Abstract
“Time is brain”, every second matters when treating stroke patients. Every second, approximately 32,000 neurons are lost after a stroke, equivalent to 3.6 years of aging. Around 137000 people die of a brain stroke in the USA per year. 87% are due to ischemic stroke. Reducing the time to classify a stroke is important for administering accurate treatment. Ischemic and hemorrhagic strokes must be classified at the earliest, as the treatment paths are fundamentally different with respect to the onset time. This paper explains why multiple modalities are required to achieve a faster and more accurate treatment path for a stroke patient. The proposed model aims to develop an automated system and analyze the best classification model. This uses CT images to classify the stroke and MRI to determine the stroke onset time using machine-learning models. During the classification of hemorrhagic and ischemic strokes, CT images were preprocessed and normalized, and HOG features were extracted to train the model. This model achieved a 94.41% accuracy on the testing data. Using a dataset of 1878 patients who had either ischemic or hemorrhagic stroke, the SVM architecture was able to predict the types with an AUC of 0.9754. Post classification, the MRI images DWI-FLAIR mismatch identifies if the stroke onset time was within 4.5 hours. The ResNet-18 model on 250 images with the nibabel library achieved a specificity of 0.9524 in identifying DWI-FLAIR mismatch. This paper shows how multiple modalities are required for classifying different stroke types and how various combinations of imaging types at different stages of treatment can save cost, accelerate stroke classification, assist onset time detection, and also eliminate subjective analysis when humans are involved for more accurate treatment.
Keywords: Strokes, Ischemic, Hemorrhagic, MRI, CT, Support Vector Machine, ResNet-18, CNN, Machine learning, Deep Learning algorithm, thrombectomy, acute stroke
Introduction
According to NIH, ischemic strokes contribute to 87% of total strokes worldwide, while hemorrhagic strokes make up the other 13%1,2. As per the Stroke Awareness Foundation, there are 7 million stroke survivors who live in the USA, and around two-thirds of them are disabled. 795,000 people (which is about half the size of the population of Idaho) suffer from strokes each year, which makes it the second most common illness among humans3. In the US, people have strokes every 45 seconds and die from them every 4 minutes4.
Ischemic strokes occur when blood vessels are blocked by the buildup of plaque, preventing blood from reaching the brain, or when a clot develops in the cardiovascular system and travels to the brain. On the other hand, hemorrhagic strokes are ruptures in blood vessels that spill blood into the intracranial area of the brain5. In Acute Ischemic Stroke, Vascular Neurologists need to either eliminate plaque or clots inside the arteries to free up the bloodstream to treat strokes. Blood carries oxygen and nutrients to the brain, which keeps all cognitive functions alive. When blood does not reach the relevant areas in the brain, humans are prone to disabilities like impaired speech, physical restrictions, and possibly death5. Despite the modern-day advances in medical imaging, accurate and rapid classification of stroke subtypes and stroke timeline remains a huge clinical challenge6. AI image analysis conducted on MRI (Magnetic Resonance Imaging) and CT (Computed Tomography Scan) scans can help analyze and treat patients more quickly7, reduce brain damage, and improve survival rates. “Time is brain”, a common phrase used to emphasize the rapid and irreparable loss of human nervous tissue as the stroke progresses 8. Therapeutic interventions should be pursued immediately5. As per studies, the CT scans are faster, 5-10 min, compared to MRI, that take 20-60 min. CT is less expensive9 than MRI scans, and availability10 of CT is more than that of MRI.
The major success of stroke treatment lies in 3 prime factors, namely: classification to either hemorrhagic or ischemic stroke, as the treatment and medication depend on this classification. Secondly, determining the last wellness time in ischemic stroke, as a patient arrives before or after a 4.5-hour period11. Lastly, the size of the infarct versus penumbra helps neurologists choose eligible patients for thrombectomy12. Clot-buster drugs are also used for ischemic strokes; patients need to take the drug before a 4 1 ⁄ 2 hour period, or else the treatment can do more harm than good13. MRI and CT need to make sure the stroke is not hemorrhagic, or else this drug can lead to the death of the patient. Hemorrhagic strokes are surgically treated differently because doctors reduce bleeding and pressure of excessive fluids in the brain. Surgeons may be able to repair blood vessels to save patients from blood loss. Even though manual interpretation of stroke imaging has been resource-intensive, this method is still time-consuming and is subject to diagnostic error. This study addresses the importance of automation and accurate stroke treatment by deep learning models to support precise and quick decision-making for diagnosis made by radiologists.
