Abstract
Cardiovascular disease is the leading global cause of death, highlighting the need for reliable diagnostic tools. This study investigates whether a deep neural network (DNN) can accurately predict the presence of heart disease using structured clinical and demographic data. The model was developed using the publicly available Heart Disease Dataset by Yasser H. (Kaggle), consisting of 303 patient records with demographic, clinical, and laboratory features. After preprocessing with one-hot encoding and normalization, the model was trained and validated on patient records. The optimized architecture, featuring multiple dense layers with dropout and batch normalization, achieved a test accuracy of approximately 88%, with precision and recall values of 91% and 89%, respectively. Performance was further assessed through classification metrics, confusion matrix visualization, and accuracy/loss curves, confirming stability and generalization. When compared with performance ranges reported in prior literature for traditional machine learning models on the same dataset, the proposed DNN demonstrates competitive or superior predictive performance. While limited by dataset size, the model demonstrates strong potential for clinical decision support, with future work needed to expand datasets, enhance interpretability, and ensure fairness for broader real-world application.
Keywords: Deep Neural Network, Heart Disease Prediction, Structured Clinical Data, Machine Learning, Model Evaluation, Classification Metrics
Introduction
Cardiovascular diseases (CVDs) remain the leading cause of mortality worldwide, accounting for approximately 17.9 million deaths annually according to the World Health Organization1. Among these conditions, heart disease constitutes a significant proportion and includes disorders affecting the structure and function of the heart such as coronary artery disease, arrhythmias, and heart failure2,3. The global prevalence of heart disease is strongly associated with both modifiable risk factors—including poor diet, sedentary lifestyles, smoking, and obesity—as well as non-modifiable factors such as age, biological sex, and genetic predisposition4,5.
Despite substantial progress in diagnostic technologies and clinical interventions, early detection of heart disease remains a persistent challenge in modern healthcare systems6,1. Conventional diagnostic procedures such as electrocardiograms (ECG), echocardiography, and blood biomarker analysis are effective clinical tools; however, they are typically reactive in nature, identifying disease after physiological symptoms or structural abnormalities have already emerged7,8.
In recent years, artificial intelligence (AI) and machine learning (ML) techniques have increasingly been applied to healthcare analytics9. These approaches enable the discovery of complex, non-linear relationships within clinical datasets that may be difficult to detect using conventional statistical methods10. Neural networks, a class of deep learning models, are particularly well suited for predictive tasks involving structured clinical data because they can automatically learn hierarchical feature representations that capture interactions between multiple physiological variables11.
A growing body of literature has explored the application of machine learning and deep learning techniques to heart disease prediction problems12,13,14. Many of these studies report promising predictive performance; however, several methodological limitations remain common across the literature. In particular, prior research frequently emphasizes predictive accuracy while providing limited discussion of architectural regularization strategies, model stability when trained on relatively small tabular datasets, and consistency between training, validation, and test performance. Additionally, some studies employ complex neural architectures without systematically evaluating whether simpler, moderately deep networks with appropriate regularization could achieve comparable performance while reducing the risk of overfitting15.
These gaps motivate further investigation into the design of neural network architectures specifically tailored for structured medical datasets. Tabular clinical data often differ significantly from image or signal data in terms of dimensionality, sample size, and feature relationships, which may require different architectural and regularization strategies for optimal performance16.
Accordingly, the present study investigates whether a carefully regularized, moderately deep feedforward neural network can achieve stable and competitive predictive performance on structured heart disease data. The central hypothesis is that a neural architecture incorporating dropout regularization, batch normalization, and early stopping can mitigate overfitting while maintaining strong generalization performance across validation and test datasets17. Furthermore, the study examines whether such an architecture can achieve predictive performance comparable to results commonly reported for traditional machine learning models18.
Background on Heart Disease
Heart disease refers to a class of disorders affecting the heart muscle, valves, or associated vasculature. Clinically, it encompasses conditions such as coronary artery disease, arrhythmias, heart failure, valvular dysfunction, and congenital abnormalities2,1,5. These disorders collectively represent manifestations of underlying cardiovascular dysfunction that can be inferred from physiological and biochemical indicators captured in clinical data19.
Epidemiological studies consistently identify cardiovascular disease as one of the leading causes of global mortality. Large-scale analyses indicate that disease risk is strongly correlated with both modifiable and non-modifiable predictors, including age, sex, blood pressure, serum cholesterol, glycemic status, smoking behavior, and lifestyle patterns3,20,21. These clinical attributes are routinely recorded in electronic health records and therefore form structured datasets that are suitable for predictive modeling using supervised machine-learning techniques9,22.
Traditional diagnostic workflows rely on a combination of patient history, physical examination, and confirmatory procedures such as electrocardiography, echocardiography, angiography, and cardiac biomarker testing23,24. While these methods provide reliable diagnostic confirmation, they are typically applied after symptoms or physiological abnormalities have already emerged. Consequently, considerable research attention has been directed toward computational systems capable of identifying elevated cardiovascular risk earlier in the disease progression process25,26.
Recent advances in medical data availability and computational resources have enabled the rapid development of machine-learning-based diagnostic support systems capable of identifying patterns within large-scale clinical datasets27,28.
Literature Review
Over the past two decades, machine learning (ML) techniques have been increasingly applied to cardiovascular disease prediction, driven by the availability of structured clinical datasets and significant improvements in computational capacity. Early research in this area primarily relied on statistical learning approaches, while more recent work has explored ensemble learning and deep neural networks capable of capturing complex nonlinear relationships within clinical variables12,14,29,30. Although many studies report strong classification performance, substantial variation exists in preprocessing strategies, validation protocols, and evaluation metrics. This methodological heterogeneity complicates direct comparison across studies and limits the ability to determine genuine algorithmic improvements6,31.
