Abstract
Background: Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), responsible for the COVID-19 pandemic, mutates quickly and develops resistance to established therapies, including neutralizing antibodies and vaccines. SARS-CoV-2 enters human cells via a host receptor, angiotensin-converting enzyme 2 (ACE2), through its spike protein. The affinity of the spike protein binding to the ACE2 receptor is a key determinant for the virulence of SARS-CoV-2 variants.
Methods: This study evaluated the evolving binding levels of the SARS-CoV-2 spike protein receptor binding domains (RBD) with human ACE2 protein. Expression vectors of membrane-bound RBD domain-containing proteins were constructed. RBD domain plasmids were transiently transfected into HEK293T cells. Fluorescence-labeled ACE2 fusion protein was then used to assess the binding levels to different RBD domain transfected HEK293T cells using flow cytometry. Additionally, etesevimab, an anti-SARS-CoV-2 monoclonal antibody derived from a patient recovering from the original strain, was evaluated for the interaction with RBD-domain transfectants and the ability to block ACE2 from binding to RBD.
Results: A decreased binding level of ACE2-RBD interaction was observed when SARS-CoV-2 evolved from the original strain to the Delta strain, followed by an increased binding levels for Omicron BA.1 and BA.2 strains. Etesevimab bound to the RBD domain of the original and Delta strains strongly, lost binding to Omicron BA.1 variant but regained some binding level to an Omicron BA.2 strain. Etesevimab maintained some blocking capacity for spike protein-hACE2 interactions for all SARS-CoV-2 variants tested.
Conclusion: Assessing etesevimab’s binding and blocking capacity might help determine its ability to treat emerging SARS-CoV-2 strains.
Keywords: SARS-CoV-2, COVID-19, ACE2, neutralizing antibody, etesevimab
Introduction
COVID-19 has left its mark as one of the most widespread diseases in recent history. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a strain of coronavirus that causes COVID-19. As of March 11, 2025, the World Health Organization (WHO) (https://www.who.int/) has reported 778 million confirmed cases of COVID-19 and 7.1 million deaths worldwide (WHO et al., 2025). SARS-CoV-2 engages human cells via a host receptor ACE2 through its spike protein. The affinity of the spike protein RBD to the ACE2 receptor thus holds a key determinant for the virulence of SARS-CoV-2 variants1.
Before the availability of vaccines and anti-viral small molecule drugs, SARS-CoV-2 neutralizing antibody or antibody cocktail emerged as the first viral specific treatment receiving emergency use authorization (EUA) by the US FDA, including casirivimab plus imdevimab2 (EUA issued November 2020), bamlanivimab (EUA issued November 2020), and bamlanivimab plus etesevimab3 (EUA issued February 2021), sotrovimab (EUA issued May 2021) and bebtelovimab (EUA issued February 2022). While the vaccine boosts host immunity over time, SARS-CoV-2 specific neutralizing antibody provides immediate, passive immunity that limits disease progression, and reduces hospitalization and death in high-risk populations. Neutralizing antibodies slow down viral spreading by preventing the virus entry into host cells. Etesevimab, a neutralizing antibody derived from a patient recovered from COVID-19 Wuhan strain, targets the RBD’s receptor-binding motif (RBM), which includes residues 438–506, and blocks RBD-ACE2 interactions4. Notably, the EUA for bamlanivimab monotherapy was revoked in April 2021 due to reduced efficacy against emerging variants, highlighting the rapid adaptation required for effective antibody therapies, and an antibody cocktail composed of antibodies targeting distinctive domains on SARS-CoV-2 is preferred over a single neutralizing antibody to reduce the chance of evolving resistance.
As a single-stranded RNA virus, SARS-CoV-2 mutates quickly and develops resistance to established therapies, including neutralizing antibodies5 and vaccines6. When SARS-CoV-2 omicron and its subvariants became dominant strains in late 2021, almost all EUA approved neutralizing antibody or cocktail can no longer neutralize the virus effectively, leading to the eventual revokes of all EUAs by the FDA. Fortunately, omicron variant specific vaccines and other small molecule anti-viral drug such as Paxlovid, have become available and remained effective against prevailing strains as of today.
