Abstract
Hereditary spherocytosis (HS) is a genetically inherited disease affecting red blood cells. Due to the genetic component that causes the disease, common treatments focus on immediate life improvements for those afflicted with HS such as blood transfusions, splenectomy, and phototherapy. Studies have revealed that patients with HS exhibit protein mutations associated with five genes: Ankyrin-1 (ANK1, Type 1), spectrin ![]() erythrocytic (SPTB, Type 2), spectrin
erythrocytic (SPTB, Type 2), spectrin ![]() erythrocytic 1 (SPTA1, Type 3), band 3 with monoclonal antibody (AE1, Type 4), and erythrocyte membrane protein-band 4.2 (EPB42, Type 5). Because mutations in the ANK1 and SPTB genes are most common, this investigation considers the prevalence and classification of Type 1 and Type 2 variants that result in an HS diagnosis. This study provides further evidence that variants in both genes result in a functional change to the structure of red blood cells that correlates to the disease’s presence by analyzing gene mutations likely to cause disease against those less likely to be pathogenic for Type 1 and Type 2. Finally, this research analyzes the frequency of observed ANK1 versus SPTB mutations, the frequency of mutation types, and the likelihood that each mutation on a given gene results in a pathogenic outcome. Predicting the precise mutation location across the protein sequence is difficult as the analysis demonstrates that genes can mutate along the entire protein length. The pathogenic mechanism is unclear, and further studies in this area will contribute to a more accurate HS diagnosis in the future.
erythrocytic 1 (SPTA1, Type 3), band 3 with monoclonal antibody (AE1, Type 4), and erythrocyte membrane protein-band 4.2 (EPB42, Type 5). Because mutations in the ANK1 and SPTB genes are most common, this investigation considers the prevalence and classification of Type 1 and Type 2 variants that result in an HS diagnosis. This study provides further evidence that variants in both genes result in a functional change to the structure of red blood cells that correlates to the disease’s presence by analyzing gene mutations likely to cause disease against those less likely to be pathogenic for Type 1 and Type 2. Finally, this research analyzes the frequency of observed ANK1 versus SPTB mutations, the frequency of mutation types, and the likelihood that each mutation on a given gene results in a pathogenic outcome. Predicting the precise mutation location across the protein sequence is difficult as the analysis demonstrates that genes can mutate along the entire protein length. The pathogenic mechanism is unclear, and further studies in this area will contribute to a more accurate HS diagnosis in the future.
Introduction
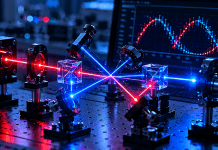
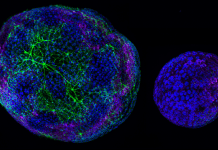
HS is a genetic condition associated with red blood cells (RBCs). Patients living with HS often feel fatigued and cannot maintain strenuous cardiac activity for prolonged periods because their red blood cells are not as efficient in transporting oxygen throughout the body. The spleen identifies the spherical RBCs as defective and filters them from the bloodstream, leading to anemia. Because HS is a genetic condition, patients must adapt their lifestyle to remain healthy. HS is diagnosed in one out of every two thousand individuals. However, the number of individuals with HS is likely far greater than those counted as many more asymptomatic cases likely remain undiagnosed1.While RBCs typically exhibit a rounded disk shape, as displayed in Figure 1, HS cells exhibit a spherical morphology, as shown by the arrow in the same picture, which causes the body to perceive the red blood cell as damaged and triggers the spleen to filter them out of the bloodstream2.

Because the spleen must filter large quantities of RBCs, the patient can develop an enlarged spleen over time. HS also results in fragile RBCs that burst more easily. RBCs typically distribute oxygen throughout the body, with HS patients often experiencing a reduction in the blood’s oxygen-carrying capacity, leading to symptoms such as anemia and fatigue3.
HS is often diagnosed through blood tests and visually through microscopic investigation. Treatment options vary depending on the patient’s HS severity. Focus is often placed on managing and reducing symptoms in less severe cases. These treatments may include folate or Vitamin B9 supplements to stimulate blood cell formation, blood transfusions to increase healthy red blood cell and hemoglobin levels, or gallbladder removal, should the patient develop gallstones. In more severe cases, spleen removal (splenectomy) is recommended4.Figure 2 provides a red blood cell cytoskeleton model with a biconcave disc shape supporting the lipid bilayer. The enlarged region highlights the arrangement of proteins within this structure, particularly on the ANK1 and SPTB proteins that, when defective, cause HS5.

