food and Energy Security, 1(1), pp.9–28. doi:https://doi.org/10.1002/fes3.3. [↩]
- Halpern, J., O’Hara, S.E., Doxzen, K.W., Witkowsky, L.B. and Owen, A.L. (2019). Societal and Ethical Impacts of Germline Genome Editing: How Can We Secure Human Rights? The CRISPR Journal, 2(5), pp.293–298. doi:https://doi.org/10.1089/crispr.2019.0042. [↩]
- Philips, J.G., Martin-Avila, E. and Robold, A.V. (2022). Horizontal gene transfer from genetically modified plants – Regulatory considerations. Frontiers in Bioengineering and Biotechnology, 10. doi:https://doi.org/10.3389/fbioe.2022.971402. [↩]
- Barrangou, R. and Doudna, J.A. (2016). Applications of CRISPR technologies in research and beyond. Nature Biotechnology, [online] 34(9), pp.933–941. doi:https://doi.org/10.1038/nbt.3659. [↩]
- Cameron, D.E., Bashor, C.J. and Collins, J.J. (2014). A brief history of synthetic biology. Nature Reviews Microbiology, [online] 12(5), pp.381–390. doi:https://doi.org/10.1038/nrmicro3239. [↩]
- Frangoul, H. (2021). CRISPR-Cas9 Gene Editing for Sickle Cell Disease and β-Thalassemia. New England Journal of Medicine, [online] 384(3). doi:https://doi.org/10.1056/nejmoa2031054. [↩]
- Stadtmauer, E.A. (2020). CRISPR-engineered T cells in patients with refractory cancer. Science, [online] 367(6481). doi:https://doi.org/10.1126/science.aba7365. [↩]
- Sterner, R.C. and Sterner, R.M. (2021). CAR-T Cell therapy: Current Limitations and Potential Strategies. Blood Cancer Journal, [online] 11(4), pp.1–11. Available at: https://www.nature.com/articles/s41408-021-00459-7. [↩] [↩]
- Wei, S., Li, X., Lu, Z., Zhang, H., Ye, X., Zhou, Y., Li, J., Yan, Y., Pei, H., Duan, F., Wang, D., Chen, S., Wang, P., Zhang, C., Shang, L., Zhou, Y., Yan, P., Zhao, M., Huang, J. and Bock, R. (2022). A transcriptional regulator that boosts grain yields and shortens the growth duration of rice. Science, 377(6604). doi:https://doi.org/10.1126/science.abi8455. [↩]
- Martinez-Feria, R., Simmonds, M.B., Bilge Ozaydin, Lewis, S., Schwartz, A., Pluchino, A., McKellar, M., Gottlieb, S.S., Kayatsky, T., Vital, R., Mehlman, S.E., Caron, Z., Colaianni, N.R., Jean-Michel Ané, Maeda, J., Infante, V., Karlsson, B.H., McLimans, C., Vyn, T. and Hanson, B. (2024). Genetic remodeling of soil diazotrophs enables partial replacement of synthetic nitrogen fertilizer with biological nitrogen fixation in maize. Scientific Reports, [online] 14(1). doi:https://doi.org/10.1038/s41598-024-78243-3. [↩]
- Guo, C., Ma, X., Gao, F. and Guo, Y. (2023). Off-target Effects in CRISPR/Cas9 Gene Editing. Frontiers in Bioengineering and Biotechnology, [online] 11(1143157). doi:https://doi.org/10.3389/fbioe.2023.1143157. [↩]
- Kohn, D.B., Chen, Y.Y. and Spencer, M.J. (2023). Successes and Challenges in Clinical Gene Therapy. Gene Therapy, [online] 30, pp.1–9. doi:https://doi.org/10.1038/s41434-023-00390-5. [↩]
- Ahmad Humayan Kabir, Zakaria, M., Ahmed, B. and Mohammad Golam Mostofa (2024). Current, faltering, and future strategies for advancing microbiome-assisted sustainable agriculture and environmental resilience. New crops, pp.100013–100013. doi:https://doi.org/10.1016/j.ncrops.2024.100013. [↩]
- Toju, H., Peay, K.G., Yamamichi, M., Narisawa, K., Hiruma, K., Naito, K., Fukuda, S., Ushio, M., Nakaoka, S., Onoda, Y., Yoshida, K., Schlaeppi, K., Bai, Y., Sugiura, R., Ichihashi, Y., Minamisawa, K. and Kiers, E.T. (2018). Core microbiomes for sustainable agroecosystems. Nature Plants, 4(5), pp.247–257. doi:https://doi.org/10.1038/s41477-018-0139-4. [↩]
- Gillmore, J.D., Gane, E., Taubel, J., Kao, J., Fontana, M., Maitland, M.L., Seitzer, J., O’Connell, D., Walsh, K.R., Wood, K., Phillips, J., Xu, Y., Amaral, A., Boyd, A.P., Cehelsky, J.E., McKee, M.D., Schiermeier, A., Harari, O., Murphy, A. and Kyratsous, C.A. (2021). CRISPR-Cas9 In Vivo Gene Editing for Transthyretin Amyloidosis. New England Journal of Medicine, [online] 385(6). doi:https://doi.org/10.1056/nejmoa2107454. [↩]
- Böck, D., Rothgangl, T., Villiger, L., Schmidheini, L., Matsushita, M., Mathis, N., Ioannidi, E., Rimann, N., Grisch-Chan, H.M., Kreutzer, S., Kontarakis, Z., Kopf, M., Thöny, B. and Schwank, G. (2022). In vivo prime editing of a metabolic liver disease in mice. Science Translational Medicine, [online] 14(636). doi:https://doi.org/10.1126/scitranslmed.abl9238. [↩]
- Qi, C., Gong, J., Li, J., Liu, D., Qin, Y., Ge, S., Zhang, M., Peng, Z., Zhou, J., Cao, Y., Zhang, X., Lu, Z., Lu, M., Yuan, J., Wang, Z., Wang, Y., Peng, X., Gao, H., Liu, Z. and Wang, H. (2022). Claudin18.2-specific CAR T cells in gastrointestinal cancers: phase 1 trial interim results. Nature Medicine, [online] 28(6), pp.1189–1198. doi:https://doi.org/10.1038/s41591-022-01800-8. [↩]
- Wen, A., Havens, K.L., Bloch, S.E., Shah, N., Higgins, D.A., Davis-Richardson, A.G., Sharon, J., Rezaei, F., Mohiti-Asli, M., Johnson, A., Abud, G., Ane, J.-M., Maeda, J., Infante, V., Gottlieb, S.S., Lorigan, J.G., Williams, L., Horton, A., McKellar, M. and Soriano, D. (2021). Enabling Biological Nitrogen Fixation for Cereal Crops in Fertilized Fields. ACS Synthetic Biology, 10(12), pp.3264–3277. doi:https://doi.org/10.1021/acssynbio.1c00049. [↩]
- Cook, N.M., Gobbato, G., Jacott, C.N., Marchal, C., Hsieh, C.Y., Lam, A.H.C., Simmonds, J., del Cerro, P., Gomez, P.N., Rodney, C., Cruz-Mireles, N., Uauy, C., Haerty, W., Lawson, D.M. and Charpentier, M. (2025). Autoactive CNGC15 enhances root endosymbiosis in legume and wheat. Nature. doi:https://doi.org/10.1038/s41586-024-08424-7. [↩]