Abstract
Human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CMs) are a promising tool to model and investigate human heart disease in vitro. To achieve this goal, differentiation protocols must be optimized to produce pure populations of somatic cells, such as cardiomyocytes (CMs). CHIR99021 (CHIR) is a small molecule that inhibits the enzyme glycogen synthase kinase-3 beta (GSK-3β), activating the Wnt/β-catenin pathway and genes involved in cardiac cell growth and differentiation. Therefore, CHIR is commonly used to initiate directed CM differentiation from hiPSCs. However, the optimal concentration of CHIR varies for different cell lines, making maximum differentiation efficiency a significant hurdle. Mutations in Rotatin (RTTN), a gene encoding a centrosomal protein, have been linked to defective heart maturation and cardiovascular disease, but this protein and other centrosomal proteins must be explored further using hiPSC-CMs. Based on conventional lab protocol, we hypothesized that 12 μM CHIR would have the highest differentiation efficiency. In this study, we used immunostaining protocols to stain and image CC2 CMs (wild-type cells) and LVIP CMs (containing a patient-specific compound heterozygous RTTN mutation), allowing protein localization to be observed through imaging and manual counting. This study suggests a concentration of 6 μM CHIR promotes optimal CM differentiation efficiency for both CC2 and LVIP hiPSCs, challenging our hypothesis. The findings of this study are crucial to enhance understanding regarding the optimal differentiation of cardiomyocytes from hiPSCs. Moreover, they constitute a step toward future research into the role of centrosomal proteins in cardiovascular disease.
Keywords: Human induced pluripotent stem cells (hiPSCs), Cardiomyocytes (CMs), CHIR99021 (CHIR), Wnt pathway, Differentiation, Rotatin (RTTN), Centrosome, Cardiovascular disease
Introduction
Cardiovascular disease is the leading cause of death in the US and other developed countries1. One of its most severe consequences is myocardial infarction, which is more commonly known as a heart attack. This leads to the loss of cardiomyocytes (CMs), which are the contractile muscle cells of the heart2,3. Unlike other tissues, adult cardiomyocytes have limited regenerative capacity because the cells are terminally differentiated and do not undergo mitosis. As a result, dead cells are replaced by scar tissue rather than new muscle cells. This leads to impaired cardiac function and, if left untreated, heart failure4,5. Therefore, finding ways to model and research the mechanisms of cardiovascular disease is crucial. One such method is pluripotent stem cells, which are often used to model the heart in vitro and to investigate disease pathology6.
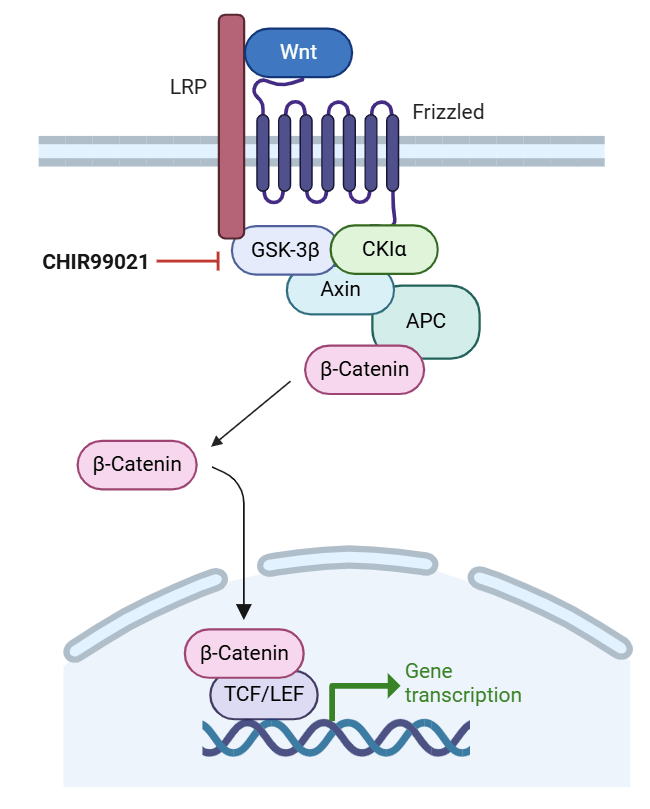
A key compound used in several hiPSC differentiation protocols is CHIR99021 (CHIR), a small molecule inhibitor of glycogen synthase kinase-3β (GSK-3β)7,8. Specifically, CHIR prevents GSK-3β from phosphorylating β-catenin. This stabilizes this molecule and allows it to translocate to the nucleus. Then, it activates Wnt/β-catenin target genes through TCF/LEF transcription factors8 (Figure 1). These Wnt gene targets then affect the expression of pluripotency and developmental factors and are important for cell fate determination and stem cell maintenance9. CHIR has been used to direct human induced pluripotent stem cells (hiPSCs) down the mesoderm lineage, a critical step in cardiomyocyte differentiation10. Its reported optimal concentration varies from as low as 7-8 μM to 12-18 μM, depending on factors such as which cell line is used and cell culture conditions11,12,13. Successful CM differentiation has not been achieved with CHIR concentrations below this threshold. However, the inconsistency between protocols needs to be further refined, as maximum differentiation efficiency is crucial for achieving pure populations of CMs and other cell types.