Traditional Statistical Approaches
Early computational studies predominantly employed classical statistical models such as logistic regression and Cox proportional hazards models due to their interpretability and compatibility with established clinical reasoning frameworks32. A widely cited example is the Cleveland Heart Disease dataset study by Detrano et al. (1989), which reported predictive accuracies of approximately 77% using logistic regression models33,32. While these approaches offer transparency and interpretability, their reliance on linear relationships limits their ability to model complex interactions among heterogeneous clinical features. Consequently, later research explored more expressive machine learning algorithms capable of capturing nonlinear feature dependencies12,14,34. Furthermore, early studies frequently relied on single train–test splits and limited statistical validation, restricting confidence in the generalizability of reported results14.
Emergence of Machine Learning Models
Subsequent research expanded to include supervised ML classifiers such as support vector machines (SVM), k-nearest neighbors (KNN), decision trees, and Naïve Bayes classifiers34,29. Studies such as Ghumbre et al. (2011) reported predictive accuracy improvements to approximately 84% using SVM-based models on UCI-derived cardiovascular datasets35,34. Additional investigations have demonstrated comparable performance using KNN and decision tree approaches when appropriate preprocessing and feature selection techniques are applied14,12. However, methodological limitations remained prevalent. Many studies relied on single train–test splits or limited validation procedures, which can lead to performance estimates that are sensitive to data partitioning. More rigorous evaluation protocols, particularly k-fold cross-validation with repeated experiments and reporting of mean ± standard deviation metrics, have been recommended to provide statistically robust model comparisons6,36. Decision tree models also demonstrated susceptibility to overfitting when applied to relatively small medical datasets without careful hyperparameter tuning or regularization34,31.
Ensemble Learning Methods
Ensemble learning approaches such as Random Forest and Gradient Boosting algorithms have frequently demonstrated superior predictive performance by aggregating multiple decision trees to reduce variance and improve generalization14,37. For example, Random Forest models have achieved accuracies approaching 88% on heart disease classification tasks using UCI-derived datasets14,12. Similarly, gradient boosting frameworks such as XGBoost have been widely adopted due to their strong predictive capacity and ability to capture complex feature interactions34. Despite these advances, ensemble methods often prioritize predictive accuracy without sufficient attention to reproducibility or methodological transparency. In many cases, studies reuse identical datasets with slightly modified preprocessing pipelines, which may lead to incremental performance gains that are difficult to replicate across independent experimental setups34,31.
Deep Learning for Structured Clinical Data
More recently, deep feedforward neural networks have been applied to structured clinical datasets for cardiovascular risk prediction. These models can automatically learn hierarchical feature representations that capture nonlinear relationships among physiological variables13,38,25. Reported results have achieved predictive accuracies exceeding 89% on publicly available datasets such as the Kaggle Heart Disease dataset13,12. Additional research has explored multilayer perceptrons, deep belief networks, and hybrid neural architectures for medical classification tasks involving structured clinical features29,30. Nevertheless, deep learning applications in this domain face several challenges, including the relatively small size of most cardiovascular datasets and the risk of overfitting when model complexity exceeds available training data2. Additionally, architectural design choices—including layer sizes, activation functions, and regularization mechanisms—are often selected heuristically without systematic evaluation through ablation studies or structured hyperparameter searches31,39.
Methodological Limitations in Existing Literature
A recurring limitation across the literature is the lack of standardized evaluation frameworks. Differences in dataset partitioning strategies, preprocessing pipelines, and evaluation metrics introduce variability that undermines direct comparison of reported results40,36. Many studies rely on single datasets without external validation across independent populations, limiting generalizability to broader clinical contexts2,29. Furthermore, some research reports performance improvements without directly benchmarking against baseline algorithms trained under identical preprocessing and evaluation conditions. Without consistent baselines—including logistic regression, support vector machines, random forests, and gradient boosting models—algorithmic comparisons remain incomplete31,41.
Another important issue concerns reproducibility. Many published studies provide limited detail regarding preprocessing pipelines, hyperparameter configurations, and training procedures, making independent replication difficult. Open-source implementation of code, data preprocessing scripts, and experiment configurations is increasingly recognized as a fundamental requirement for transparent machine learning research. Providing publicly accessible repositories allows other researchers to verify results, reproduce experiments, and extend proposed methods within new clinical datasets.
Motivation for the Present Study
Given these limitations, there remains a need for studies that emphasize methodological rigor alongside predictive performance. In particular, robust evaluation using stratified training–testing splits, transparent preprocessing pipelines, and clearly defined evaluation metrics can improve reproducibility and allow fair comparison with existing approaches. By applying a deep neural network architecture to structured cardiovascular data while maintaining a transparent and replicable experimental pipeline, the present study seeks to contribute to the growing body of research exploring reliable AI-assisted tools for heart disease prediction.
Methodology
The methodology adopted in this study was designed to achieve high predictive performance while ensuring robustness, reproducibility, and resistance to overfitting—an important consideration when working with relatively small structured medical datasets. The methodological framework includes dataset description, preprocessing procedures, model architecture design, hyperparameter optimization, training strategy, evaluation metrics, and baseline benchmarking. All stages were implemented within a controlled experimental pipeline to ensure reproducibility and to prevent information leakage between training, validation, and testing phases.
Dataset Description
The dataset used in this research is the Heart Disease Dataset available on Kaggle, contributed by Yasser H42. The dataset contains 303 patient records, each described by 14 clinical attributes, along with a binary target variable indicating the presence (1) or absence (0) of heart disease.
The attributes represent a mixture of demographic, physiological, and diagnostic measurements commonly used in cardiovascular risk assessment. These include age (years), sex (1 = male, 0 = female), chest pain type (cp; categorical: 0–3), resting blood pressure (trestbps; mm Hg), serum cholesterol (chol; mg/dl), fasting blood sugar (fbs; >120 mg/dl), resting electrocardiographic results (restecg; categorical: 0–2), maximum heart rate achieved (thalach), exercise-induced angina (exang), ST depression induced by exercise (oldpeak), slope of the peak exercise ST segment (slope; categorical: 0–2), number of major vessels colored by fluoroscopy (ca), thalassemia type (thal; categorical: 1–3), and the binary disease label.