The current study evaluated the evolving binding levels of the RBD of the spike protein from the SARS-CoV-2 variants (original Wuhan strain, Delta, Omicron BA.1 and BA.2) with human ACE2 protein. One neutralizing antibody, etesevimab, was also evaluated for binding to the RBD-domain of SARS-CoV-2 variants, and for its ability to block ACE2 binding. While previous studies7, have investigated the RBD-ACE2 interaction, they have not comprehensively assessed how different SARS-CoV-2 mutations impact both receptor binding and neutralization by a therapeutic antibody. This study fills that gap by systematically evaluating binding levels across multiple variants and impacting on etesevimab, providing insights into the molecular evolution of SARS-CoV-2 and its implications for antibody-based therapies.
2. Results
2.1 Binding of ACE2-mIg fusion protein to HEK293T transfected with RBD variants
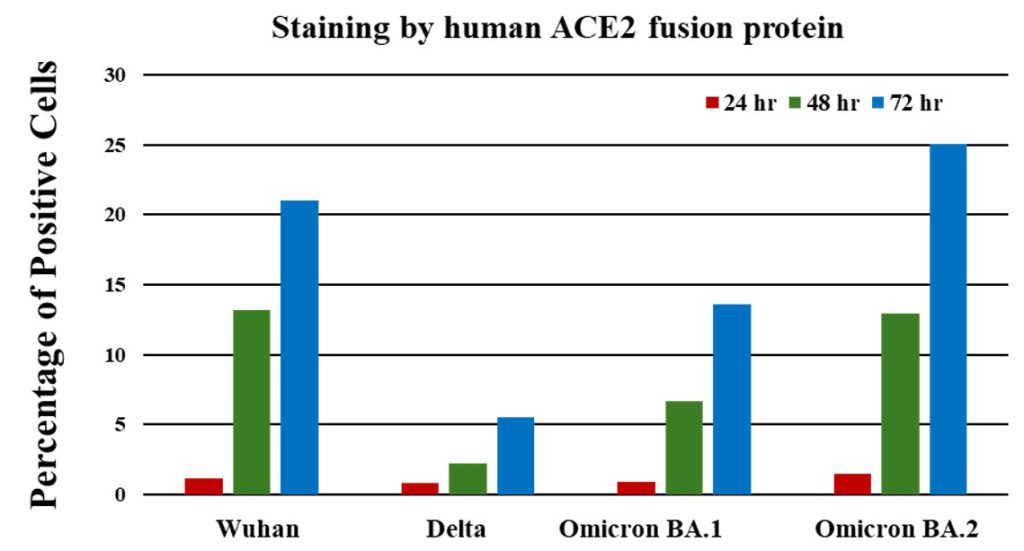
An increased binding of ACE2-mIg fusion protein to HEK293T cells transfected with RBD variants was observed over time and peaked on day 3 (Figure 1), indicating the transient expression reached peak around day 3. An initial decreased binding level of ACE2-RBD interaction was observed when SARS-CoV-2 evolved from the original Wuhan strain to the delta strain, then followed by increased binding levels for omicron BA.1 and BA.2 strains, with omicron BA.2 strain has the highest binding level.