This study aims to evaluate which concentration of CHIR99021 most effectively promotes cardiomyocyte differentiation in two hiPSC lines: CC2, a wild-type control, and LVIP, a patient-specific line with a compound heterozygous mutation in the Rotatin gene (RTTN). More specifically, this cell line contains a G1321D missense mutation from the father and an in-frame deletion from the mother. One study has found that mutations in RTTN cause infantile dilated cardiomyopathy through impaired CM maturation but the involvement of other centrosomal proteins must be explored14. Through this study, effective differentiation protocols can be developed for future studies aimed at investigating centrosomal protein localization in developing CMs. Based on publicly available protocols, we hypothesized that 12 μM CHIR would result in the most efficient differentiation in both lines15,16,17. However, we found that 6 μM CHIR was by far the most effective concentration of those tested. The results of this study represent steps towards challenging conventional paradigms, allowing for optimized differentiation of hiPSC-CMs. Our findings provide us with a valuable tool for future research aimed at investigating the effects of impaired centrosomal protein function on cardiac development.
In this study, immunostaining protocols were used to stain cardiomyocyte nuclei to quantify differentiation efficiency. Automated image analysis was conducted using the Cytation 10 imaging system as well as ImageJ software to manually count the number of nuclei with cardiac staining. Although this study does investigate differentiation efficiency for two cell lines, different trends could be observed if conducted using other cell lines. Additionally, data analysis could be influenced by bias as the researcher knew which concentration of CHIR each set of images was from. However, the researcher conducting the analysis did not know what to expect, so confirmation bias was limited.
Methods
Stem cell culture
Human iPSCs (hiPSCs) were cultured on six-well plates coated with growth factor-reduced Matrigel (Corning) in an incubator at 37°C, 5% CO2. Stem cells were maintained in mTeSR Plus medium (Stem Cell Technologies) until approximately 70% confluency was reached, at which point cells were split into new wells using ReLeSR passaging reagent (Stem Cell Technologies) with 2 μM ROCK inhibitor (Thiazovivin/TZV; Selleck Chemicals) added to prevent cells from undergoing apoptosis while in suspension. All hiPSC lines were validated for pluripotency approximately every three months using karyotypic analysis.
Both CC2 and LVIP lines were from males and originate from fibroblasts (skin fibroblasts for the CC2 line, and cardiac fibroblasts for the LVIP line). For the CC2 line, reprogramming followed the protocol outlined in this study18. For the LVIP line, reprogramming followed the protocol outlined in this study14.
Cardiomyocyte-directed differentiation
Cardiac differentiation of hiPSCs was performed using the small molecule-based method, where small molecules are added to hiPSCs at specific time points to induce differentiation15,19,20. hiPSCs were grown until approximately 80-90% confluence, a time point designated as Day 0. At this point, the medium was changed to RPMI 1640 (Life Technologies) plus B27 without insulin supplement (Life Technologies), supplemented with 6-12 μM CHIR99021 (Selleck Chemicals) for 24 hours. On day 1, the medium was changed to RPMI 1640 plus B27 without insulin or CHIR. On day 3, the medium was changed to RPMI 1640 plus B27 minus insulin, supplemented with 2 μM Wnt-C59, a small molecule Wnt inhibitor (Sigma). Medium was changed on day 5 with RPMI 1640 plus B27 without insulin. On Day 7, medium was changed to RPMI 1640 plus B27 with insulin and changed with this medium every other day thereafter until samples were collected.
Immunofluorescence
CM samples were collected at day 30 to 40 and transferred onto Millicell EZ slides (MilliporeSigma), fixed in 4% paraformaldehyde (PFA) solution for 8 minutes, and washed with phosphate-buffered saline (PBS) + 1.5 g/L glycine. Once ready for staining, all cells were blocked and permeabilized with 10% normal goat serum, 0.5% bovine serum albumin (BSA), and 0.5% Triton X-100 in PBS for 1 hour at room temperature. After washing, primary antibodies were diluted in antibody solution (1% normal goat serum, 0.5% BSA, 0.05% Triton X-100, all dissolved in PBS), added to the samples, and incubated overnight at 4°C. The primary antibody used in this study was mouse monoclonal cardiac Troponin T (Abcam AB8295). The next day, cells were washed, and secondary antibodies were diluted in antibody solution at 1:200 (with 0.05% Triton X-100) and added for 2 hours at room temperature in the dark. For the CC2 line, the secondary antibody used was polyclonal AlexaFluor-568-conjugated goat anti-mouse IgG (Invitrogen A21124). For the LVIP line, the secondary antibody used was polyclonal AlexaFlour-488-conjugated goat anti-mouse IgG (Invitrogen A21042). 4’,6-diamidino-2-phenylindole (DAPI, Thermo Fisher Scientific) was then added immediately at a concentration of 1:1000 to label DNA for 10 minutes in the dark. Stained slides were washed 3 times in PBS +1.5 g/L glycine, and coverslips (VWR) were added using ProLong Gold Antifade Mountant (Thermo Fisher Scientific). Immunofluorescence was utilized as it allows protein localization to be clearly viewed, so that differentiation efficiency could be determined by staining for proteins specific to cardiomyocytes.
