Kashvi Tacker1, Xiaoliang Ba1
1 Cambridge Centre for International Research (CCIR)
Abstract
Tuberculosis (TB) remains a major public health challenge in India, which accounts for the highest burden of drug-resistant TB globally. In recent years, the rise of drug-resistant tuberculosis (DR-TB) continues to undermine national TB control efforts, with increasing complexity in resistance mechanisms and transmission patterns. This data analysis and review aimed to compile genomic data and epidemiological reports from 2001 to 2022 to evaluate resistance trends and genomic patterns to better understand the evolving resistance landscape and inform policy responses. Publicly available whole genome sequences of M. tuberculosis isolates from India were analysed alongside peer-reviewed literature. Resistance-associated mutations, lineage distributions, and temporal trends were examined to identify shifts in dominant strains and resistance profiles. Policy interventions and diagnostic advances during this period were also reviewed. The analysis reveals a sharp increase in Pre-XDR TB, now surpassing MDR TB the national genomic data set over time. Resistance was largely driven by mutations in genes such as katG, rpoB, and gyrA, with rare mutations (ethA, tlyA, thyX) increasingly detected in recent years. Lineage analysis showed a growing dominance of Lineage 2 (Beijing), associated with high levels of fluoroquinolone resistance and rapid community transmission. Expanded diagnostic coverage and national initiatives improved case detection, though challenges in treatment adherence and private sector regulation remain. The growing genetic diversity and drug resistance pose significant challenges to TB control in India. Continued genomic surveillance, alongside strengthened treatment adherence and targeted public health responses, is essential to avoid the spread of resistant TB strains and achieve national elimination goals.
Keywords: Drug-resistant tuberculosis, Multidrug-resistant TB (MDR-TB), Pre-extensively drug-resistant TB (Pre-XDR TB), Mycobacterium tuberculosis, Genomic surveillance, India, Prevalence
Introduction
Tuberculosis (TB) is a prevalent infectious disease that is caused by the bacterium Mycobacterium tuberculosis (M. tuberculosis). Active tuberculosis develops when tubercle bacilli overcome host immune control in individuals with latent infection, allowing the bacilli to multiply rapidly and cause TB disease1. It is one of the oldest and deadliest diseases known to affect humans. Despite being preventable and curable, TB continues to represent a serious public health concern for the human population2. According to the Global Tuberculosis Report 2023, an estimated total of 10.8 million developed TB worldwide, resulting in approximately 1.09 million TB-related deaths. Most TB cases in 2023 were in the WHO regions of South-East Asia (45%)3, with India accounting for 26 percent of the global TB burden as of 2024, making it the country with the highest burden4.
In India, the National Strategic Plan (2017–2025) set a national goal of eliminating tuberculosis by 20255. However, achieving this target has proven increasingly challenging as the bacterium develops resistance to Rifampicin and Isoniazid, which are the first-line antibiotics used for treatment, as well as several second line antibiotics6. Treating drug-resistant tuberculosis places a significant burden on healthcare systems, primarily due to prolonged treatment durations, increased costs, complex drug regimens, and poorer treatment outcomes7.
Drug resistance in M. tuberculosis arises through a combination of intrinsic bacterial defenses and acquired genetic mutations. The organism’s lipid-rich cell envelope and efflux mechanisms reduce intracellular drug concentrations8, while chromosomal mutations impair antibiotic binding, metabolic pathways, or drug activation, thereby diminishing treatment efficacy9,10. Collectively, these mechanisms enable rapid adaptation under selective drug pressure, highlighting the genetic basis of resistance evolution.
Accordingly, drug-resistant TB is classified by the World Health Organization into the following: mono-resistant strains are resistant to a single first-line drug (such as isoniazid-resistant tuberculosis, HR-TB, or rifampicin-resistant tuberculosis, RR-TB);multidrug-resistant (MDR) strains are resistant to at least isoniazid and rifampicin; pre‑extensively drug-resistant (Pre‑XDR) strains are MDR with additional resistance to any fluoroquinolone; and extensively drug-resistant (XDR) strains are MDR, resistant to any fluoroquinolone, and at least one Group A drug (such as bedaquiline or linezolid)11.
One technique that has gained increasing traction and transformed the study of drug-resistant tuberculosis in both research and public health settings is whole-genome sequencing (WGS). This method allows simultaneous detection of known and emerging resistance mutations, enabling high-resolution analysis of resistance-associated mutations, strain diversity, and transmission dynamics methods12,13 . This provides a more comprehensive and accurate framework for investigating resistance evolution and spread than traditional genotyping methods.