The round red blood cell shape attributed to HS has largely been correlated to mutations in five different genes. These genes (and associated proteins) can be categorized as Type 1: ANK1 (ankyrin), Type 2: SPTB (![]() -spectrin), Type 3: SPTA1 (
-spectrin), Type 3: SPTA1 (![]() -spectrin), Type 4: SLC4A1 (band-3 protein), and Type 5: EPB42 (protein 4.2). The disease-causing mutations result in gene function impairment, protein misfolding, or insufficient binding between the proteins at the cell membrane, preventing the proper membrane shape from forming6.
-spectrin), Type 4: SLC4A1 (band-3 protein), and Type 5: EPB42 (protein 4.2). The disease-causing mutations result in gene function impairment, protein misfolding, or insufficient binding between the proteins at the cell membrane, preventing the proper membrane shape from forming6.
Other studies demonstrate that HS results from defects of any protein components involved in the cell membrane structure and skeletal network vertical linkages. As shown in Figure 3, ANK1 (anchoring protein) interacts with essential membrane proteins, including SPTB and cytoskeletal proteins7.

Fifty to sixty percent of HS cases are estimated to be linked to ANK1 gene mutations, making it the most likely of the five to exhibit changes7. ANK1 mutations occur primarily with frameshift, nonsense, and splice site mutation types. Researchers have found that ANK1 mutations are almost always autosomal dominant when investigating all HS patients, but autosomal recessive inheritance can occur. ANK1 mutations can be unique to an individual, but common mutation sites in a family can be inherited. Some research has shown that HS can be induced by the haploinsufficiency of ANK1, demonstrating a complex pathogenic relationship between ANK1 and HS6.
The second most common gene tied to HS mutations is SPTB, representing up to twenty percent of the patient population8. SPTB plays a different but equally important role to ANK1 in ensuring erythrocyte membrane stability. SPTB deficiency is commonly inherited in an autosomal dominant manner. This research demonstrates that heterozygous SPTB mutations represent about twenty-five percent with common mutation types, including splice site, frameshift, and nonsense mutations, often leading to defects in mRNA processing and truncated SPTB6.Although other studies demonstrate that specific genetic mutations are associated with an HS diagnosis, less investigation is focused on the genetic mutations that result in protein structural changes and the associated functional relationship as it applies to HS. This relationship includes the type of disease-causing mutations and the affected protein structures involved. This paper focuses primarily on the ANK1 and SPTB genes and the effect of mutations within each related protein structure, which results in an HS diagnosis. The ANK1 and SPTB genes are chosen because relatively few publications have investigated their involvement in HS and because the combination of ANK1 and SPTB mutations is estimated to comprise at least seventy-five percent of the genetic mutations associated with HS. Bioinformatics methodologies and publicly available data sources are used to determine how mutations in the protein-coding sequences of the genes can be related to HS and its exhibited phenotypes.
Results
To conceptualize the Ankyrin-1 and ![]() -Spectrin protein structures, refer to Figures 4 and 5, obtained using the UniProt tool9.
-Spectrin protein structures, refer to Figures 4 and 5, obtained using the UniProt tool9.


 -spectrin protein structure10.
-spectrin protein structure10.Using the ClinVar database11, gene mutations on ANK1 and SPTB are extracted and categorized using The American College of Medical Genetics and Genomics (ACMG) guidelines: (i) pathogenic, (ii) likely pathogenic, (iii) uncertain significance, (iv) likely benign, or (v) benign12. Clinically, both “pathogenic” and “likely pathogenic” variants are treated equally because they are likely to cause disease13, and, for this analysis, these two categories are compared against the other three categories.
As summarized in Figure 6, more than twice as many total mutations linked to HS have been found in ANK1 (244) compared to SPTB (92). This implies that someone with an HS diagnosis is more likely to have an ANK1 genetic mutation than an SPTB one.