Gene expression analysis
Total RNA was isolated from hiPSC-CMs cultured on 6-well plates using the RNeasy Mini Kit (Qiagen, Cat. #74106). Complementary DNA (cDNA) was then generated using the High-Capacity cDNA Reverse Transcription Kit with RNase Inhibitor (Applied Biosystems, Cat. #4374966) according to the manufacturer’s protocol. Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) plates were prepared in 384-well format as follows: 10 μL TaqMan Universal Master Mix II with UNG (Applied Biosystems, Cat. #4440045), 8 μL H₂O, 1 μL TaqMan Gene Expression Assay (Applied Biosystems), and 1 μL cDNA per well. The qRT-PCR plates were run on a QuantStudio 7 Real-Time PCR System (Applied Biosystems) for 50 amplification cycles, with four replicates per sample/primer pair. Relative expression levels of target genes TNNT2 and ACTN2 were determined by normalizing their Ct values to the Ct value of the housekeeping gene GAPDH, and fold changes were calculated using the 2⁻ΔΔCt method.
Statistical analysis
Images were taken using the Cytation 10 Imaging Reader (BioTek). To analyze the data, the number of DAPI+ and cTnT+ cells was manually counted using ImageJ Fiji software. This allowed the researchers to determine differentiation efficiency, which was the measure of optimal differentiation in this study. All graphs were made using Microsoft Excel. Statistical significance was evaluated with a standard unpaired student’s t-test (2-tailed) when comparing two groups or 1-way ANOVA with Tukey’s post-test for multiple comparison analysis. For all tests, a p-value less than 0.05 was considered statistically significant. All data are presented as mean ± SEM. In this study, the independent variable was the CHIR concentration added to the cells (6, 8, 10, and 12 μM CHIR). The dependent variable was differentiation efficiency, measured by the percentage of DAPI-stained nuclei that were also positive for cTnT, a specific marker for CMs.