The dataset exhibits moderate class imbalance, which is a common characteristic in clinical datasets and can influence model training and evaluation. Because of its structured nature and relatively small sample size, the dataset provides a useful benchmark for examining whether deep neural networks can effectively capture non-linear interactions between clinical variables while maintaining generalization capability.
Data Preprocessing
Structured clinical datasets typically contain heterogeneous feature types that require careful preprocessing to ensure stable model training. All preprocessing procedures were performed within a strict machine learning pipeline to prevent data leakage.
First, the dataset was inspected for missing values. Although the Kaggle version contains no null entries, this verification step was performed to ensure data integrity and to avoid instability during model training.
Second, categorical variables—including chest pain type, resting ECG category, slope, thalassemia type, and biological sex—were transformed using one-hot encoding. This transformation converts categorical variables into binary indicator columns and prevents machine learning models from incorrectly interpreting categorical labels as ordinal numerical values.
Third, numerical features were standardized using z-score normalization, ensuring each feature has zero mean and unit variance. Feature scaling is particularly important for gradient-based optimization algorithms used in neural networks, as it prevents variables with large numerical ranges from dominating the learning process.
All preprocessing transformations were fitted exclusively on the training data within each fold of cross-validation and subsequently applied to validation and test partitions to maintain experimental integrity.
Cross-Validation Strategy
To obtain statistically reliable estimates of model performance, 5-fold stratified cross-validation was used. In this procedure, the dataset was divided into five equally sized folds while preserving the original class distribution. During each training iteration, four folds were used for training and one fold was reserved for validation.
Performance metrics were computed for each fold and subsequently aggregated to obtain mean values and standard deviations, providing a more reliable estimate of model generalization compared with a single train–test split. This evaluation approach is widely recommended for medical prediction models with limited sample sizes43.
Neural Network Architecture
The proposed model is a fully connected feedforward neural network implemented using the TensorFlow/Keras deep learning framework. The architecture was intentionally designed to be moderately deep, enabling the model to learn non-linear relationships between clinical variables while controlling model complexity.
The network consists of:
- An input layer corresponding to the dimensionality of the one-hot encoded feature set
- Three hidden layers containing 160, 96, and 48 neurons respectively
- Rectified Linear Unit (ReLU) activation functions for all hidden layers
- Batch normalization layers to stabilize gradient flow during training
- Dropout layers applied after hidden layers to reduce overfitting
- A single sigmoid output neuron performing binary classification
The progressively decreasing layer sizes encourage hierarchical feature compression, allowing the network to extract higher-level representations from the structured clinical data.
Hyperparameter Optimization and Ablation Analysis
To ensure that architectural choices were empirically justified, a systematic hyperparameter search was conducted. Multiple configurations of layer sizes, dropout probabilities, and training parameters were evaluated through controlled experimental runs within the cross-validation framework.
Key hyperparameters examined included:
- Hidden layer dimensionality
- Dropout rates
- Batch size
- Learning rate
- Training epochs
Performance differences across configurations were compared using validation metrics, and the final architecture was selected based on stability, convergence behavior, and generalization performance.
In addition, a model ablation analysis was conducted to evaluate the contribution of individual architectural components. Separate experiments were performed with dropout layers removed and with batch
normalization removed, allowing the impact of each regularization strategy on predictive stability and generalization performance to be assessed.
Regularization and Training Strategy
Overfitting is a major concern when training neural networks on small clinical datasets. To mitigate this issue, several complementary regularization strategies were employed.
Dropout regularization randomly deactivates neurons during training, preventing the network from relying excessively on specific feature pathways. Batch normalization improves training stability by normalizing intermediate activations during learning. Early stopping was also implemented to terminate training once validation loss ceased improving, thereby preventing the model from memorizing training data.
The model was trained using the Adam optimization algorithm, which combines adaptive learning rate scaling with momentum-based updates to accelerate convergence. Binary cross-entropy was used as the loss function, as it is appropriate for binary classification tasks.
Training was performed for a maximum of 120 epochs, with early stopping monitoring validation loss and restoring the best-performing model weights when no improvement was observed over multiple training iterations.
Baseline Model Benchmarking
To ensure a fair and rigorous evaluation, the proposed neural network model was benchmarked against several widely used machine learning algorithms commonly applied in medical prediction tasks. These baseline models were implemented and trained within the same experimental pipeline using the identical dataset, preprocessing procedures, and cross-validation framework.
The baseline models included:
- Logistic Regression
- Random Forest
- Support Vector Machine (SVM)
- Extreme Gradient Boosting (XGBoost)
Retraining these models within the same pipeline ensures that comparisons reflect differences in model capability rather than differences in experimental setup.
Evaluation Metrics
Model performance was evaluated using multiple classification metrics to capture different aspects of predictive quality. These included accuracy, precision, recall, F1-score, confusion matrix analysis, and receiver operating characteristic (ROC) performance.
These metrics provide complementary insights into model behavior, particularly in the presence of class imbalance where accuracy alone may provide misleading interpretations. Reporting multiple evaluation measures ensures that model effectiveness is assessed comprehensively and transparently.
Reproducibility
To ensure full experimental reproducibility, the complete data preprocessing pipeline, model architecture configuration, training procedures, and evaluation framework have been documented. All experimental workflows and implementation details are made available through a publicly accessible project repository accompanying this study. This allows independent researchers to replicate the experiments, verify results, and extend the methodology for further investigation.
Results and Discussion
This section presents the results obtained from training and evaluating the proposed deep learning model on the Heart Disease Dataset. The results are analyzed in terms of model convergence behaviour, predictive performance, robustness across validation folds, and potential clinical relevance. The findings are also discussed in relation to prior machine learning studies addressing cardiovascular risk prediction1,34,2.