genetic mutations of the cases analyzed within the ClinVar database as categorized by ACMG category guidelines: (i) pathogenic, (ii) likely pathogenic, (iii) uncertain significance, (iv) likely benign, or (v) benign12. Clinically, both “pathogenic” and “likely pathogenic” variants are treated equally because they are likely to cause disease13, and, for this analysis, these two categories are compared against the other three categories. The difference between pathogenic and likely pathogenic is the amount of data that has linked the variant to the manifestation on HS. Because there is no quantitative definition of the term ‘likely,’ the AMCG proposed that ‘likely pathogenic’ and ‘likely benign’ apply to 90% certainty of a variant either being disease-causing or benign to provide a standard definition12.
Figure 6 also displays the percentage of each mutation classification for a given gene. Only about half (49%) of Type 1 are categorized as pathogenic or likely pathogenic, but almost three-quarters (73%) of the Type 2 mutations have this classification. In summary, once an HS patient acquires an SPTB mutation, that gene is more likely to be pathogenic than with an ANK1 mutation.
Given the total number of ANK1 mutations (244 cases) is more than double those associated with SPTB (92 cases), the total pathogenic and likely pathogenic values are also about twice in quantity for ANK1 (120) versus SPTB (67), as displayed in Figure 6.
In Figure 7, the type of variant for each genetic mutation is considered. Again, ANK1 mutations represent more than two and a half times the number of SPTB mutations.

genetic variants of the cases within the ClinVar database14. Note: when the variant type in the ClinVar database is recorded as Insertion (1 case in HS Type 1) or N/A (2 cases in HS Type 1 and 1 case in HS Type 2), the data is excluded from the chart.
Interestingly, the ANK1 and SPTB mutations are similar in percentage when viewed by variant types, with over three-quarters representing substitution errors and five percent or less representing duplication. This is illustrated in Figure 7.
When one considers the location of the ANK1 and SPTB mutations along the protein sequence, mutations seem to be distributed evenly, with no significant clusters observed in any region. Figures 18 and 9 represent each mutation’s “pathogenic” and “likely pathogenic” placement for Types 1 and 2, respectively.


While no location along the ANK1 or the SPTB proteins appears to be significant in the mutations that result in an HS diagnosis, the results do demonstrate that ANK1 mutations are more common than SPTB mutations, aligned with previous reported results. For ANK1 mutations, only about half of the mutations are likely to result in an HS diagnosis, but almost three quarters of the SPTB mutations result in the disease. Because the ANK1 mutations are more commonly observed than the SPTB ones, a patient with HS is more likely to exhibit a pathogenic or likely pathogenic mutation on the ANK1 gene than the SPTB one.
Discussion
HS is not a well-known disease compared to other inherited blood conditions, resulting in minimal research focus. Worldwide, this disease is considered rare, and a relatively small population inherits this disease. This opens a tremendous opportunity for scientists willing to focus on this disease because additional understanding could benefit patients suffering from this affliction. Specifically, future experiments could target several areas to gain a deeper understanding of HS. Research opportunities include investigating the relationship between protein structure and the other three types of mutation, developing a kind of medicine or treatment for HS patients, and analyzing how one mutation in the protein structure can affect a different kind of gene. This paper considers the effects of ANK1 and SPTB mutations, but future analysis of SPTA1, AE1 and EPB42 mutations might reveal relationships not observed on ANK1 and SPTB genes. This additional investigation could reveal a deeper understanding of mutations and underlying causes of HS that are not observed when focus is placed on ANK1 and SPTB mutations only.
As stated in the Results, this investigation demonstrates that ANK1 genes exhibit mutations more than twice as often as SPTB ones. Scientists focusing on HS Type 1 and associated mutations might target their research outcomes to benefit most HS patients quickly. Because ANK1 acts as an anchoring protein, this region might play a significant role in the development of HS. Because ANK1 interacts with SPTB and other cytoskeletal proteins, the research also supports the conclusion that SPTB mutations also lead to HS symptoms.
Researchers might take an alternate approach and, rather than targeting ANK1, instead target SPTB mutations. This could be a logical choice if the experiment focuses specifically on “pathogenic” or “likely pathogenic” categories since a higher percentage of Type 2 gene mutations exhibit these classifications. For a given SPTB mutation, there is a higher likelihood that it will result in a pathogenic response than for an ANK1 mutation.
Another area for researchers to consider is investigating whether other red blood cell disease management strategies might be applicable in treatments or cures for HS too. In this case, researchers might consider the underlying causes of diseases that result in a cell membrane structural change and whether a similar treatment methodology might be applied to HS. For example, sickle cell anemia results in misshapen red blood cells. An experiment to see whether there might be any similarities between the diseases where a known treatment in one disease might benefit patients suffering from the other.