2.2 Binding of etesevimab to HEK293T transfected with RBD variants
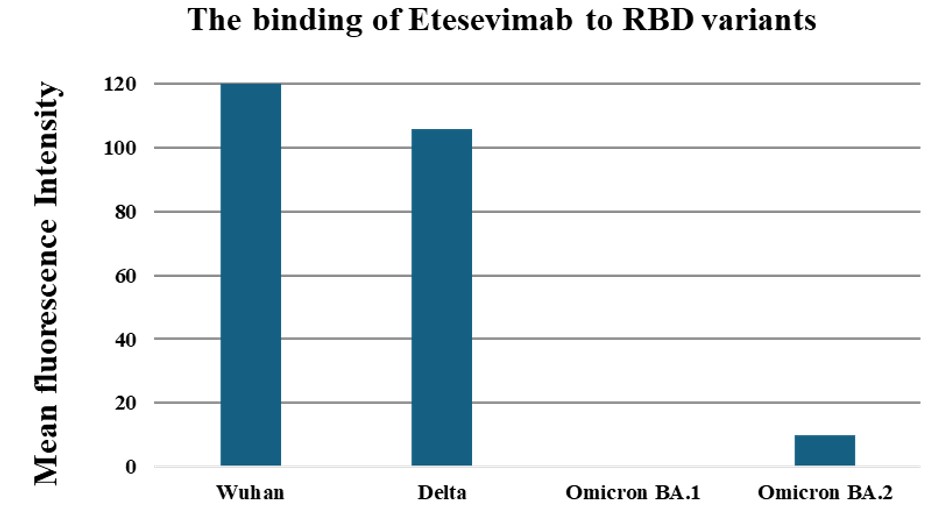
Etesevimab is a neutralizing antibody derived from a patient recovered from COVID-19 Wuhan strain. As the SARS-CoV-2 evolved from Wuhan to delta strain, a decreased binding of etesevimab to the spike protein was observed. Etesevimab lost binding to Omicron BA.1 completely, however regained some binding to a later Omicron BA.2 strain (Figure 2).
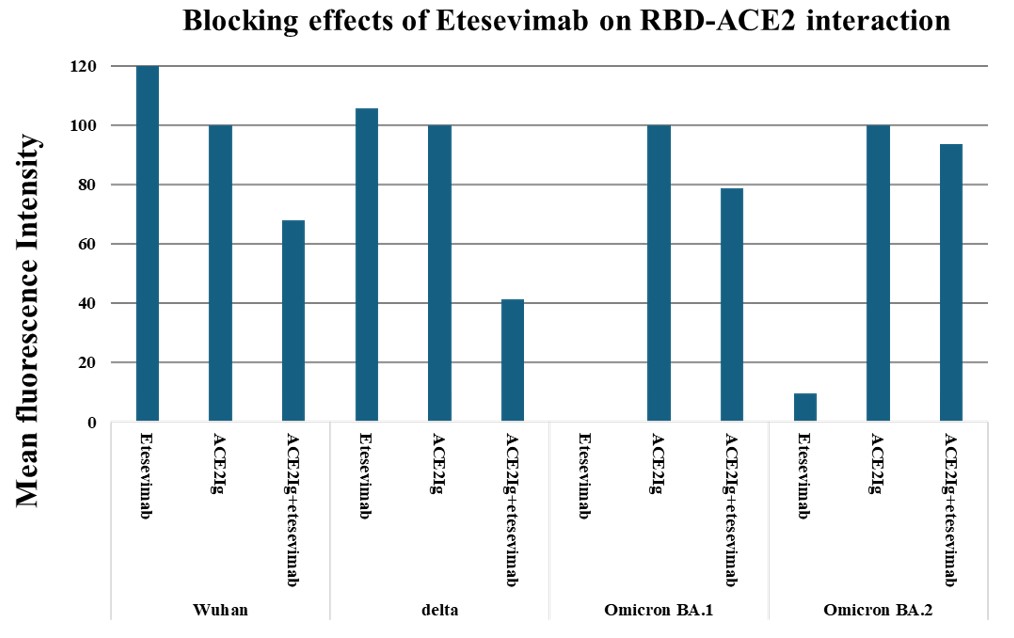
As the virus evolved, the binding levels of etesevimab to omicron RBD domain decreased and therefore its capacity to block the binding between the omicron RBD and ACE2 fusion protein also decreased (Figure 3). Notably, although etesevimab lost binding to BA.1 strain RBD completely, it maintained some blocking capacity for RBD-ACE2 interaction.