Results
6 μM CHIR led to optimal hiPSC-CM differentiation efficiency in the CC2 line, with higher concentrations leading to decreased efficiency
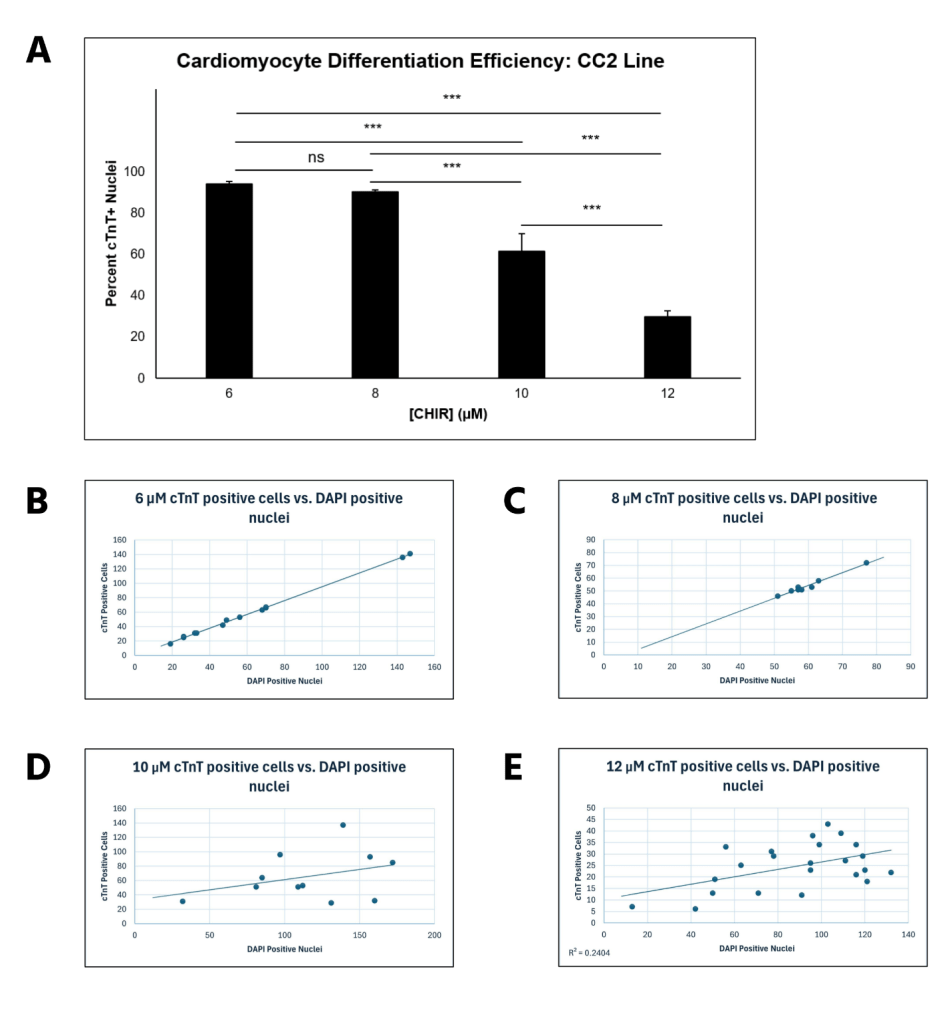
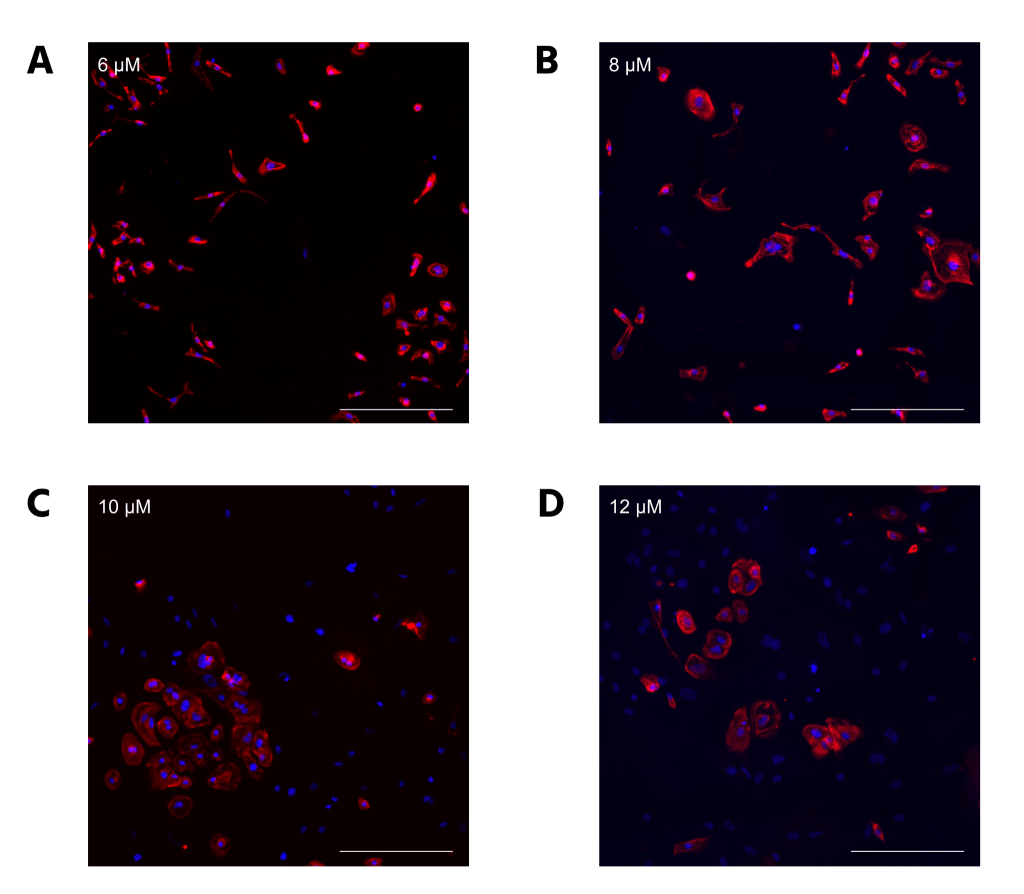
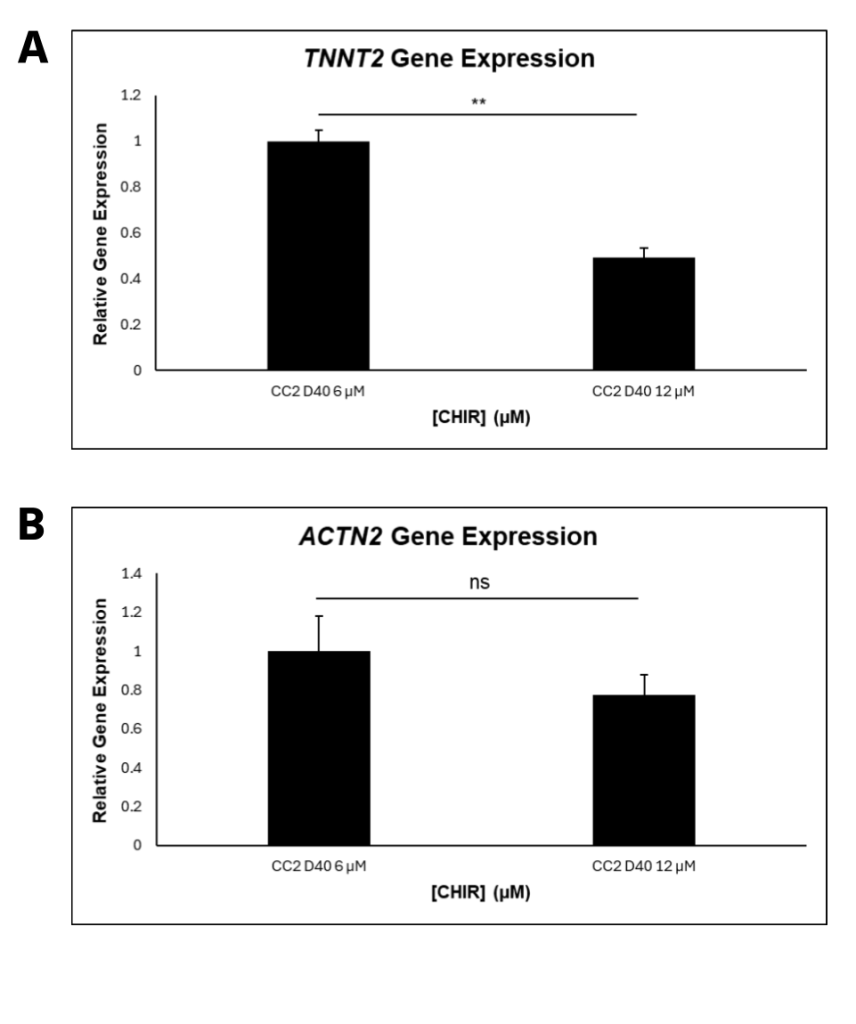
CC2 cells demonstrated a statistically significant decline in differentiation efficiency from 6 μM to 12 μM CHIR, measured through the percentage of DAPI-stained nuclei that were cTnT positive (Figure 2A). However, the decrease in efficiency from 6 to 8 μM CHIR was not statistically significant when a 1-way ANOVA test was conducted. The scatterplots (Figure 2B-E) demonstrate that hiPSCs treated with higher CHIR concentrations also had the highest variation, verified by calculating R2 values. Hence, 6 μM CHIR not only led to the highest differentiation efficiency, but also the highest consistency in the CC2 line. Because the dots represent different differentiation attempts, the scatterplots further suggest that between differentiation attempts, higher CHIR concentrations lead to less consistency in differentiation. The quantitative data is also supported by immunofluorescence, with a decrease in differentiation efficiency as CHIR concentration increases (Figure 3). At higher concentrations, a lower number of nuclei (blue) stain positive for cTnT (red). qRT-PCR experiments (Figure 4) also support the immunostaining results. Relative gene expression of TNNT2, which encodes the cTnT protein, statistically decreased from 6 to 12 μM CHIR, indicating that fewer cells were cardiomyocytes at 12 μM CHIR relative to 6 μM CHIR. Although the difference in relative gene expression for ACTN2 was not statistically significant, a 22.54% decrease in relative expression could be biologically relevant, and statistical significance was likely not achieved due to a relatively low sample size.