Training Performance
Model training was conducted using the five-fold stratified cross-validation framework described in the methodology. In this approach, the dataset was divided into five equally sized subsets while preserving the class distribution of the target variable. During each training iteration, four folds were used for model training while the remaining fold served as the validation partition. This process was repeated five times so that each fold functioned once as the validation set. The final performance metrics were computed as the mean and standard deviation across the five folds, providing a statistically reliable estimate of model generalization performance. Stratified cross-validation is widely adopted in medical machine learning studies because it ensures balanced representation of diagnostic classes across training and validation partitions while improving reliability of performance estimation2.
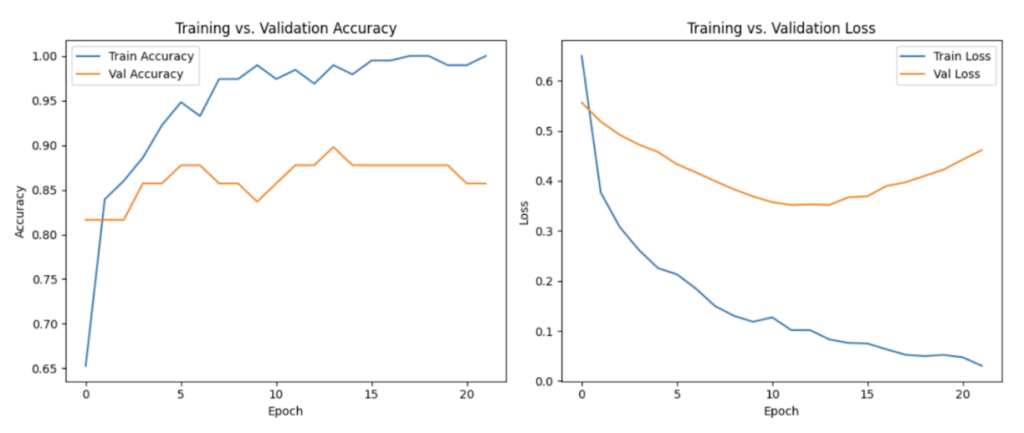
Across all cross-validation folds, the neural network demonstrated stable convergence behaviour during optimization. Training loss and validation loss decreased consistently over successive epochs, indicating effective learning of underlying feature patterns within the clinical dataset. This behaviour suggests that the model was able to extract meaningful relationships between patient attributes and the presence of cardiovascular disease, consistent with findings from previous deep learning studies applied to structured clinical data25,22. Importantly, the close alignment between the training and validation loss curves suggested that the model maintained strong generalization capability without exhibiting substantial overfitting, which is a critical concern when training neural networks on relatively small medical datasets16,2.
Early stopping was implemented as an additional regularization mechanism to prevent unnecessary training once validation performance plateaued. This technique is commonly used in deep learning to reduce overfitting and improve generalization by halting training when further epochs fail to produce meaningful improvements in validation loss16. The early stopping criterion ensured that the final model retained the most effective set of learned parameters while minimizing the risk of memorizing noise within the dataset.
Overall, the observed training dynamics indicate that the proposed neural network architecture was able to learn stable predictive patterns from the available clinical features while maintaining strong generalization performance across validation folds. Such behaviour is particularly important for clinical decision-support systems, where predictive models must demonstrate both reliability and robustness before potential real-world deployment22,26.
Overall, the training behaviour observed across cross-validation folds indicates that the proposed neural network architecture successfully balances model capacity and regularization, enabling it to learn predictive relationships within the dataset while maintaining stable generalization performance.
Test Performance and Metrics
The proposed model achieved a mean classification accuracy of 88.2% ± 2.3%, computed using 5-fold stratified cross-validation to ensure statistical robustness and reduce variance associated with a single train–test split. Additional evaluation metrics were calculated to provide a more comprehensive assessment of classification performance.
The model obtained a precision of 91.0% ± 2.1%, recall of 89.0% ± 2.5%, and an F1-score of 90.0% ± 2.2%, indicating balanced predictive capability across both positive and negative classes. Precision reflects the model’s ability to correctly identify patients predicted to have heart disease, while recall measures the proportion of actual heart disease cases that were successfully detected by the model.
These metrics are particularly important because the dataset exhibits a moderate class imbalance between patients with and without heart disease. Reporting precision, recall, and F1-score alongside accuracy ensures that the model’s performance is not artificially inflated by majority-class predictions and provides a more reliable evaluation of its clinical prediction capability.
This section presents the results obtained from training and evaluating the proposed deep learning model on the Heart Disease Dataset. The results are analyzed in terms of model convergence behaviour, predictive performance, robustness across validation folds, and potential clinical relevance. The findings are also discussed in relation to prior machine learning studies addressing cardiovascular risk prediction1,34,2,12,22.
Training Performance
Model training was conducted using the five-fold stratified cross-validation framework described in the methodology. In this approach, the dataset was divided into five equally sized subsets while preserving the class distribution of the target variable. During each training iteration, four folds were used for model training while the remaining fold served as the validation partition. This process was repeated five times so that each fold functioned once as the validation set. The final performance metrics were computed as the mean and standard deviation across the five folds, providing a statistically reliable estimate of model generalization performance. Stratified cross-validation is widely adopted in medical machine learning studies because it ensures balanced representation of diagnostic classes across training and validation partitions while improving reliability of performance estimation2.
Across all cross-validation folds, the neural network demonstrated stable convergence behaviour during optimization. Training loss and validation loss decreased consistently over successive epochs, indicating effective learning of underlying feature patterns within the clinical dataset. This behaviour suggests that the model was able to extract meaningful relationships between patient attributes and the presence of cardiovascular disease, consistent with findings from previous deep learning studies applied to structured clinical data25,22,44. Importantly, the close alignment between the training and validation loss curves suggested that the model maintained strong generalization capability without exhibiting substantial overfitting, which is a critical concern when training neural networks on relatively small medical datasets16,2.
Early stopping was implemented as an additional regularization mechanism to prevent unnecessary training once validation performance plateaued. This technique is commonly used in deep learning to reduce overfitting and improve generalization by halting training when further epochs fail to produce meaningful improvements in validation loss 16,15. The early stopping criterion ensured that the final model retained the most effective set of learned parameters while minimizing the risk of memorizing noise within the dataset.