Discussion
In the wake of the COVID19 global pandemic, the patient-derived neutralizing antibody or antibody cocktail emerged as the first viral specific treatment approved by the US FDA. In February 2021, the FDA granted emergency use authorization for bamlanivimab plus etesevimab to treat mild to moderate COVID-19 as the cocktail limits disease progression and reduces hospitalization and death in high-risk populations. Due to its fast-mutating nature as an RNA virus, SARS-COV-2 evolved quickly and developed resistance to established treatment. In December 2023, FDA revoked the emergency use authorization of bamlanivimab plus etesevimab as the antibody cocktail can no longer neutralize SARS-COV-2 Omicron stains (BA.1, BA.2 and BA.5) effectively.
In the current study, the scientific basis for the approval and the revocation of EUA was demonstrated. In a set of cell-based experiments, the binding of the receptor binding domain of SARS-CoV-2 variants with human ACE2 protein was investigated. An increased binding levels for ACE2-RBD interaction was observed in the Omicron strains, as previously reported7. Etesevimab showed strong binding to the RBD domains of the Wuhan and Delta strains, but not to the Omicron strains. Consistently, etesevimab blocked the ACE2-RBD interaction more effectively for Wuhan and Delta strains than for the Omicron strains. The accumulated mutations in the RBD domain of the Omicron strains likely changed the configuration of the spike protein, leading to the greatly reduced binding for etesevimab. Notably, mutations at positions K417 and E484 have been associated with reduced neutralization efficacy of etesevimab. Interestingly, although etesevimab lost binding to Omicron BA.1 strain completely, it maintained some blocking capacity. One possible explanation for this observation is that while as assessed by flow cytometry, Omicron BA.1 lost apparent binding to RBD, but it retained some weak binding to RBD undetectable by the flow cytometry method, or regained some binding to RBD in the presence of ACE2, which led to some blocking capacity for RBD-ACE2 interaction.
There are several limitations of the study. First of all, the binding analysis was based on transient transfected RBD domains on HEK293T cells. Variable protein expression levels are expected from different transient transfection and thus impact the accuracy of the binding analysis. Stably transfected HEK293T cells would be preferred. Secondly, the membrane expressed RBD fusion protein as well as the human ACE2-mIg fusion protein do not represent the exact native configurations of the viral spike protein and membrane expressed human ACE2 receptor. A pseudo-virus with complete spike protein and a human cell line with ACE2 expression would be the better system to study virus entry and fusion with host cells.
4. Conclusion
In summary, the current study used a cell based binding analysis system to study the interactions of SARS-CoV-2 variants spike protein receptor binding domain with human ACE2 protein as well as the binding of SARS-CoV-2 variants by an FDA approved neutralizing antibody, etesevimab. The results demonstrated that as SARS-CoV-2 evolved, Omicron variants have higher binding levels to the human ACE2 receptor and escaped the recognition by etesevimab. As SARS-CoV-2 continues to evolve, previously discontinued neutralizing antibodies should be continuously monitored for their neutralizing effects and might be repurposed for new emerging strains if they regain binding and blocking capacity.
5. Materials and Methods
5.1 Plasmids, fusion proteins and monoclonal antibodies.
Human expression plasmid containing a signal peptide “MFVFLVLLPLVSSQC” fused with SARS-CoV-2 (Wuhan strain) Spike Protein receptor binding domain and a his-tag (pCAGGS-RBD-his) was provided by TopAlliance Biosciences. The RBD domain included in the plasmid consists of residues 319 – 541 AA in S1 subunit of the spike protein. Additional variants (Delta, Omicron BA.1 and BA.2) were made using PCR based cloning methods. Paired primers were used to generate specific DNA sequences encoding various COVID-19 RBD domains. PCR products were then digested with (EcoR I/Xho I) and ligated using T4 DNA ligase into pCAGGS-RBD-his expression plasmids. Primers used in the PCR included: PRIMER_TA276, AGATCTGCTAGCTCGAGTCAGTGGTGGTGGTGGTGGTGAAAG; PRIMER_TA277, TTGTGATCAAAGGAGATGAAGTGAGACAAATCGCACC; PRIMER_TA320, AATCCATCCACTCCGTTGCAAGGTGTTG. PrimeSTAR GXL DNA polymerase from TaKaRa was used. Detailed PCR protocol included: step 1. 98℃ for 5 minutes; step 2. 30cycles of 98℃ for 10 seconds, 60℃ for 15 seconds and 68℃ for 4 minutes; step 3. 68℃ for 5 minutes.