One well-known public education message to identify the symptoms of a stroke is F.A.S.T.14 F stands for face; A stands for Arm; S stands for speech; and T stands for Time to call. Stroke patients will feel the face numbness of drag, and their arms will become weak; speech may also be slurred. When these symptoms arrive during sleep, a patient must try to recall their last time of wellness when they woke up from sleep, so doctors can understand when the stroke happened and calculate their onset time15. The faster one can get treated, the better the probability of a positive outcome. Even a 2-hour delay can be a huge determinant in the patient’s health. The MRI and CT scans need to make sure the stroke isn’t hemorrhagic, or else the drug can lead to the death of patients.
Embolic strokes start as small blood clots that move around the bloodstream. These clots start an ischemic stroke in a certain area by blocking arteries, as the clot moves to other areas closer to the brain. To remove the clots, neurosurgeons dissolve them by using medications such as alteplase and Tenecteplase in a surgical method called thrombotic therapy13,16. The people who have the biggest clots are the most in need of mechanical thrombectomy, which is an endovascular technique that removes clots throughout areas using a device that physically removes the clot. Patients with symptoms of stroke should have immediate attention to neuro images with non-contrast computed tomography or magnetic resonance imaging. Although this method is precise, the model is limited by the experiences and availability of staff and the infrastructure of hospitals.
As per existing research a DL model with high sensitivity and fewer false positives can greatly reduce the workload of radiologists17,18. This novice model focuses on achieving high sensitivity. Also, recent studies further demonstrated that deep learning-based assistance systems not only improve diagnostic accuracy, but also significantly reduce reading times19.
Methods
This is an experimental research to include multiple modalities to diagnose and determine accurate treatment for stroke patient. In this model, 1878 brain CT scan images of ischemic and hemorrhagic strokes were used for this research. This machine learning model is trained to identify if the stroke the patient is suffering from is Ischemic or hemorrhagic, so that doctors can understand what treatment route to follow. The two datasets that were used were selected because of their public availability and open source medical relevance, along with image quality. In Fig 1, this paper proposes a Proof of Concept to use multimodalities to decide the treatment path.