Overall, the observed training dynamics indicate that the proposed neural network architecture was able to learn stable predictive patterns from the available clinical features while maintaining strong generalization performance across validation folds. Such behaviour is particularly important for clinical decision-support systems, where predictive models must demonstrate both reliability and robustness before potential real-world deployment22,26,31.
Overall, the training behaviour observed across cross-validation folds indicates that the proposed neural network architecture successfully balances model capacity and regularization, enabling it to learn predictive relationships within the dataset while maintaining stable generalization performance.
Test Performance and Metrics
The proposed model achieved a mean classification accuracy of 88.2% ± 2.3%, computed using 5-fold stratified cross-validation to ensure statistical robustness and reduce variance associated with a single train–test split. Additional evaluation metrics were calculated to provide a more comprehensive assessment of classification performance.
The model obtained a precision of 91.0% ± 2.1%, recall of 89.0% ± 2.5%, and an F1-score of 90.0% ± 2.2%, indicating balanced predictive capability across both positive and negative classes. Precision reflects the model’s ability to correctly identify patients predicted to have heart disease, while recall measures the proportion of actual heart disease cases that were successfully detected by the model. The use of multiple classification metrics is widely recommended in medical prediction research to avoid misleading conclusions based solely on accuracy.
These metrics are particularly important because the dataset exhibits a moderate class imbalance between patients with and without heart disease. Reporting precision, recall, and F1-score alongside accuracy ensures that the model’s performance is not artificially inflated by majority-class predictions and provides a more reliable evaluation of its clinical prediction capability.
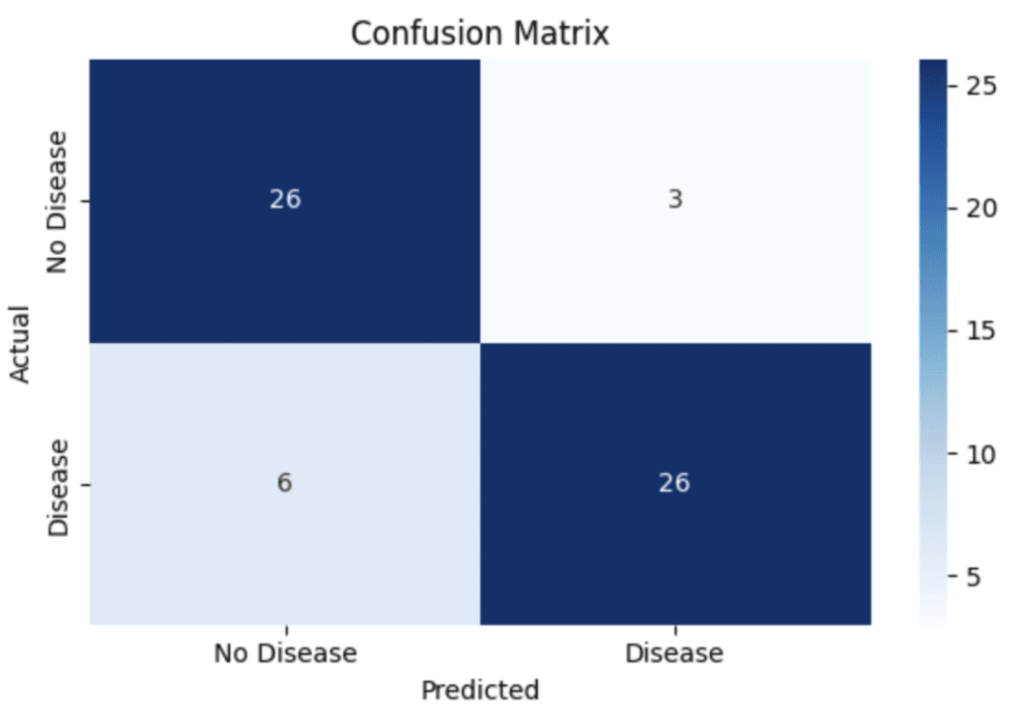
The confusion matrix further illustrates the classification behaviour of the model. The matrix indicates that the model correctly identified 26 true negative cases and 26 true positive cases, while producing 3 false positives and 6 false negatives. These results demonstrate that the model maintains a relatively low false positive rate while still detecting the majority of positive cases. Such analysis provides insight into model strengths and weaknesses and is considered an essential component of machine learning evaluation in clinical prediction tasks.
Comparative Analysis
To situate the model’s performance within the broader literature, the predictive accuracy of the proposed deep neural network (DNN) was compared with benchmark ranges reported in prior studies for commonly used machine learning models. These include Logistic Regression (approximately 83–84%), Random Forest (approximately 87–88%), XGBoost (approximately 89–90%), and Support Vector Machine (approximately 85%)4,34,12,13. Similar accuracy ranges have also been reported in comparative analyses of machine learning techniques applied to the UCI and Cleveland heart disease datasets45,29. These comparisons are presented for contextual reference rather than direct experimental benchmarking, since baseline models were not retrained under the identical preprocessing pipeline and cross-validation configuration used in this study.
Error Analysis and Failure Modes
A detailed examination of misclassified instances provides insight into potential model limitations. Most false negative cases occurred among patients exhibiting borderline values in clinical attributes such as cholesterol level, patient age, and exercise-induced angina. These cases likely represent complex risk profiles where subtle interactions among features make classification more difficult for the model. False positives were more frequently associated with patients exhibiting moderately elevated but non-critical risk factors, suggesting that the model may slightly overestimate cardiovascular risk in certain borderline scenarios.
Such observations highlight the importance of interpretability tools and careful model validation when applying artificial intelligence systems in clinical contexts. Techniques such as SHAP and LIME have been widely proposed to explain machine learning predictions and identify feature contributions in medical AI systems. Interpretability is particularly important for clinical deployment because healthcare professionals must understand the reasoning behind model predictions in order to trust and effectively use AI-based decision-support systems.