6 μM CHIR led to optimal hiPSC-CM differentiation efficiency in the LVIP line, with 12 μM leading to decreased efficiency
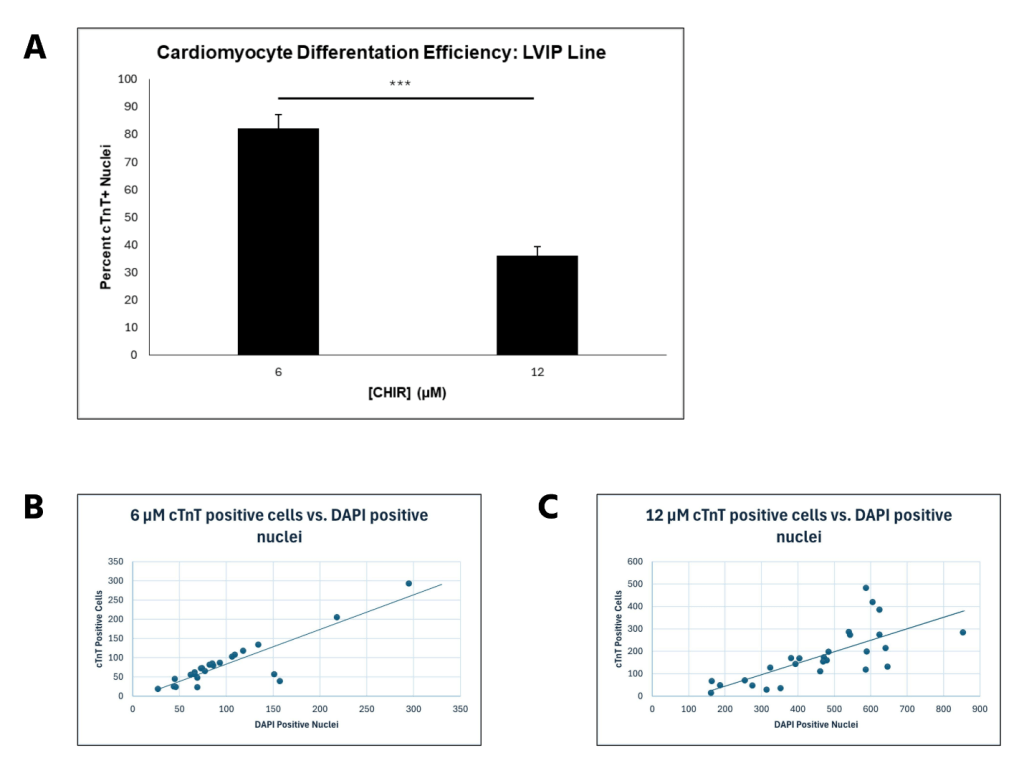
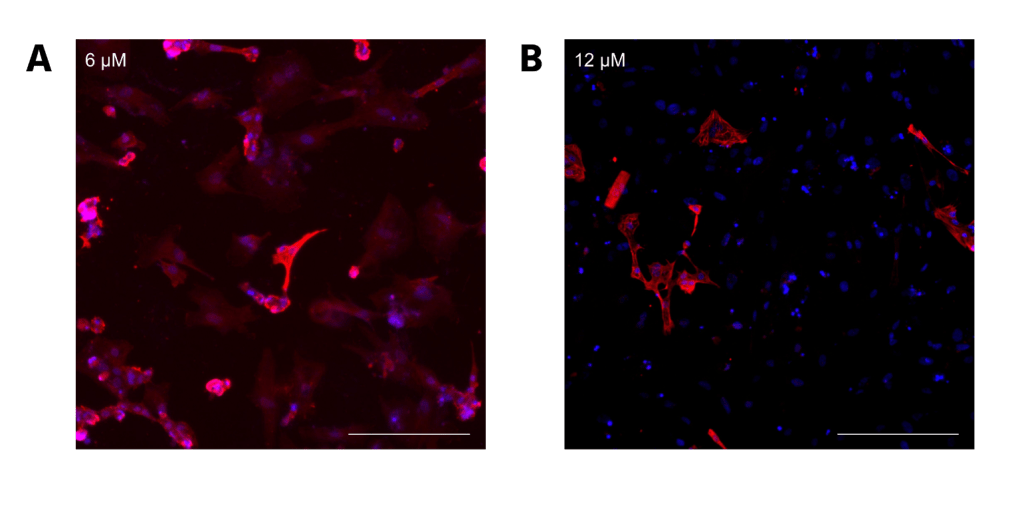
For LVIP cells containing the RTTN mutation, a similar trend to CC2 cells was observed, with average differentiation efficiency decreasing from 82.22% to 36.07% (P < 0.00000001) as CHIR concentration increased from 6 to 12 μM (Figure 5A). However, intermediate CHIR concentrations were not evaluated, so it is possible that an altered trend could be observed with values such as 8 and 10 μM CHIR. The scatterplots demonstrate that differentiation efficiency for cells treated with 12 μM CHIR did not fit the trend line as well as it did for cells treated with 6 μM CHIR. Moreover, the correlation was relatively low compared to the CC2 cell line, verified by the R2 values for both scatterplots. Therefore, the data for the LVIP cell line also support greater differentiation consistency when lower CHIR concentrations are used, but less than that of the CC2 line. (Figure 5B, C). These observations are further supported by visual representations of the immunostainings (Figure 6), depicting a decrease in differentiation efficiency, as a lower percentage of cells stained for cTnT when transitioning from 6 to 12 μM CHIR.