Despite these advances, most existing genomic studies of drug-resistant TB have been geographically limited or focused on specific resistance categories and outbreaks. Even fewer have systematically examined temporal trends in resistance mutations and profiles, leaving gaps in understanding how drug-resistant TB evolves within a country over time. Therefore, this study applies a genomic epidemiology framework to characterize its evolution in India over the past twenty-one years, identifying patterns, trends, and key drivers of resistance. By analysing publicly available genomic data, it aims to increase knowledge on how drug-resistant TB has responded to treatment strategies and public health interventions since prior research has not fully explored these areas in depth. To better comprehend the nuanced interplay of genetic mutations driving resistance and the phylogenetic relationships among drug-resistant strains, a detailed genomic analysis of drug-resistant TB cases identified between 2001 and 2022 in India was conducted. Insights from this analysis can help inform more effective treatment approaches, and have broader implications for global TB control efforts, particularly in high-burden countries. Addressing this knowledge deficit is critical, as a comprehensive genomic perspective can clarify whether rising drug resistance is driven by de novo mutations, clonal expansion, or changes in detection practices. This is further underscored in India, where understanding lineage-specific resistance patterns is essential given the presence of highly transmissible M. tuberculosis lineages.
Methodology
This study is a retrospective analysis on publicly available genomic sequences of M. tuberculosis that were isolated over the past twenty-one years from India to determine changes in resistance profiles, differences in resistance mutations, and how they have evolved. In addition, the study also examines the relationship between global treatment policies and observed resistance patterns, conducting an evaluation on a national scale.
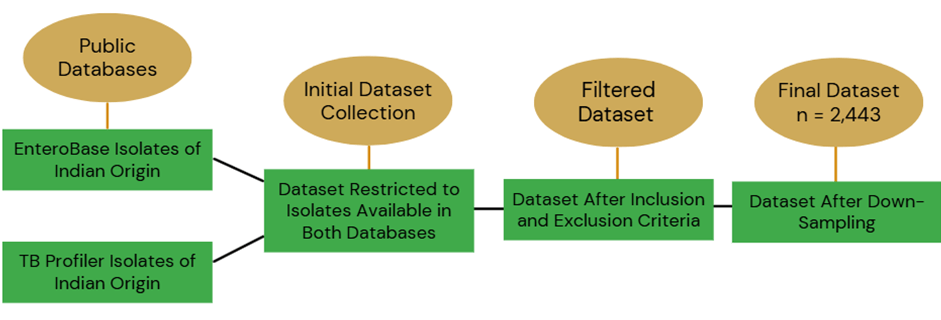
For this research, a set of clinical M. tuberculosis isolates with high-quality WGS data was curated, forming a global dataset using public databases, such as the National Centre for Biotechnology Information (NCBI) and EnteroBase (https://enterobase.warwick.ac.uk/). The dataset for resistance mechanisms analysis was generated based on specific strain inclusion and exclusion criteria to ensure data reliability. Only isolates that were present in both the Enterobase and TB Profiler14,15 databases were included to confirm that the data for each isolate can be cross verified. The filters applied included: Species = “Mycobacterium tuberculosis“, Country = “India”, and Collection date ≥2001 and ≤ 2022. All isolates with complete sequencing runs and publicly available accession numbers were retained as presented in Figure 1.
Whole-genome sequencing data had to meet a minimum quality threshold, with a median depth of at least 30× and a GC content of 65%, to ensure accurate variant detection. The dataset was limited to isolates collected between 2001 and 2022 and originating specifically from India. Isolates were defined as being of Indian origin if the country of isolation was recorded as India in the associated metadata. Sequences lacking clear geographic metadata or with ambiguous country annotations were excluded from the analysis. Both drug-susceptible and drug-resistant strains were selected, encompassing a range from mono-resistant to multidrug-resistant and pre-extensively drug-resistant tuberculosis. Low-quality or incomplete sequences were excluded to maintain dataset integrity.