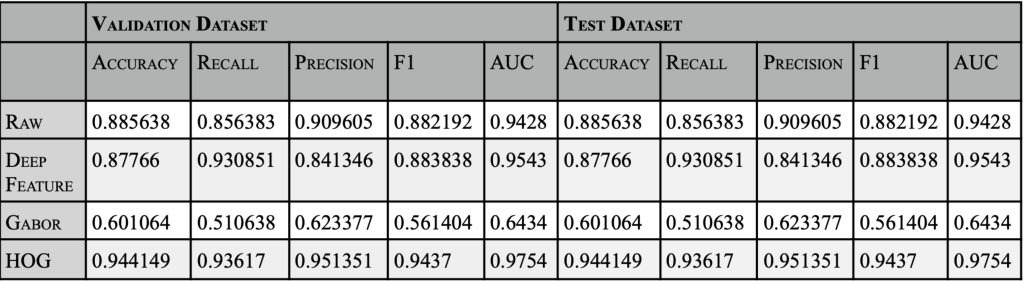
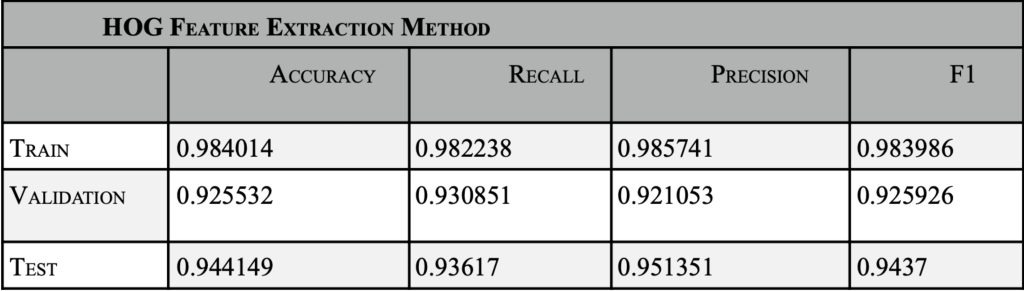
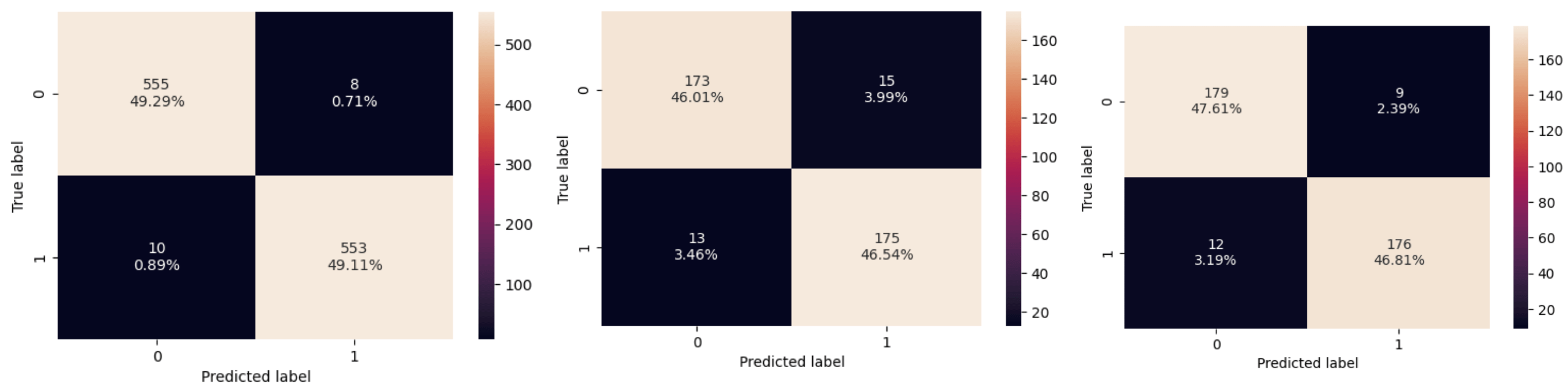
Dataset 1 uses SVM (Support Vector Machine)20 to classify patients into either having an ischemic or hemorrhagic stroke. The training database CT-scan-image-data set, found on Kaggle, was uploaded by NOSHIN TASNIA21. There are an equal number of ischemic and hemorrhagic images to avoid bias in training. Then data is split up into train/test/validation, 60% of the 1878 (1126) ischemic and hemorrhagic images were sent to train, 20% (376) images were sent to test, and 20% (376) images were sent to validation for both ischemic and hemorrhagic. The model preprocessed the image acquisitions to 256 by 256 pixels, to form all images to the same size. SVM is a supervised learning method used for classification, regression, and outlier detection. The model was trained using 4 different feature extraction methods – raw images, deep features, Gabor, and HOG (Histogram of Gradient Radiance). Hyperparameter tuning was performed by using GridsearchCV, with cv=5, param_grid = { ‘C’: [0.1, 1, 10, 100], ‘gamma’: [1, 0.1, 0.01, 0.001], ‘kernel’: [‘linear’, ‘rbf’, ‘poly’, ‘sigmoid’]}. The best hyperparameter for RAW was identified as {‘C’: 10, ‘gamma’: 0.001, ‘kernel’: ‘rbf’}, which produced a validation dataset accuracy of 90.96%. For Deep feature with cv=5 best parameters: {‘C’: 10, ‘gamma’: 0.001, ‘kernel’: ‘rbf’} with validation dataset accuracy of 83.51%.Using the Gabor features extraction yielded a validation dataset accuracy of 0.606383. HOG feature extraction resulted in a validation dataset accuracy of 92.55%. For Data Leakage is prevented before HOG feature extraction using a pipeline = Pipeline([(“scaler”, StandardScaler()),(“pca”, PCA(whiten=True)),(“svm”, SVC(probability=True)) ]). The proposed model performs feature normalization and PCA-based dimensionality reduction. Raw image classification used 65536 features, but when feature extraction methods were applied, the model used only relevant features. HOG feature extraction used 50% less features (34596) than raw image training.