Discussion
Stem cells have emerged as a promising tool for modeling diseases and investigating therapeutics. Their potential use for treating cardiovascular disease has gained recent attention, as the majority of adult cardiomyocytes are terminally differentiated and cannot regenerate after damage, thereby opening the door for stem cell-derived therapies4. To study this concept further, proper and efficient differentiation of stem cells into CMs is necessary, as this process can differ for specific cell lines. Additionally, research has shown that centrosomal proteins may play a role in the development of cardiovascular disease, but further research is needed to uncover the mechanisms. In this study, we show that 6 μM CHIR99021 promotes the best differentiation efficiency for both wild-type CC2 cells and LVIP cells, used here as a model of infantile dilated cardiomyopathy. The results of this study disprove the hypothesis that 12 μM CHIR provides the best differentiation efficiency. This study also provides an effective differentiation protocol to generate human CMs for the study of cardiovascular development and disease.
Our results indicate that 6 μM CHIR was the optimal concentration for both cell lines, resulting in the highest differentiation efficiency. As CHIR concentration increased, particularly at 12 μM, differentiation efficiency significantly decreased. This observation is consistent with research showing that excessive Wnt/β-catenin pathway activation leads to abnormal cell behavior such as uncontrolled proliferation and impaired differentiation. Interestingly, this abnormal behavior is associated with the development of colorectal cancer21. The dose-dependent decline in differentiation efficiency observed in our study highlights the balance required when using small molecules like CHIR for stem cell differentiation. The two cell lines displayed different differentiation efficiencies at 6 μM CHIR (94.28% for CC2 hiPSC-CMs and 82.22% for LVIP hiPSC-CMs), suggesting that genetic differences, such as mutations affecting centrosomal proteins, can alter the cellular response to differentiation cues. Moreover, centrosomal proteins are known to regulate microtubules and cellular signaling22, so their localization could play a role in regulating stem cell differentiation. The lower differentiation efficiency in LVIP cells could also be linked to altered cellular responses to Wnt signaling, which might be less effective in these cells due to the RTTN mutation. These results highlight the importance of understanding the genetic background of hiPSC lines when designing differentiation protocols.
The variability in differentiation efficiency at higher CHIR concentrations—particularly in the 10 μM and 12 μM groups—raises an important limitation regarding experimental reproducibility. The large standard deviations and low R2 values in the scatterplots suggest that inconsistencies in the CHIR treatment process may have contributed to the observed data variability. This variability could be due to several factors, such as differences in the quality of the hiPSCs, minor discrepancies in pipetting, or genetic differences within the hiPSC populations. One study suggests that cell culture conditions and stages of the cell cycle can impact the differentiation efficiency of hiPSCs to CMs, so these factors may have also played a role in our study23. These findings further underscore the need to optimize experimental protocols to minimize such variability, ensuring consistent and reliable results for stem cell differentiation.
While 6 μM CHIR led to optimal results for both cell lines in this study, future research should investigate whether this concentration is universally effective across other hiPSC lines. Additionally, intermediate and lower concentrations of CHIR should be evaluated to determine whether differentiation efficiency is affected at these concentrations. These studies would help identify the precise concentrations needed to maximize differentiation for various cell lines. Future studies on the ideal concentrations of other small molecules used in CM differentiation, such as IWP-2, Wnt-C59, XAV-939, and DMH110, would also help ensure the maximum number of cells are successfully differentiated. Future research could expand the study to include in vivo models or cardiac organoid systems24, which provide more accurate representations of the human body to understand how CHIR treatment influence cardiac differentiation. In vivo models could also help determine whether the effects observed in this study are consistent with those in a more complex, three-dimensional environment, where cellular interactions and tissue structures are better represented25. Optimizing differentiation in organoid systems would allow for more detailed analysis of how differentiating CMs integrate into developing heart tissue and interact with other cardiac cell types.