Sequencing data were processed by using appropriate bioinformatics tools and pipelines, including FastQC for quality control and TB-Profiler for both lineage identification and drug resistance profiling. Bioinformatic operations were conducted on the Galaxy web platform16. Resistance-associated mutations were identified using TB-Profiler (v4.1.1) and validated against known resistance databases. This tool was additionally used in determining different lineages. It aligns the input reads to the M. tuberculosis H37Rv reference genome using BWA-MEM as well as identifies variants associated with drug resistance to then generate a report with all the required information. The link to the complete list of accessions for all 2,443 isolates is available at: raw.githubusercontent.com/KashviT007/mtb-India-resistance-dataset/refs/heads/main/MTB_India_2001_2022_Accession_List.csv
Drugs assessed were first-line (including Isoniazid, Rifampicin, Ethambutol, Pyrazinamide), Streptomycin, fluoroquinolones (including Moxifloxacin, Levofloxacin), Bedaquiline, Linezolid, second-line injectables (including Kanamycin, Amikacin, Capreomycin), Ethionamide, and Para-aminosalicylic acid. Notably, as Bedaquiline and Linezolid are relatively newer and clinically critical drugs for the management of drug-resistant tuberculosis, resistance to these agents was assessed using a targeted genomic approach. Resistance was determined using TB-Profiler, which identifies mutations in specific loci known to confer resistance. For Bedaquiline, the loci analysed were atpE, Rv0678 (mmpR5), and occasionally pepQ. For Linezolid, the loci included rplC and rrl. Detected mutations were cross-referenced against TB-Profiler’s curated resistance database, derived from the WHO Catalogue of Mutations in M. tuberculosis.
Data availability varied substantially across the study period. In earlier years, particularly in the early 2000s, fewer than 500 Indian M. tuberculosis isolates meeting the inclusion criteria were available in public databases; in these cases, all eligible isolates were included. In contrast, during 2019–2022, the number of available isolates exceeded 1,000. To prevent disproportionate weighting of these high-density years and to ensure balanced temporal representation across the 22-year period, a maximum of 500 isolates per year was imposed for years with extensive data availability. This threshold was selected as a pragmatic compromise that preserves sufficient statistical power to capture resistance patterns and lineage diversity within each year, while preventing years with disproportionately high sequencing activity from dominating longitudinal analyses. To reduce sampling bias and ensure that samples were chosen randomly, the following method was used: First, all samples from the specific year were obtained by filtering for that year. Then, a new column was created in which Excel’s random (=RAND ()) function was used to assign an arbitrary number to each of the filtered rows. After generating the random numbers, the values were pasted in the column to freeze them so they wouldn’t change upon recalculation. The filtered rows were then sorted in ascending order based on the random number column. Finally, the top 500 rows from the sorted list were selected as the random subsample.
Ethical
Since all genomic data analysed was publicly available, no ethical approval was required for this study.
Results
Between 2001 and 2022, a total of 2,443 TB cases were analysed in the dataset, with an increasing rate of drug-resistant tuberculosis subtypes over the two decades. The most common resistance to M.tuberculosis is Pre-XDR TB, which accounted for 30.5% of all recorded cases (n=746). This was then followed by MDR-TB at 15.2% (n=371), HR-TB at 7.2% (n = 176), XDR-TB at 3.9% (n = 96), and RR-TB at 1.8% (n = 44). Although drug-sensitive TB declined in proportion over time, it remained the most frequent, at 37.2% (n=908). To formally assess this trend, a chi-square test for trend was performed, with study years grouped into three ordered periods: 2001–2009, 2011–2016, and 2017–2022. The analysis demonstrated a statistically significant decline in the proportion of drug-sensitive TB across these periods (χ² trend =55.72, p <0.0001) indicating that this decrease is unlikely to have occurred by chance.