Dataset-2: This is a proof of concept to determine DWI-FLAIR mismatch22. A total of 250 images were used with a split of 60% images for training, 20% for validation, and 20% images for testing. Here model uses a dual stream architecture to extract features from both the MRI modalities of DWI (Diffusion Weighted Imaging) & FLAIR ( Fluid Attenuated Inversion Recovery). As the MRI images are single-channel, we need to modify the images to 3-channel RGB images that can be processed by ResNet-18. Each ResNet outputs a 512 dimension feature-vector. The model learns both modalities together, which is important to identify DWI-FLAIR mismatch. These 2 vectors are fed to the various layers in the model. The Linear layer makes the model learn from the most meaningful 250 patterns. The ReLU layer adds non-linearity. To avoid overfitting, a dropout layer is introduced to disable 50% of neurons randomly. The final linear layer classifies into 2 classes: if there is a mismatch (infarct found) or no-mismatch(no-infarct). The model was trained using cross-entropy loss and optimized with the Adam optimizer at a learning rate of 1×10-4 with a batch size of 32 and 4 epochs. Training was performed using shuffled mini-batches, and GPU hardware was used for acceleration when available. The source of the second dataset is the Ischemic Stroke Lesion Segmentation ISLES challenge 202223. Since the datasets used were publicly available, there was no ethical approval or patient consent that was required.

Results

Discussion
This study emphasizes the importance of multiple modalities in arriving at an accurate treatment. Different modalities are used at different stages of analysis. The CT scans are enough to identify if the stroke is hemorrhagic or ischemic. The model in this study has compared the results of image analysis using just raw images and compared them to using different feature extraction methods, viz., deep learning, gabor and HOG. The HOG feature extraction and F1 scores used in this model are on-par with state-of-the art human radiologist predictions24. Based on the results in Fig. 2, the PCA( Principal Component Analysis) SVM is more memory efficient and avoids storing large numbers of training parameters, and when applied with HOG feature extraction, yields a F1 score of 94%. HOG feature extraction used 50% less features than raw image training, hence saving memory. The use of such Machine Learning models will aid the radiologist during high peak hours and night shifts25, when awaiting a decision from one radiologist, which causes a delay in treatments. On average, the radiologist may take 10-20 mins to conclude to classify haemorrhagic stroke patients, while this model may perform it below 2 min26. One of the limitations of CT scan images can be asymmetrical CT brain images. Asymmetrical CT can result in improper head positioning in the CT gantry, which can compromise the diagnostic value.27
The main motive of this study is to apply different machine learning algorithms in a single case of stroke classification and treatment. The model proves why different modalities are needed for a single stroke case. For haemorrhagic classification, CT is sufficient. Once the Ischemic Stroke is predicted, the next step would be to find the Stroke Onset Time. The CT scans in dataset1 only capture structural changes like tissue density, but cannot identify density changes over time. Hence, MRI images are used to identify the stroke onset time. As seen in dataset1, SVM performs better on non-linear CT scans, but stroke onset time needs progressive ischemic changes on non-linear spatial data that changes by time. The ResNet-18 model on dataset2 performs a classification on non-linear data to identify DWI-FLAIR mismatch. The input to this CNN model16 is an MRI image. The MRI images are not flattened and have high spatial data. The high number of features can be automatically learned by a CNN-based ResNet-18 model. While an SVM classification on a CT scan required a feature extraction method like HOG (Fig. 6) to be used before classifying. For identifying the stroke onset time in dataset-2, based on Fig. 7, the model achieved an accuracy of 96% on test data. The DWI-FLAIR mismatch implies that the stroke onset time occurred before 4.5 hours of arrival at the hospital. The patient, if treated within 4.5 hours, has a better chance of survival and fewer disabilities28. The model implemented detects the mismatch and change between the DWI and FLAIR imaging, which shows that the stroke occurred between the onset time.
The novice model proposed a pathway, as shown in Fig.1, to accurately treat promptly. These findings highlight the accuracy and efficiency that machine learning can offer to aid radiologists in treating stroke patients, which directly impacts patient survival rates. The study strongly suggests that the goal of designing a computational method must include CT and multi-modal MRI imaging in identifying stroke onset time and determining treatment type. The algorithm leads to aiding decision-making for a patient to receive Thrombectomy, alteplase tPA28, or an alternate method29. The results shared are performed on publicly available datasets and not on real patient data. The future of this research will focus on expanding the dataset with more perfusion CT scans30,31,32 to improve mapping the lesions of infarct and penumbra more accurately, since they are the critical areas in which the stroke is prominent in the brain. A key limitation of this study is the computation. Currently, the limited GPU, RAM, and memory cause a delay in the output of results, which can be improved with higher-end infrastructure. The model took ~3-4 hours to train. But once trained, the trained SVM model takes 0.8 sec inference time, while the ResNet-18 takes 1.8 sec inference time to test 1 unseen image. The trained model gives results under 2 sec. The actual time to train is proportional to the image data and the hardware. There is scope in implementing advanced deep learning architectures such as ResNet-50, U-Net, or EffeinetNet, one could further improve the segmentation precision and have a reduction in processing time.33 Hence, this project can be planned to be implemented on larger real datasets, revamped hardware, and alternative and refined CNN algorithms.