Spending too much time investigating specific locations within the ANK1 and SPTB genes to identify “pathogenic” or “likely pathogenic” mutations is not as likely to provide answers as the analysis did not reveal any localized clusters. As a result, there are no targeted locations for scientists to focus their studies along the length of either protein. Since five known genes mutate to result in an HS diagnosis, the other three could be evaluated similarly to determine whether any other localized correlations might exist.
Because the data is extracted from the ClinVar database11, any bias or errors in the database’s submitted content would impact this study’s conclusions’ accuracy. The ClinVar database allows global information to be freely submitted from clinical testing labs, researchers, database curators, and expert panels. While this incorporates a variety of worldwide sources, the results could be subject to errors and omissions based on the bias of those individuals sharing the results. Future studies could obtain data from other sources to compare those with the findings presented here. Specific populations of HS patients may also exhibit variation in the most prevalent genetic mutations, which would not be identified in this research since all data is combined regardless of ethnicity, age, sex, and other characteristics.
Scientists should continue to pursue an understanding of genetic mutations and their effect on HS. Once these are more thoroughly understood, targeted treatments might become available for HS that are not available currently. In the meantime, most patients can live relatively normal lives with HS with minor lifestyle accommodations. In the future, severe HS cases might be best treated through targeted genetic treatments that will only become available through more research and clinical trials.
Methods
First, the UniProt tool9 is used to generate the Ankyrin 1 and Spectrin protein structures. The following steps are used to obtain the structure of these proteins:
- Enter the gene name into the search box at the initial landing web page9.
- Refine search criteria (see left panel) by selecting a relevant organism, e.g., ‘Human.’
- Select from the entry display list for the protein of interest, e.g., ankyrin-1 or
 -Spectrin.
-Spectrin. - Select a further search parameter on the left of the page, e.g., ‘Structure’ to find the 3D image of the protein.
This investigation next gathers information about HS and previous research using Google Scholar and other databases. Next, Uniprot9 and ClinVar11 are accessed; both are considered well-respected data repositories for gene variant submissions of experimental results. ClinVar is a freely available archive for interpretations of the clinical significance of variants for reported conditions. Submitters to ClinVar include clinical testing labs, researchers, database curators, and expert panels. ClinVar is increasingly becoming the central repository of interpreted genomic variants11.
The variant data is obtained using the ClinVar Data Miner web-based browsing tool14, selected for ease of interrogating the ClinVar data repository11. ClinVar Miner provides summaries of ClinVar data that can be sorted using built-in filters. The ‘Variants by gene’ filter option is initially chosen at the ClinVar Data Miner website. ‘Variants by gene’ displays the variant classification of submitted genes and enables the sorting of genes by the number of variants per gene. A filter is included to discriminate between variants within or near a single gene. For the purposes of this research paper, only the condition with ‘Hereditary spherocytosis Type 1’ and ‘Hereditary spherocytosis Type 2’ are chosen to avoid any confusion with ClinVar submissions that do not use consistent naming conventions.
Once selected, a further filter option is automatically presented to refine the specific gene, i.e., ANK1 and SPTB. Selecting the gene name’s hyperlink reveals a summary page highlighting the condition (disease) and the significance of the variant submitted to the ClinVar database. The ‘HS Type 1’ is chosen for ANKI, and ‘HS Type 2’ is selected for SPTB. Each condition shows a breakdown of variants by significance level (e.g., pathogenic or benign). The list of variants for each significance level is displayed by choosing the hyperlinked numerical summary total value. These are then downloaded in the.csv file format. The process is repeated for each significance level and consolidated into a single table, as indicated in Appendix 1, for further analysis.
Once the ANK1 and SBPN Genetic Variants are extracted from the ClinVar database, each is categorized into HS Type 1 (ANK1) or HS Type 2 (SPTB). The category is determined from the database where the information is extracted. To populate the “Variant Type” column, the following categories are determined from the “Variant Name (HGVS)” column:
- Substitution is selected when one letter (nucleotide) of the DNA code is replaced (substituted) by one other letter. A substitution is indicated using the ”>” symbol. For example, “c.4375C>T” reflects that the C nucleotide at position c.4375 changed to a T15.