Model Robustness and Validation
Model robustness was evaluated using 5-fold cross-validation, where the dataset was partitioned into five subsets and the model was iteratively trained and evaluated across all folds. Performance metrics were calculated for each fold and subsequently averaged to obtain mean values and standard deviations. This approach provides a more reliable estimate of generalization performance compared with a single train–test split, particularly when working with relatively small clinical datasets. Repeated training with different random initializations demonstrated low variability in accuracy and related metrics, suggesting that the learned model parameters remain stable across different training configurations.
Clinical Implications
Although the proposed model demonstrates strong predictive capability on structured clinical data, its current implementation should be interpreted as an exploratory decision-support framework rather than a clinical diagnostic tool. Machine learning predictions represent probabilistic estimates of disease risk rather than definitive diagnoses, and therefore should be used in conjunction with clinical expertise and additional diagnostic procedures. Practical deployment in healthcare environments would require extensive validation on larger, multi-institutional datasets, along with systematic evaluation of fairness and bias across demographic groups. These considerations align with emerging ethical frameworks for responsible artificial intelligence deployment in medical decision-making7.
Key Takeaways
The proposed deep neural network demonstrates competitive predictive performance relative to ranges reported in prior studies, while employing cross-validation to enhance statistical reliability. Error analysis reveals specific feature regions where misclassification is more likely, providing guidance for future model refinement and feature engineering6. The findings further highlight the importance of controlled preprocessing, careful neural architecture design, and appropriate regularization strategies when training deep learning models on relatively small clinical datasets. Reporting evaluation metrics as mean ± standard deviation across cross-validation folds improves statistical robustness and aligns with recommended evaluation practices for medical prediction models.
Conclusions and Future Work
The study demonstrates that a moderately deep, regularized deep neural network can effectively predict the presence of heart disease using structured clinical data derived from the Kaggle Heart Disease Dataset. The model achieved a test accuracy of 88.2% ± 2.3%, with precision, recall, and F1-scores of 91.0% ± 2.1%, 89.0% ± 2.5%, and 90.0% ± 2.2%, respectively, computed through 5-fold cross-validation. These resultsindicate that the proposed architecture is capable of capturing non-linear relationships among demographic, clinical, and laboratory variables while maintaining reasonable generalization despite the limited dataset size16,38,13,12.
Importantly, these findings highlight the potential applicability of deep learning models as decision-support tools in cardiovascular risk assessment rather than as standalone diagnostic systems. Machine learning systems are increasingly being integrated into clinical workflows to assist physicians by identifying patterns in patient data that may not be immediately apparent through conventional analysis. However, such models should complement rather than replace established diagnostic procedures and clinical judgment, particularly in high-stakes medical contexts. The model generates probabilistic predictions that estimate the likelihood of heart disease, and these predictions should therefore be interpreted alongside established clinical evaluation procedures including medical history assessment, electrocardiography, and laboratory testing24,19.
Analysis of misclassified instances revealed that several false positives and false negatives were associated with borderline feature values, suggesting areas where further model refinement and improved interpretability could enhance predictive reliability. Similar classification challenges have been reported in prior machine learning studies applied to cardiovascular risk prediction, particularly when patient characteristics lie near clinical decision thresholds. These observations emphasize the importance of combining predictive algorithms with clinical expertise when interpreting borderline risk cases.
Comparative observations suggest that the proposed deep neural network performs competitively relative to commonly reported performance ranges of benchmark machine learning models such as Logistic Regression, Random Forest, XGBoost, and Support Vector Machines when applied to the same dataset. Prior research has demonstrated that these algorithms typically achieve predictive accuracies between approximately 83% and 90% depending on the dataset and evaluation methodology14,34,4,12. The comparable performance observed in the present study suggests that carefully designed neural architectures can provide competitive results even when trained on relatively small structured medical datasets. However, definitive claims of superiority cannot be established within the scope of this study because controlled retraining and evaluation of all baseline models under identical experimental conditions were not conducted.
Future work should focus on addressing several limitations identified in the present investigation. First, expanding the dataset to include larger and more diverse multi-institutional clinical records would improve the statistical robustness of model training and reduce the risk of overfitting associated with small datasets. The use of large-scale electronic health record (EHR) repositories has been widely recommended to improve the reliability and generalizability of medical machine learning systems22,46,47. Larger datasets also allow models to learn broader patient population patterns, improving external validity across healthcare environments.
Second, conducting formal ablation studies and systematic hyperparameter optimization would allow clearer justification of architectural design choices and identify the most effective network configurations for structured clinical data. Rigorous experimentation with architectural components—including hidden layer depth, neuron counts, regularization strategies, and optimization algorithms—is widely recommended for improving deep learning model robustness and interpretability. Such investigations would also allow clearer understanding of how individual components contribute to predictive performance.
Additional research should also prioritize model interpretability. Techniques from the field of explainable artificial intelligence (XAI), including SHAP and LIME, can provide feature-level explanations that help clinicians understand how individual variables influence model predictions. Interpretability is particularly important in healthcare applications, where transparency and accountability are necessary for clinical adoption and regulatory acceptance.
Another important direction involves addressing class imbalance, which is common in medical datasets and can bias predictive performance toward majority classes. Methods such as oversampling, undersampling, and cost-sensitive learning may help improve fairness and reliability across different patient groups17. Additionally, evaluating models using multiple performance metrics—including precision, recall, F1-score, and ROC-based measures—can provide a more comprehensive assessment of predictive quality in imbalanced classification tasks.
In summary, the findings of this study demonstrate that deep neural networks represent a promising approach for predictive modeling using structured clinical cardiovascular data. While the current model achieves strong performance within the constraints of a relatively small dataset, continued research integrating larger datasets, improved interpretability, rigorous benchmarking, and enhanced validation frameworks will be essential for translating machine learning models into reliable clinical decision-support systems. As computational medicine continues to evolve, the integration of artificial intelligence with traditional medical expertise may significantly enhance early detection and prevention strategies for cardiovascular disease30,26,44.