References
- V. L. Feigin et al., “Global, regional, and national burden of stroke and its risk factors, 1990–2019,” The Lancet Neurology, vol. 20, no. 10, pp. 795–820, 2021. doi: 10.1016/S1474-4422(21)00252-0. [↩]
- Y. Wang et al., “Traditional and machine learning models for predicting haemorrhagic transformation in ischaemic stroke: A systematic review and meta-analysis,” Systematic Reviews, vol. 14, no. 1, p. 46, 2025. doi: 10.1186/s13643-025-02771-w. [↩]
- V. L. Feigin, M. Brainin, B. Norrving, S. O. Martins, J. Pandian, P. Lindsay, M. F. Grupper, and I. Rautalin, “World Stroke Organization: Global stroke fact sheet 2025,” International Journal of Stroke, vol. 20, pp. 132–144, 2025. [↩]
- S. S. Virani et al., “Heart disease and stroke statistics—2023 update: A report from the American Heart Association,” Circulation, vol. 147, pp. e93–e621, 2023. [↩]
- C. J. van Asch et al., “Incidence, case fatality, and functional outcome of intracerebral haemorrhage over time,” The Lancet Neurology, vol. 9, no. 2, pp. 167–176, 2010. doi: 10.1016/S1474-4422(09)70340-0 [↩] [↩] [↩]
- B. Wang, B. Jiang, D. Liu, and R. Zhu, “Early predictive accuracy of machine learning for hemorrhagic transformation in acute ischemic stroke: Systematic review and meta-analysis,” Journal of Medical Internet Research, vol. 27, e71654, 2025. doi: 10.2196/71654. [↩]
- J. J. Nukovic et al., “Neuroimaging modalities used for ischemic stroke diagnosis and monitoring,” Medicina, vol. 59, no. 11, p. 1908, 2023. doi: 10.3390/medicina59111908. [↩]
- J. L. Saver, “Time is brain—quantified,” Stroke, vol. 37, no. 1, pp. 263–266, 2006. doi: 10.1161/01.str.0000196957.55928.ab [↩]
- G. Martinez, J. M. Katz, A. Pandya, J. J. Wang, A. Boltyenkov, A. Malhotra, A. I. Mushlin, and P. C. Sanelli, “Cost-effectiveness study of initial imaging selection in acute ischemic stroke care,” Journal of the American College of Radiology, vol. 18, no. 6, pp. 820–833, 2021. doi: 10.1016/j.jacr.2020.12.013. [↩]
- R. Smith-Bindman et al., “Use of diagnostic imaging studies and associated radiation exposure for patients enrolled in large integrated health care systems,” JAMA, vol. 307, no. 22, pp. 2400–2409, 2012. [↩]
- N. G. Ferrone et al., “Ten-year trends in last known well to arrival time in acute ischemic stroke patients: 2014–2023,” Stroke, vol. 56, pp. 591–602, 2025. [↩]
- H. Chen, S. Chaturvedi, D. Gandhi, and M. Colasurdo, “Stroke thrombectomy for large infarcts with limited penumbra: Systematic review and meta-analysis of randomized trials,” American Journal of Neuroradiology, vol. 46, pp. 915–920, 2025. [↩]
- N. Logallo et al., “Tenecteplase versus alteplase for management of acute ischemic stroke (NOR-TEST): A randomized controlled trial,” The Lancet Neurology, vol. 16, no. 10, pp. 781–788, 2017. [↩] [↩]
- D. O. Kleindorfer et al., “2021 guideline for the prevention of stroke in patients with stroke and transient ischemic attack,” Stroke, vol. 52, no. 7, pp. e364–e467, 2021. [↩]
- W. J. Powers et al., “2019 guidelines for the early management of patients with acute ischemic stroke,” Stroke, vol. 50, no. 12, pp. e344–e418, 2019 [↩]
- E. M. Z. Akay et al., “A deep learning analysis of stroke onset time prediction and comparison to DWI-FLAIR mismatch,” NeuroImage: Clinical, vol. 40, 103544, 2023. doi: 10.1016/j.nicl.2023.103544. [↩] [↩]