- Deletion occurs when one or more letters of the DNA code are missing (deleted). A deletion is indicated using “del.” For example, the notation “c.4375_4379del” means that the nucleotides from position c.4375 to c.4379 (CGATT) are missing (deleted). They are also documented as c.4375_4379delCGATT15.
- Duplication is chosen when one or more letters of the DNA code are present twice (doubled, duplicated). A duplication is indicated using “dup.” As an example, “c.4375_4385dup” means that the nucleotides from position c.4375 to c.4385 (CGATTATTCCA) are present twice (duplicated)15.
- Insertion occurs when one or more letters in the DNA code are new (inserted). An insertion is indicated using “ins” An example is “c.4375_4376insACCT,” which reflects that the new sequence “ACCT” is found inserted between positions c.4375 and c.437615.
- Finally, deletion/insertion (indel) is used when one or more letters in the DNA code are missing and replaced by several new letters. A deletion/insertion is indicated using “delins.” In the case of “c.4375_4376delinsAGTT,” the nucleotides from position c.4375 to c.4376 (CG) are missing (deleted) and replaced by the new sequence “AGTT”15.
Once the data (by condition and significance) is collected, it is then transferred into Microsoft Word and Excel spreadsheets to generate stacked bar plots, pie charts, and tables.
Acknowledgements
Thank you for the guidance of my mentor, Mr. Christopher Beaudoin from the University of Cambridge and Ms. Stephanie Leow and Mr. Tyler Moulton of Empowerly in developing this research paper.
Appendix
ClinVar raw data is categorized for ANK1 and SPTB by significance and variant category14.
| Gene | Condition | Significance | Variant Name (HGVS) | Variant Type |
|---|---|---|---|---|
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4105A>G (p. Lys1369Glu) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1024C>T (p. Gln342Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1331_1338del (p. Leu444fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.151G>T (p. Glu51Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1591C>T (p. Gln531Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1630dup (p. Met544fs) | duplication |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1912del (p. Arg638fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.1A>G (p. Met1Val) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2092del (p. Gln698fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2165C>A (p. Ser722Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2278C>T (p. Gln760Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2423del (p. Gly808fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2521C>T (p. Gln841Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2647del (p. Leu883fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2659C>T (p. Gln887Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2749_2750del (p. Ser917fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.2863C>T (p. Arg955Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3106dup (p. Gln1036fs) | duplication |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3351C>A (p. Tyr1117Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3841C>T (p. Gln1281Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3855+1G>A | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3936G>A (p. Trp1312Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.3986_4001del (p. Leu1329fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4267C>T (p. Arg1423Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB):c.4368del (p.Ile1456fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4417C>T (p. Gln1473Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.467G>C (p. Arg156Pro) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.472C>T (p. Gln158Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4842+1G>C | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4873C>T (p. Arg1625Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.493C>T (p. Gln165Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4942C>T (p. Gln1648Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.4969G>T (p. Glu1657Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5059dup (p. Glu1687fs) | duplication |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5099del (p. Asp1700fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5114G>A (p. Trp1705Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5266C>T (p. Arg1756Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5464G>T (p. Glu1822Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5623C>T (p. Gln1875Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5737dup (p. Arg1913fs) | duplication |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.5788G>T (p. Glu1930Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.604T>C (p. Trp202Arg) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.6059_6060del (p. Val2020fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.6536_6537del (p. Val2179fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.85G>T (p. Glu29Ter) | substitution |
| SPTB | HS Type 2 | Pathogenic | NM_001355436.2(SPTB): c.999_1000del (p. Leu334fs) | deletion |
| SPTB | HS Type 2 | Pathogenic | SPTB, EX22-23DEL | deletion |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.1182+2T>C | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.1465G>T (p. Glu489Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.146C>T (p. Ala49Val) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.1795+1G>A | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.1908del (p. Lys637fs) | deletion |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.2863C>T (p. Arg955Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.3082C>T (p. Gln1028Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.3784_3787del (p. Lys1262fs) | deletion |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.3877A>T (p. Lys1293Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.3916C>T (p. Arg1306Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.4178del (p. Lys1393fs) | deletion |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.474+1G>A | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.4973+5G>A | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.5178+1G>A | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.5446A>T (p. Lys1816Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.5455G>T (p. Glu1819Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.5953C>T (p. Gln1985Ter) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.6055T>C (p. Ser2019Pro) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.6119C>T (p. Thr2040Ile) | substitution |
| SPTB | HS Type 2 | Likely Pathogenic | NM_001355436.2(SPTB): c.647G>A (p. Arg216Gln) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.3679C>T (p. Pro1227Ser) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.2519G>A (p. Arg840His) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.774G>A (p. Thr258=) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.413T>C (p. Ile138Thr) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.5680G>A (p. Gly1894Arg) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.6074T>G (p. Leu2025Arg) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.754G>A (p. Asp252Asn) | substitution |