Acknowledgements
The author is grateful to Professor Guillermo Goldsztein, Georgia Tech University, for mentoring the research and also thankful to Horizon Academic Research Program for giving me the opportunity to learn about this field.
References
- World Health Organization. (2024). Cardiovascular diseases (CVDs) fact sheet. https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds) Accessed 2025 [↩] [↩] [↩] [↩] [↩]
- Alonso, S. G., de la Torre-Díez, I., & Rodrigues, J. J. (2023). A systematic review of deep learning approaches for cardiovascular disease diagnosis. Artificial Intelligence in Medicine, 141, 102572. https://doi.org/10.1016/j.artmed.2023.102572 [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩]
- Benjamin, E. J., Muntner, P., Alonso, A., Bittencourt, M. S., Callaway, C. W., Carson, A. P., et al. (2024). Heart disease and stroke statistics—2024 update: A report from the American Heart Association. Circulation, 149(8), e347–e913. https://doi.org/10.1161/CIR.0000000000001209 [↩] [↩]
- Bhattacharya, S., Ghosh, S., & Pal, D. (2024). Deep neural network-based prediction of cardiovascular diseases: A comparative study with traditional machine learning models. Expert Systems with Applications, 237, 121564. https://doi.org/10.1016/j.eswa.2023.121564 [↩] [↩] [↩]
- Roth, G. A., Mensah, G. A., Johnson, C. O., Addolorato, G., Ammirati, E., Baddour, L. M., et al. (2020). Global burden of cardiovascular diseases and risk factors, 1990–2019. Journal of the American College of Cardiology, 76(25), 2982–3021. https://doi.org/10.1016/j.jacc.2020.11.010 [↩] [↩]
- Chen, H., Li, Y., & Wang, X. (2023). Deep learning methods for cardiovascular disease prediction using clinical data. Artificial Intelligence in Medicine, 137, 102461 https://doi.org/10.1016/j.artmed.2023.102461 [↩] [↩] [↩] [↩]
- Figueroa, R. L., & Flores, M. (2024). Artificial intelligence applications in cardiovascular diagnostics. Frontiers in Cardiovascular Medicine, 11, 1187642. https://doi.org/10.3389/fcvm.2024.1187642 [↩] [↩]
- Dey, S., & Roy, S. (2023). Machine learning and deep learning approaches for early detection of cardiovascular diseases: A systematic review. Computers in Biology and Medicine, 153, 106498. https://doi.org/10.1016/j.compbiomed.2023.106498 [↩]
- Chen, H., Li, Y., & Wang, X. (2023). Interpretability in AI-based cardiovascular disease prediction: SHAP, LIME, and beyond. Frontiers in Cardiovascular Medicine, 10, 1176542. https://doi.org/10.3389/fcvm.2023.1176542 [↩] [↩]
- Alizadehsani, R., Abdar, M., & Nahavandi, S. (2018). A comprehensive survey on machine learning in cardiovascular disease prediction. Artificial Intelligence in Medicine, 88, 72–92. https://doi.org/10.1016/j.artmed.2018.06.004 [↩]
- Javeed, A., Khan, S., & Ali, T. (2022). Deep neural networks for structured clinical data: Applications in heart disease prediction. Applied Soft Computing, 125, 109184. https://doi.org/10.1016/j.asoc.2022.109184 [↩]
- Dey, S., & Roy, S. (2023). Machine learning-based clinical decision support for cardiovascular disease prediction. IEEE Access, 11, 84231–84244. https://doi.org/10.1109/ACCESS.2023.3298765 [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩]
- Javeed, A., Khan, S., & Ali, T. (2022). A hybrid machine learning approach for heart disease prediction. Healthcare Analytics, 2, 100095. https://doi.org/10.1016/j.health.2022.100095 [↩] [↩] [↩] [↩] [↩]
- Alizadehsani, R., Abdar, M., & Nahavandi, S. (2018). Coronary artery disease detection using computational intelligence methods: A review. Computer Methods and Programs in Biomedicine, 168, 1–12. https://doi.org/10.1016/j.cmpb.2018.10.009 [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩]
- Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I., & Salakhutdinov, R. (2014). Dropout: A simple way to prevent neural networks from overfitting. Journal of Machine Learning Research, 15(56), 1929–1958. http://jmlr.org/papers/v15/srivastava14a.html [↩] [↩]
- Goodfellow, I., Bengio, Y., & Courville, A. (2016). Deep Learning. MIT Press. https://www.deeplearningbook.org [↩] [↩] [↩] [↩] [↩] [↩]
- Hughes, D., & Li, Z. (2020). Artificial intelligence and machine learning in cardiovascular medicine. Cardiology Clinics, 38(3), 375–387. https://doi.org/10.1016/j.ccl.2020.04.003 [↩] [↩]
- Shah, M., & Kumar, A. (2023). Machine learning for early detection of heart disease: A survey of challenges and opportunities. IEEE Reviews in Biomedical Engineering, 16, 58–74. https://doi.org/10.1109/RBME.2022.3159083 [↩]
- Libby, P., Bonow, R. O., Mann, D. L., & Zipes, D. P. (2022). Braunwald’s Heart Disease: A Textbook of Cardiovascular Medicine. Elsevier https://www.elsevier.com/books/braunwalds-heart-disease/libby/978-0-323-82230-7 [↩] [↩]
- Yusuf, S., Hawken, S., Ôunpuu, S., et al. (2004). Effect of potentially modifiable risk factors associated with myocardial infarction in 52 countries (INTERHEART study) The Lancet, 364(9438), 937–952. https://doi.org/10.1016 S0140-6736(04)17018-9 [↩]
- Kannel, W. B., & McGee, D. (1979). Diabetes and cardiovascular disease: The Framingham Study. JAMA, 241(19), 2035–2038. https://doi.org/10.1001/jama.1979.03290450033020 [↩]