During cardiomyocyte maturation, the centrosome undergoes structural reorganization, where its components re-localize from the centriole to the nuclear envelope in a process known as centrosome reduction. Moreover, centrosomal changes are associated with CM terminal differentiation, indicating they play a crucial role in CM maturation26. These changes suggest that centrosomal proteins are crucial not only for cellular function but also for heart development and potentially cardiac disease14. In general, the relationship between cardiac disease and centrosomal proteins is largely unexplored. Through this study, we developed an effective differentiation protocol to explore the localization of these proteins in healthy and diseased cardiomyocytes.
In conclusion, our study provides valuable insights into the optimal CHIR99021 concentration for cardiomyocyte differentiation from hiPSCs. By examining the effects of varying concentrations and exploring the differential responses of specific cell lines, protocols can be refined to enhance the efficiency and reproducibility of CMs. Furthermore, the hypothesis of centrosomal protein localization as a potential regulator of differentiation efficiency offers an exciting avenue for future studies aimed at investigating the molecular mechanisms of cardiac development and disease, especially in pediatric cases. Ultimately, these findings pave the way for optimal hiPSC differentiation in future studies, thereby opening the door for discoveries at the cellular and molecular level in human cardiomyocytes.
References
- S. S. Martin, A. W. Aday, N. B. Allen, Z. I. Almarzooq, C. A. M. Anderson, P. Arora, C. L. Avery, C. M. Baker-Smith, N. Bansal, A. Z. Beaton, Y. Commodore-Mensah, M. E. Currie, M. S. V. Elkind, W. Fan, G. Generoso, B. B. Gibbs, D. G. Heard, S. Hiremath, M. C. Johansen, D. S. Kazi, D. Ko, M. H. Leppert, J. W. Magnani, E. D. Michos, M. E. Mussolino, N. I. Parikh, S. M. Perman, M. Rezk-Hanna, G. A. Roth, N. S. Shah, M. V. Springer, M.-P. St-Onge, E. L. Thacker, S. M. Urbut, H. G. C. Van Spall, J. H. Voeks, S. P. Whelton, N. D. Wong, S. S. Wong, K. Yaffe, L. P. Palaniappan. 2025 heart disease and stroke statistics: A report of US and global data from the American Heart Association. Circulation, 151, e41–e660 (2025). [↩]
- M. A. Laflamme, C. E. Murry. Heart regeneration. Nature, 473, 326–335 (2011). [↩]
- E. A. Woodcock, S. J. Matkovich. Cardiomyocytes structure, function and associated pathologies. The International Journal of Biochemistry & Cell Biology, 37, 1746-1751 (2005). [↩]
- O. Bergmann, R. D. Bhardwaj, S. Bernard, S. Zdunek, F. Barnabé-Heider, S. Walsh, J. Zupicich, K. Alkass, B. A. Buchholz, H. Druid, S. Jovinge, J. Frisén. Evidence for cardiomyocyte renewal in humans. Science, 324, 98-102 (2009). [↩] [↩]
- D. Fan, A. Takawale, J. Lee, Z. Kassiri. Cardiac fibroblasts, fibrosis and extracellular matrix remodeling in heart disease. Fibrogenesis & Tissue Repair, 5, 15 (2012). [↩]
- J. Gulati, M. Zhu, J. Gilbreth, S. Wang. The use of stem cells in cardiac pathologies: a review. Georgetown Medical Review, 7, 1-13 (2024). [↩]
- R. Hesselbarth, T. U. Esser, K. Roshanbinfar, S. Schrüfer, D. W. Schubert, F. B. Engel. CHIR99021 promotes hiPSC-derived cardiomyocyte proliferation in engineered 3D microtissues. Advanced Healthcare Materials, 10, e2100926 (2021). [↩]
- S. J. Kühl, M. Kühl. On the role of Wnt/β-catenin signaling in stem cells. Biochimica et Biophysica Acta (BBA), 1830, 2297-22306 (2013). [↩] [↩]
- S. Y. Sokol. Maintaining embryonic stem cell pluripotency with Wnt signaling. Development, 138, 4341–4350 (2011). [↩]
- D. M. Lyra-Leite, O. Gutiérrez-Gutiérrez, M. Wang, Y. Zhou, L. Cyganek, P. W. Burridge. A review of protocols for human iPSC culture, cardiac differentiation, subtype-specification, maturation, and direct reprogramming. STAR Protocols, 3, 101560 (2022). [↩] [↩]
- R. G.C. Maas, S. Lee, M. Harakalova, C. J.B. Snijders Blok, W. R. Goodyer, J. Hjortnaes, P. A.F.M. Doevendans, L. W. Van Laake, J. van der Velden, F. W. Asselbergs, J. C. Wu, J. P.G. Sluijter, S. M. Wu, J. W. Buikema. Massive expansion and cryopreservation of functional human induced pluripotent stem cell-derived cardiomyocytes. STAR Protocols, 2, 100334 (2021). [↩]