| SPTB | HS Type 2 | Uncertain | NM_001355436.2(SPTB): c.836A>C (p. Lys279Thr) | substitution |
| SPTB | HS Type 2 | Likely Benign | None | n/a |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.2154A>C (p. Ile718=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.1316G>A (p. Ser439Asn) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.300+7T>C | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4860T>C (p. Ile1620=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4293A>G (p. Arg1431=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.3451A>G (p. Asn1151Asp) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.876+5A>G | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4476T>C (p. Leu1492=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4482G>A (p. Val1494=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4641G>A (p. Ala1547=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.1269G>A (p. Leu423=) | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.300+23C>T | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4002+26T>C | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4474-42C>T | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.4563+12G>C | substitution |
| SPTB | HS Type 2 | Benign | NM_001355436.2(SPTB): c.648-49G>A | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4153C>T (p. Arg1385Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5497C>T (p. Arg1833Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1124T>G (p. Leu375Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1305+1G>A | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1365T>G (p. Tyr455Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1427_1430dup (p. Ala478fs) | duplication |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1519dup (p. Leu507fs) | duplication |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1602+1G>C | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1801-17G>A | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1891G>T (p. Glu631Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.1915del (p. Leu639fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2029C>T (p. Gln677Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2098-1G>T | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2102del (p. Gly701fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2164C>T (p. Gln722Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2389-1G>A | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2390_2393del | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2393_2403del (p. Val798fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2394_2397del (p. Ser799fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.2803C>T (p. Arg935Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3073G>T (p. Gly1025Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3092_3095del (p. Gln1031fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3157C>T (p. Arg1053Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.319C>T (p. Gln107Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3275del (p. Gln1092fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3325C>T (p. Gln1109Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3504_3514del (p. Ser1169fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3555G>A (p. Trp1185Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3629+1G>C | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3629+2T>C | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3639_3649dup (p. Pro1217fs) | duplication |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3754C>T (p. Arg1252Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.382_386del (p. Lys128fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3850del (p. Asp1284fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.3984+2T>C | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4000C>T (p. Arg1334Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4040_4041del (p. Met1347fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4057C>T (p. Gln1353Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4098C>A (p. Cys1366Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.409C>T (p. Gln137Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4140del (p. Leu1382fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4157dup (p. Tyr1386Ter) | duplication |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4204C>T (p. Gln1402Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4306C>T (p. Arg1436Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4387_4390del (p. Asn1463fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4541del (p. Tyr1514fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.4819_4820del (p. Ser1607fs) | deletion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5005G>T (p. Glu1669Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5044C>T (p. Arg1682Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5071C>T (p. Gln1691Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5097-33G>A | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5152C>T (p. Gln1718Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5163G>A (p. Trp1721Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5192C>G (p. Ser1731Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.5436_5437insCAGGG (p. Glu1813fs) | Insertion |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1): c.841C>T (p. Arg281Ter) | substitution |
| ANK1 | HS Type 1 | Pathogenic | NM_000037.4(ANK1):c.886del (p.Ala296fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1135del (p. Cys379fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.1381_1382del (p.Ala461fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1405-9G>A | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1452del (p. Asn484fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1488dup (p. Asn497fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1585C>T (p. Gln529Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1801-17G>A | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.181del (p. Val61fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1895C>A (p. Ser632Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1900C>T (p. Gln634Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1972C>T (p. Gln658Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.1A>G (p. Met1Val) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2004del (p. Leu669fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2023dup (p. Val675fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2104dup (p. Tyr702fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2283del (p. Asn761fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2296-2A>C | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2296-2A>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.2320_2350del (p.Ala774fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.2485del (p.Ala829fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2558+2T>C | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2632G>T (p. Glu878Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2768del (p. Gly923fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2823del (p. Thr942fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2960+1G>A | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.2961-2A>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3059_3066del (p. His1020fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3123del (p. Ser1042fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3123dup (p. Ser1042fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3196_3199dup (p. Ser1067fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3269del (p. Leu1090fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3325C>T (p. Gln1109Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3493_3496dup (p. Asp1166fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3533-2A>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3550C>T (p. Gln1184Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3623_3624del (p. Ser1208fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3630-1G>A | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3775del (p. Tyr1259fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3777C>G (p. Tyr1259Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3859-2A>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3893_3894del (p. Ser1298fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.3928C>T (p. Gln1310Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4051del (p. Asp1351fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.4092_4101del (p.Pro1365fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.409C>T (p. Gln137Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4184-2A>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4259-1G>T | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4267del (p. Arg1423fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4390+1G>A | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4771G>T (p. Glu1591Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4779_4780del (p. Asp1594fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.4813del (p.Ala1605fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4835del (p. Gly1612fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4855G>T (p. Glu1619Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.4886del (p. Asp1629fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.5026del (p. His1676fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.5076dup (p. Thr1693fs) | duplication |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.5096+2T>G | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.5108G>A (p. Trp1703Ter) | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.5323_5324del (p. Arg1775fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.712-2A>T | substitution |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1):c.725del (p.Pro242fs) | deletion |
| ANK1 | HS Type 1 | Likely Pathogenic | NM_000037.4(ANK1): c.86del (p. Leu29fs) | deletion |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3224C>T (p. Thr1075Ile) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3813G>A (p. Glu1271=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2211C>G (p. Pro737=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.654C>A (p. Asn218Lys) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*1708G>C | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5544+91C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3399C>T (p. Thr1133=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.981C>T (p. Tyr327=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2403G>A (p. Lys801=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2735+10G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1506C>T (p. Ala502=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5544+46G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5097-34C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*8C>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2830G>A (p. Ala944Thr) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2495G>A (p. Arg832Gln) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2745G>A (p. Val915=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*821C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5395-1162C>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.4606C>T (p. Arg1536Cys) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.997G>A (p. Asp333Asn) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1484A>G (p. Asn495Ser) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3532+13C>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2589C>T (p. Pro863=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2960+13G>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3102C>T (p. Asn1034=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2132A>G (p. Tyr711Cys) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*992T>C | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1153C>T (p. Arg385Cys) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1404+15C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2196+6G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.4022C>T (p. Ser1341Leu) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*1667T>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1483A>C (p. Asn495His) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*395T>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2167C>A (p. His723Asn) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1467G>A (p. Leu489=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3668T>C (p. Val1223Ala) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1203C>T (p. Thr401=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3570A>C (p. Gly1190=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2982G>A (p. Pro994=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1926C>T (p. Ala642=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2097+15C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.4136C>T (p. Pro1379Leu) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*857C>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1197G>A (p. Ala399=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5520C>T (p. Ala1840=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2370G>T (p. Thr790=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.*1892G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1305+21G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1602+18G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.1999-17C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2197-17G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2352C>T (p. Asp784=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.2461+20C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3115+9G>A | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3298G>A (p. Val1100Ile) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3342G>A (p. Pro1114=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3829G>A (p. Val1277Met) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.3984+10C>T | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.4538-6A>G | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.4974C>T (p. Asp1658=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5233G>A (p. Gly1745Ser) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5598G>A (p. Gly1866=) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_000037.4(ANK1): c.5600C>T (p. Ala1867Val) | substitution |