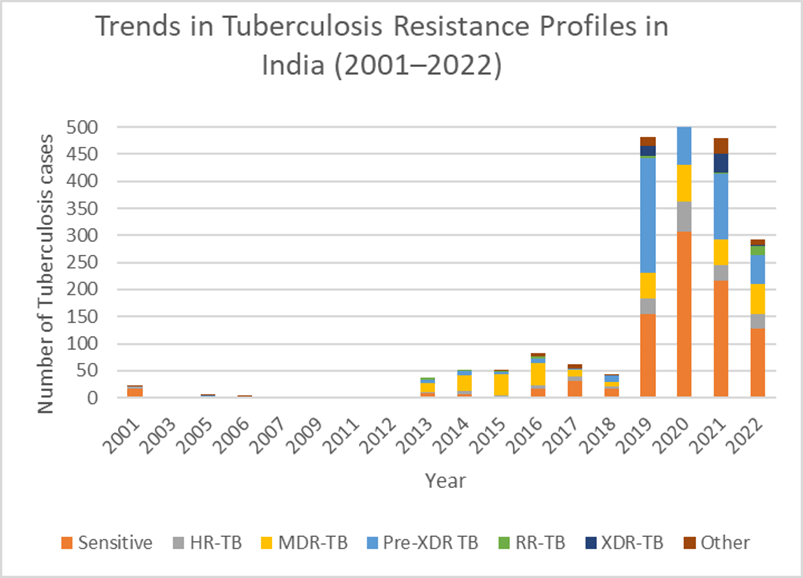
During this analysis, temporal trends were also identified. During the first decade, between 2001 and 2012, drug-resistant TB cases were occasional and infrequent with HR-TB and MDR-TB appearing sporadically, while most cases remained drug-sensitive. However, the year 2013 marked a noticeable increase in MDR-TB cases, alongside the emergence of Pre-XDR-TB and its similar increase. However, sensitive TB began to decline proportionally: from 24.3% (95% CI:10.5–38.1%) in 2013 to 19.5% (95% CI:10.9–28.1%) in 2016. The period which saw a dramatic escalation was 2017 to 2020. In 2019, Pre-XDR-TB cases spiked to 212, representing nearly half (44.1% (95% CI: 40.0% to 48.5%)) of all TB cases that year. MDR-TB remained high, and XDR-TB was noticed for the first time in the dataset. While sensitive TB still accounted for a large proportion, its dominance diminished as resistant forms accelerated. Notably, HR-TB cases additionally rose sharply to 29 in 2019 and 56 in 2020. Similarly, a chi-square test of association was performed comparing HR-TB cases in the pre-2019 period with the 2019–2020 period. The analysis confirmed that HR-TB cases were more likely to occur in 2019–2020 than in earlier years (χ² = 4.06, p = 0.044). More recently, more complex resistant profiles have emerged. Pre-XDR-TB remained significant along with MDR-TB remaining consistent, and HR-TB persisting at high levels, suggesting continued transmission of resistance to first-line drugs. A decline was observed in XDR-TB, dropping to just 3 cases in 2022 and sensitive TB cases, falling to 128 in 2022 (χ² =8.10, p= 0.0044).
Figure 2 illustrates the evolving epidemiological profile. Until 2012, drug-sensitive TB comprised over 56% of cases annually. By contrast, 2019 onwards, other drug-resistant forms, especially Pre-XDR-TB became more prevalent, overtaking all other types of resistance, with this increase across the three study periods being statistically significant (χ² trend=53.32, p < 0.0001). The emergence of XDR-TB, although smaller, marks a concerning escalation in resistance severity. Finally, the proportion of MDR-TB remained steady throughout the 2013–2022 window, indicating persistent challenges in controlling or reducing transmission.
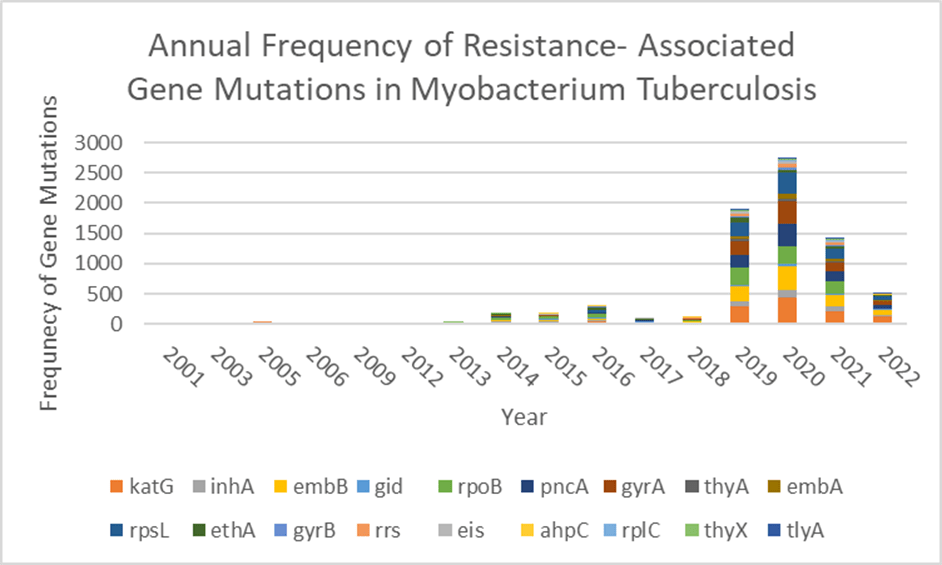
In parallel with the shifting epidemiological profiles of TB, the genetic basis of resistance also revealed significant trends. The most frequently observed resistance associated gene was katG (n=1,220), associated with isoniazid resistance, a key marker for HR-TB when mutated alone. This was closely followed by embB (n=969), responsible for ethambutol resistance, and rpoB (n=946), often linked with rifampicin resistance, a key marker for MDR and RR-TB. Subsequently, rpsL (n=887), often linked to streptomycin resistance, and pncA (n=863), involved in pyrazinamide resistance, were also frequently detected.