Cloned plasmids were all verified by sequencing. Sanger sequencing was performed with the cloned plasmid and PCR primers from both ends covering the entire inserted 4 Kb sequence.
Human ACE2 extracellular domain-mIgG2a Fc fusion protein (Biosciences et al., 2020) was produced by transient transfection in HEK293T cells (ATCC # CRL-3216) and purified by protein A column. Specifically, human ACE2 extracellular domain residues position 18-740 was fused with mouse IgG2a heavy chain residue position 98-330, containing the hinge, CH2 and CH3 domains. Etesevimab drug product was provided by TopAlliance Biosciences (20200402 et al., 2020). APC labeled mouse anti-human Ig-Fc (409306 et al., 4093) and PE-labeled goat anti-mouse Ig (405307 et al., 4053) were purchased from Biolegend (San Diego, California, USA).
5.2 HEK293T transient transfection by polyethylenimine (PEI) method
HEK293T cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine calf serum (FBS). For transient transfection, 0.8 µg of expression plasmid DNA, encoding the membrane-bound domain of Spike protein of the SARS-CoV-2 variants, was mixed with 50 µL of Opti-MEM medium. Separately, 8.0 µL of polyethylenimine (PEI) was also diluted in 50 µL of Opti-MEM medium. Both mixtures were incubated at room temperature for 5 minutes. The DNA and PEI solutions were then combined, mixed thoroughly, and incubated for 15–30 minutes to allow for complex formation. A total of 100 µL of the plasmid DNA-PEI mixture was added to each well of cells. PEI used in the study was PEI MAX® (24765 et al., 2476), linear polyethylenimine hydrochloride (MW 40,000) from Polysciences. A transfection control is provided by co-transfection with a GFP-containing control plasmid in each transfection.
5.3 Expression and Binding Analysis by Flow Cytometry
Flow cytometry analysis was conducted using FACS Canto II following manufacturer’s manual. Expression of RBD variants were accessed by incubation with etesevimab, a SARS-CoV-2 specific human antibody first, and followed by incubation with fluorescence labeled anti-human Ig secondary antibody. To determine the binding levels of the different COVID-19 variants to ACE-2, ACE2-mIg fusion protein (20 μg/uL, 1μg/50μL/tube) was first incubated with SARS-CoV-2 variants RBD domain transfected HEK293T cells for 30 minutes. Fluorescence labeled Anti-mouse Ig Fc secondary antibody was then added to the cell suspension. For this blocking assay, hACE2-mFc, 1.0 μg/50 μL /tube (20 μg /mL) was used, and etesevimab concentration ranging from 0, 1.6, 4.1, 10.2, 25.6, 64.0, 160.0 and 400.00 μg/mL, were used to establish a titration. Each flow cytometry study included cells without primary and secondary antibody, and cells with secondary antibody only as controls. For hACE2-mFc, the secondary antibody was PE-labeled goat anti-mouse Ig, BioLegend, Cat# 405307 (B253778 et al., 2537). For etesevimab, the secondary antibody was APC labeled mouse anti-human Ig-Fc, BioLegend, Cat# 409306 (B265810 et al., 2658). Flow cytometry data were captured using BD FACSDiva software and results were analyzed using Flowjo software. The reported staining results were the relative values after subtraction from secondary antibody only staining.