| ANK1 | HS Type 1 | Likely Benign | NM_001142446.2(ANK1): c.127-39556C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*2152T>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5096+16T>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5479-3T>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.315C>T (p. Asn105=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5265G>A (p. Val1755=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2349C>T (p. Thr783=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2073C>T (p. Gly691=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.4101C>T (p. Ala1367=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.1856G>A (p. Arg619His) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*238T>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.909+7A>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*1899G>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*1609C>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3486C>T (p. Ser1162=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_001142446.2(ANK1): c.127-39509T>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3973A>G (p. Met1325Val) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.1320G>A (p. Pro440=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5395-1147C>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.4385C>T (p. Ala1462Val) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*2022G>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3327+21C>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3224C>T (p. Thr1075Ile) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.4008G>A (p. Pro1336=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3813G>A (p. Glu1271=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2211C>G (p. Pro737=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.183G>C (p. Val61=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2296-20C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.654C>A (p. Asn218Lys) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*1402G>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2970C>T (p. Ile990=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5544+91C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.1590C>T (p. Ala530=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*774A>G | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3399C>T (p. Thr1133=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.981C>T (p. Tyr327=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2403G>A (p. Lys801=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3033C>T (p. Ser1011=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5097-34C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2495G>A (p. Arg832Gln) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5479-17T>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.5395-1162C>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3532+13C>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3102C>T (p. Asn1034=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.1467G>A (p. Leu489=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3668T>C (p. Val1223Ala) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.675C>T (p. Leu225=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2688C>T (p. Thr896=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.237C>T (p. Asn79=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*36+989dup | duplication |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.*385del | deletion |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.129+125C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.1999-17C>T | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.229-47G>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.2913G>C (p. Leu971=) | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.3115+9G>A | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.4538-52G>C | substitution |
| ANK1 | HS Type 1 | Benign | NM_000037.4(ANK1): c.909+36A>G | substitution |
| ANK1 | HS Type 1 | Uncertain | n/a |
- E. Zamora, C. A. Schaefer. Hereditary spherocytosis. National Library of Medicine: National Center for Biotechnology Information, https://pubmed.ncbi.nlm.nih.gov/30969619/ (2023). [↩]
- S. Crary. Hereditary spherocytosis. BMJ Best Practice, https://bestpractice.bmj.com/topics/en-us/1143 (2023). [↩] [↩]
- P. Mehta. What is hereditary spherocytosis? WebMD, www.webmd.com/children/what-is-hereditary-spherocytosis (2021). [↩]
- E. Westfall, C. A. Bellcross. Hereditary spherocytosis. NORD, rarediseases.org/rare-diseases/anemia-hereditary-spherocytic-hemolytic/ (2019). [↩]
- M. Risinger, T. A. Kalfa. Red cell membrane disorders: structure Meets function. Blood, 136, 1250-1261 (2020). [↩] [↩]
- B.-J. He, L. Liao, Z.-F. Deng, Y.-F. Tao, Y.-C. Xu, & F.-Q. Lin. Molecular genetic mechanisms of hereditary spherocytosis: current perspectives. Acta Haematologica, 139, 60-66 (2018). [↩] [↩] [↩]
- X. An, N. Mohandas. Disorders of red cell membrane. British Journal of Haematology, 141, 367-375 (2008). [↩] [↩] [↩]
- X. An, N. Mohandas. Disorders of the red cell membrane. British Journal of Haematology, 141, 367-375 (2008). [↩]
- The UniProt Consortium, UniProt: the universal protein knowledgebase in 2023, Nucleic Acids Research, 51, Pages D523–D531 (2023). [↩] [↩] [↩] [↩] [↩]
- The UniProt Consortium, UniProt: the universal protein knowledgebase in 2023, Nucleic Acids Research, 51, Pages D523–D531 (2023). [↩] [↩] [↩]
- ClinVar. National Library of Medicine: National Center for Biotechnology Information, www.ncbi.nlm.nih.gov/clinvar/ (2023). [↩] [↩] [↩] [↩] [↩]
- S. Richards, N. Aziz, S. Bale, K. Voelkerding, H. L. Rehm. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genetics in Medicine, 17, 405-424 (2015). [↩] [↩] [↩]
- K. East, W. Chung, K. Foreman, M. Gilmore, M. Gornick, L. Hindorff, T. Kauffman, D. Messersmith, C. Prows, E. Stoffel, J.-H. Yu, S. Plon. Guide to interpreting genomic reports: a genomics toolkit. Clinical Sequencing Exploratory Research (CSER), https://www.genome.gov/sites/default/files/media/files/2020-04/Guide_to_Interpreting_Genomic_Reports_Toolkit.pdf [↩] [↩]
- A. Henrie, S. E. Hemphill, N. Ruiz-Schultz, B. Cushman, M. T. DiStefano, D. Azzariti, S. M. Harrison, H. L. Rehm, K. Eilbeck. ClinVar miner: demonstrating the utility of a web-based tool for viewing and filtering ClinVar data. Hum Mutat. 8, 1051-1060 (2018). [↩] [↩] [↩]
- HGVS simple. Human Genome Variation Society, https://varnomen.hgvs.org/bg-material/simple/ (2023). [↩] [↩] [↩] [↩] [↩]