- H. Kempf, R. Olmer, C. Kropp, M. Rückert, M. Jara-Avaca, D. Robles-Diaz, A. Franke, D. A. Elliott, D. Wojciechowski, M. Fischer, A. R. Lara, G. Kensah, I. Gruh, A. Haverich, U. Martin, R. Zweigerdt. Controlling expansion and cardiomyogenic differentiation of human pluripotent stem cells in scalable suspension culture. Stem Cell Reports, 3, 1132-1146 (2014). [↩]
- V. C. Chen, J. Ye, P. Shukla, G. Hua, D. Chen, Z. Lin, J. C. Liu, J. Chai, J. Gold, J. Wu, D. Hsu, L. A. Couture. Development of a scalable suspension culture for cardiac differentiation from human pluripotent stem cells. Stem Cell Research, 15, 365-375 (2015). [↩]
- Y. W. Chun, M. Miyamoto, C. H. Williams, L. R. Neitzel, M. Silver-Isenstadt, A. G. Cadar, D. T. Fuller, D. C. Fong, H. Liu, R. Lease, S. Kim, M. Katagiri, M. D. Durbin, K.C. Wang, T. K. Feaster, C. C. Sheng, M. D. Neely, U. Sreenivasan, M. Cortes-Gutierrez, A. V. Finn, R. Schot, G. M.S. Mancini, S. A. Ament, K. C. Ess, A. B. Bowman, Z. Han, D. P. Bichell, Y. R. Su, C. C. Hong. Impaired reorganization of centrosome structure underlies human infantile dilated cardiomyopathy. Circulation, 147, 1291-1303 (2023). [↩] [↩] [↩]
- X. Lian, J. Zhang, S. M. Azarin, K. Zhu, L. B. Hazeltine, X. Bao, C. Hsiao, T. J. Kamp, S. P. Palecek. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/β-catenin signaling under fully defined conditions. Nature Protocols, 8, 162-175 (2012). [↩] [↩]
- X. Lian, C. Hsiao, G. Wilson, K. Zhu, L. B. Hazeltine, S. M. Azarin, K. K. Raval, J. Zhang, T. J. Kamp, S. P. Palecek. Robust cardiomyocyte differentiation from human pluripotent stem cells via temporal modulation of canonical Wnt signaling. Proceedings of the National Academy of Sciences of the United States of America, 109, e1848-1857 (2012). [↩]
- X.X. Qiu, Y. Liu, Y.F. Zhang, Y.N. Guan, Q.Q. Jia, C. Wang, H. Liang, Y.Q. Li, H.T. Yang, Y.W. Qin, S. Huang, X.X. Zhao, Q. Jing. Rapamycin and CHIR99021 coordinate robust cardiomyocyte differentiation from human pluripotent stem cells via reducing p53‐dependent apoptosis. Journal of the American Heart Association, 6, e005295 (2017). [↩]
- T. K. Feaster, A. G. Cadar, L. Wang, C. H. Williams, Y. W. Chun, J. E. Hempel, N. Bloodworth, D. W. Merryman, C. C. Lim, J. C. Wu, B. C. Knollmann, C. C. Hong. Matrigel mattress: a method for the generation of single contracting human-induced pluripotent stem cell-derived cardiomyocytes. Circulation Research, 117, 995-1000 (2015). [↩]
- P. W. Burridge, E. Matsa, P. Shukla, Z. C. Lin, J. M. Churko, A. D. Ebert, F. Lan, S. Diecke, B. Huber, N. M. Mordwinkin, J. R. Plews, O. J. Abilez, B. Cui, J. D. Gold, J. C. Wu. Chemically defined generation of human cardiomyocytes. Nature Methods, 11, 855-860 (2014). [↩]
- C. Halloin, K. Schwanke, W. Löbel, A. Franke, M. Szepes, S. Biswanath, S. Wunderlich, S. Merkert, N. Weber, F. Osten J. de la Roche, F. Polten, K. Wollert, T. Kraft, M. Fischer, U. Martin, I. Gruh, H. Kempf, R. Zweigerdt. Continuous WNT control enables advanced hPSC cardiac processing and prognostic surface marker identification in chemically defined suspension culture. Stem Cell Reports, 13, 366-379 (2019). [↩]
- H. Zhao, T. Ming, S. Tang, S. Ren, H. Yang, M. Liu, Q. Tao, H. Xu. Wnt signaling in colorectal cancer: pathogenic role and therapeutic target. Molecular Cancer, 21, 144 (2022). [↩]
- C. Arquint, A.M. Gabryjonczyk, E. A. Nigg. Centrosomes as signalling centres. Philosophical Transactions of the Royal Society of London, 369, 20130464 (2014). [↩]
- F. Laco, T. L. Woo, Q. Zhong, R. Szmyd, S. Ting, F. J. Khan, C. L.L. Chai, S. Reuveny, A. Chen, S. Oh. Unraveling the inconsistencies of cardiac differentiation efficiency induced by the GSK3β inhibitor CHIR99021 in human pluripotent stem cells. Stem Cell Reports, 10, 1851-1866 (2018). [↩]
- M. Miyamoto, L. Nam, S. Kannan, C. Kwon. Heart organoids and tissue models for modeling development. Seminars in Cell and Development Biology, 118, 119-128 (2021). [↩]
- P. Mukherjee, S. Roy, D. Ghosh, S. K. Nandi. Role of animal models in biomedical research: a review. Laboratory Animal Research, 38, 18-34 (2022). [↩]
- D. C.H. Ng, D. K. Richards, R. J. Mills, U. Y. Ho1, H. L. Perks, C. R. Tucker, H. K. Voges, J. K. Pagan, J. E. Hudson. Centrosome reduction promotes terminal differentiation of human cardiomyocytes. Stem Cell Reports, 15, 817-826 (2020). [↩]