Acknowledgements
I would like to thank my mentors, Dr. Yuming Zhao and Dr. Lanqing Huang at TopAlliance Biosciences for their guidance on study design and conducting the experiments.
References
- J. Lan, J. Ge, J. Yu, S. Shan, H. Zhou, S. Fan, Q. Zhang, X. Shi, Q. Wang, L. Zhang, and X. Wang. 2020. ‘Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor’, Nature, 581: 215-20. [↩]
- D. M. Weinreich, S. Sivapalasingam, T. Norton, S. Ali, H. Gao, R. Bhore, J. Xiao, A. T. Hooper, J. D. Hamilton, B. J. Musser, D. Rofail, M. Hussein, J. Im, D. Y. Atmodjo, C. Perry, C. Pan, A. Mahmood, R. Hosain, J. D. Davis, K. C. Turner, A. Baum, C. A. Kyratsous, Y. Kim, A. Cook, W. Kampman, L. Roque-Guerrero, G. Acloque, H. Aazami, K. Cannon, J. A. Simon-Campos, J. A. Bocchini, B. Kowal, A. T. DiCioccio, Y. Soo, G. P. Geba, N. Stahl, L. Lipsich, N. Braunstein, G. Herman, G. D. Yancopoulos, and Investigators Trial. 2021. ‘REGEN-COV Antibody Combination and Outcomes in Outpatients with Covid-19’, N Engl J Med, 385: e81. [↩]
- M. Dougan, A. Nirula, M. Azizad, B. Mocherla, R. L. Gottlieb, P. Chen, C. Hebert, R. Perry, J. Boscia, B. Heller, J. Morris, C. Crystal, A. Igbinadolor, G. Huhn, J. Cardona, I. Shawa, P. Kumar, A. C. Adams, J. Van Naarden, K. L. Custer, M. Durante, G. Oakley, A. E. Schade, T. R. Holzer, P. J. Ebert, R. E. Higgs, N. L. Kallewaard, J. Sabo, D. R. Patel, M. C. Dabora, P. Klekotka, L. Shen, D. M. Skovronsky, and Blaze- Investigators. 2021. ‘Bamlanivimab plus Etesevimab in Mild or Moderate Covid-19’, N Engl J Med, 385: 1382-92. [↩]
- R. Shi, C. Shan, X. Duan, Z. Chen, P. Liu, J. Song, T. Song, X. Bi, C. Han, L. Wu, G. Gao, X. Hu, Y. Zhang, Z. Tong, W. Huang, W. J. Liu, G. Wu, B. Zhang, L. Wang, J. Qi, H. Feng, F. S. Wang, Q. Wang, G. F. Gao, Z. Yuan, and J. Yan. 2020. ‘A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2’, Nature, 584: 120-24. [↩]
- P. Wang, M. S. Nair, L. Liu, S. Iketani, Y. Luo, Y. Guo, M. Wang, J. Yu, B. Zhang, P. D. Kwong, B. S. Graham, J. R. Mascola, J. Y. Chang, M. T. Yin, M. Sobieszczyk, C. A. Kyratsous, L. Shapiro, Z. Sheng, Y. Huang, and D. D. Ho. 2021. ‘Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7’, Nature, 593: 130-35. [↩]
- G. Lobinska, A. Pauzner, A. Traulsen, Y. Pilpel, and M. A. Nowak. 2022. ‘Evolution of resistance to COVID-19 vaccination with dynamic social distancing’, Nat Hum Behav, 6: 193-206. [↩]
- L. Li, H. Liao, Y. Meng, W. Li, P. Han, K. Liu, Q. Wang, D. Li, Y. Zhang, L. Wang, Z. Fan, Y. Zhang, Q. Wang, X. Zhao, Y. Sun, N. Huang, J. Qi, and G. F. Gao. 2022. ‘Structural basis of human ACE2 higher binding affinity to currently circulating Omicron SARS-CoV-2 sub-variants BA.2 and BA.1.1’, Cell, 185: 2952-60 e10. [↩] [↩]