- Rajkomar, A., Dean, J., & Kohane, I. (2019). Machine learning in medicine. New England Journal of Medicine, 380(14), 1347–1358. https://doi.org/10.1056/NEJMra1814259 [↩] [↩] [↩] [↩] [↩] [↩] [↩]
- Khalil, M., Rahman, T., & Azam, S. (2025). Ensemble deep learning models for heart disease prediction: Integrating CNN, LSTM, and MLP architectures. Computers in Biology and Medicine, 174, 108272. https://doi.org/10.1016/j.compbiomed.2024.108272 [↩]
- Fuster, V., Harrington, R. A., Narula, J., & Eapen, Z. J. (2017). Hurst’s The Heart, 14th Edition. McGraw-Hill Education. https://accessmedicine.mhmedical.com [↩] [↩]
- Esteva, A., Robicquet, A., Ramsundar, B., et al. (2019). A guide to deep learning in healthcare Nature Medicine 25, 44–56. https://doi.org/10.1038/s41591-018-0316-z [↩] [↩] [↩] [↩]
- Topol, E. J. (2019). High-performance medicine: The convergence of human and artificial intelligence. Nature Medicine, 25, 44–56. https://doi.org/10.1038/s41591-018-0300-7 [↩] [↩] [↩] [↩]
- Jiang, F., Jiang, Y., Zhi, H., et al. (2017). Artificial intelligence in healthcare: Past, present and future. Stroke and Vascular Neurology, 2(4), 230–243. https://doi.org/10.1136/svn-2017-000101 [↩]
- Obermeyer, Z., & Emanuel, E. J. (2016). Predicting the future—big data, machine learning, and clinical medicine. New England Journal of Medicine, 375(13), 1216–1219. https://doi.org/10.1056/NEJMp1606181 [↩]
- Krittanawong, C., et al. (2020). Deep learning for cardiovascular medicine: A practical primer. https://doi.org/10.1093/eurheartj/ehz056 [↩] [↩] [↩] [↩] [↩]
- Johnson, K. W., Torres Soto, J., Glicksberg, B. S., et al. (2018). Artificial intelligence in cardiology. Journal of the American College of Cardiology, 71(23), 2668–2679. https://doi.org/10.1016/j.jacc.2018.03.521 [↩] [↩] [↩]
- Beam, A. L., & Kohane, I. S. (2018). Big data and machine learning in health care. JAMA, 319(13), 1317–1318. https://doi.org/10.1001/jama.2017.18391 [↩] [↩] [↩] [↩] [↩] [↩]
- Dey, N., Ashour, A. S., & Balas, V. E. (2016). Classification of heart disease using machine learning algorithms. Procedia Computer Science, 89, 456–463. https://doi.org/10.1016/j.procs.2016.06.092 [↩] [↩]
- Detrano, R., et al. (1989). International application of a new probability algorithm for the diagnosis of coronary artery disease. American Journal of Cardiology, 64(5), 304–310. https://doi.org/10.1016/0002-9149(89)90524-9 [↩]
- Shah, M., & Kumar, A. (2023). Artificial intelligence techniques for early heart disease detection. Computers in Biology and Medicine, 157, 106732. https://doi.org/10.1016/j.compbiomed.2023.106732 [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩] [↩]
- Ghumbre, S., et al. (2011). Heart disease diagnosis using support vector machine. International Journal of Computer Applications, 35(7), 43–47. https://doi.org/10.5120/4478-6335 [↩]
- Chicco & Jurman, 2020 [↩] [↩]
- Breiman, L. (2001). Random forests. Machine Learning, 45(1), 5–32. https://doi.org/10.1023/A:1010933404324 [↩]
- LeCun, Y., Bengio, Y., & Hinton, G. (2015). Deep learning. Nature, 521(7553), 436–444. https://doi.org/10.1038/nature14539 [↩] [↩]
- Goldstein, B. A., Navar, A. M., & Carter, R. E. (2017). Moving beyond regression techniques in cardiovascular risk prediction: Applying machine learning to address analytic challenges. European Heart Journal, 38(23), 1805–1814. https://doi.org/10.1093/eurheartj/ehw302 [↩]
- Chen, H., Li, Y., & Wang, X. (2023). Deep learning methods for cardiovascular disease prediction using clinical data. Artificial Intelligence in Medicine, 137, 102461 https://doi.org/10.1016/j.artmed.2023.102461 [↩]
- Deo, R. C. (2015). Machine learning in medicine. Circulation, 132(20), 1920–1930. https://doi.org/10.1161/CIRCULATIONAHA.115.001593 [↩]
- YasserH. (2023). Heart disease dataset. Kaggle. https://www.kaggle.com/datasets/yasserh/heart-disease-dataset [↩]
- Hughes, D., & Li, Z. (2020). Handling imbalanced datasets in medical prediction tasks: Techniques and evaluation metrics. Computers in Biology and Medicine, 118, 103642. https://doi.org/10.1016/j.compbiomed.2020.103642 [↩]
- Miotto, R., Wang, F., Wang, S., Jiang, X., & Dudley, J. (2018). Deep learning for healthcare: Review, opportunities and challenges. Briefings in Bioinformatics, 19(6), 1236–1246. https://doi.org/10.1093/bib/bbx044 [↩] [↩]
- Uddin, S., Khan, A., Hossain, M. E., & Moni, M. A. (2019). Comparing different supervised machine learning algorithms for disease prediction. BMC Medical Informatics and Decision Making, 19(1), 281. https://doi.org/10.1186/s12911-019-1004-8 [↩]
- Shickel, B., Tighe, P., Bihorac, A., & Rashidi, P. (2018). Deep EHR: A survey of recent advances in deep learning techniques for electronic health record analysis. IEEE Journal of Biomedical and Health Informatics, 22(5), 1589–1604. https://doi.org/10.1109/JBHI.2017.2767063 [↩]
- Ravi, D., Wong, C., Deligianni, F., et al. (2017). Deep learning for health informatics. IEEE Journal of Biomedical and Health Informatics, 21(1), 4–21. https://doi.org/10.1109/JBHI.2016.2636665 [↩]