- A. Karamian and A. Seifi, “Diagnostic accuracy of deep learning for intracranial hemorrhage detection in non-contrast brain CT scans: A systematic review and meta-analysis,” Journal of Clinical Medicine, vol. 14, no. 7, p. 2377, 2025. doi: 10.3390/jcm14072377. [↩]
- K. Villringer et al., “An artificial intelligence algorithm integrated into the clinical workflow can ensure high quality acute intracranial hemorrhage CT diagnostic,” Clinical Neuroradiology, vol. 35, no. 1, pp. 115–122, 2025. doi: 10.1007/s00062-024-01461-9. [↩]
- D. W. Kang et al., “Deep learning-assisted detection of intracranial hemorrhage: Validation and impact on reader performance,” Neuroradiology, vol. 67, pp. 1511–1519, 2025. doi: 10.1007/s00234-025-03560-x [↩]
- C. Cortes and V. Vapnik, “Support-vector networks,” Machine Learning, vol. 20, no. 3, pp. 273–297, 1995. [↩]
- N. Tasnia, “Brain Stroke Prediction CT Scan Image Dataset,” Kaggle, 2023. [Online]. Available: https://www.kaggle.com/datasets/noshintasnia/brain-stroke-prediction-ct-scan-image-dataset. Accessed: Mar. 10, 2026. [↩]
- Y. Liu et al., “Artificial intelligence in ischemic stroke images: Current applications and future directions,” Frontiers in Neurology, vol. 15, 1418060, 2024. doi: 10.3389/fneur.2024.1418060. [↩]
- M. R. Hernandez Petzsche et al., “ISLES 2022: A multi-center magnetic resonance imaging stroke lesion segmentation dataset,” Zenodo, 2022. doi: 10.5281/zenodo.7153326. [↩]
- A. Al-Salahat, Y. Pirahanchi, T. Dhasakeerthi, et al., “Current state and advancements of imaging in acute ischemic stroke: A practical review,” Neurological Sciences, vol. 47, p. 29, 2026. doi: 10.1007/s10072-025-08685-8. [↩]
- T. D’Angelo et al., “Accuracy and time efficiency of a novel deep learning algorithm for intracranial hemorrhage detection in CT scans,” La Radiologia Medica, vol. 129, pp. 1499–1506, 2024. [↩]
- A. N. Khoruzhaya et al., “Standalone AI versus AI-assisted radiologists in emergency ICH detection: A prospective multicenter diagnostic accuracy study,” Journal of Clinical Medicine, vol. 14, no. 16, p. 5700, 2025. doi: 10.3390/jcm14165700 [↩]
- J. J. Downer and P. M. Pretorius, “Symmetry in computed tomography of the brain: The pitfalls,” Clinical Radiology, vol. 64, no. 3, pp. 298–306, 2009. doi: 10.1016/j.crad.2008.08.012. [↩]
- G. Thomalla et al., “Intravenous alteplase for stroke with unknown time of onset guided by advanced imaging: Systematic review and meta-analysis,” The Lancet, vol. 396, no. 10262, pp. 1574–1584, 2020. doi: 10.1016/S0140-6736(20)32163-2. [↩] [↩]
- B. C. V. Campbell and P. Khatri, “Stroke,” The Lancet, vol. 396, no. 10244, pp. 129–142, 2020. doi: 10.1016/S0140-6736(20)31179-X. [↩]
- M. Koneru et al., “Early experience with artificial intelligence software to detect intracranial occlusive stroke in trauma patients,” Cureus, 2024. doi: 10.7759/cureus.57084. [↩]
- S. A. Sheth et al., “Machine learning–enabled automated determination of acute ischemic core from computed tomography angiography,” Stroke, vol. 50, no. 11, 2019. doi: 10.1161/STROKEAHA.119.026189. [↩]
- K. C. Ho, W. Speier, S. El-Saden, and C. W. Arnold, “Classifying acute ischemic stroke onset time using deep imaging features,” AMIA Annual Symposium Proceedings, pp. 892–901, 2018. [↩]
- Y. Wang et al., “Patterns and clinical implications of hemorrhagic transformation after thrombolysis in acute ischemic stroke,” Neurology, vol. 103, no. 11, 2024. [↩]