Other notable genes included gyrA (n=854), a marker for fluoroquinolone resistance, and a defining feature of Pre-XDR TB. InhA (n = 385) also contributed to ethionamide resistance. Though less common, ethA (n=293), embA (n=211), gid (n=207), rrs (n=170), eis (n=117), thyA (n=85), gyrB (n=65), thyX (n= 63), rplC (n=33), tlyA (n=17), and ahpC (n=16) were also detected, reflecting the growing complexity and diversity of resistance mechanisms.
The prevalence of these genes rose sharply after 2013, coinciding with the increase in Pre-XDR and MDR-TB cases. This could suggest the clonal transmission of resistant strains, as evidenced by the recurring presence of identical resistance-associated mutations. Between 2013 and 2022, katG, rpoB and embB, depicted a consistent and substantial rise in frequency throughout, depicted in Figure 3. Notably, katG mutations were nearly absent prior to 2012 but rose to a peak of 429 cases in 2020 alone. The increase in occurrence of ethA, gyrB, ahpC, rplC, thyX and tlyA mutations from 2019 onwards further highlights the growing burden of multidrug-resistant TB, expanding drug resistance beyond traditional first and second-line agents, underscoring a concerning trend in the evolving genetic landscape of drug-resistant tuberculosis.
| Lineage 1 | Lineage 2 | Lineage 3 | Lineage 4 | |
| 2003 | 1 | 0 | 0 | 0 |
| 2005 | 3 | 2 | 1 | 2 |
| 2006 | 0 | 2 | 1 | 1 |
| 2007 | 0 | 0 | 2 | 0 |
| 2009 | 1 | 0 | 0 | 1 |
| 2011 | 0 | 0 | 1 | 0 |
| 2012 | 0 | 0 | 2 | 0 |
| 2013 | 21 | 9 | 6 | 2 |
| 2014 | 30 | 8 | 7 | 8 |
| 2015 | 30 | 6 | 13 | 3 |
| 2016 | 42 | 9 | 16 | 14 |
| 2017 | 46 | 7 | 6 | 6 |
| 2018 | 14 | 10 | 17 | 7 |
| 2019 | 76 | 212 | 138 | 66 |
| 2020 | 131 | 347 | 212 | 132 |
| 2021 | 113 | 185 | 160 | 84 |
| 2022 | 102 | 54 | 91 | 50 |
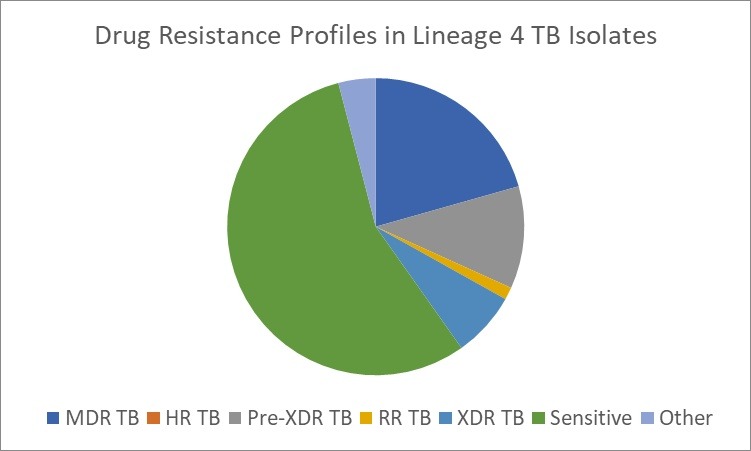
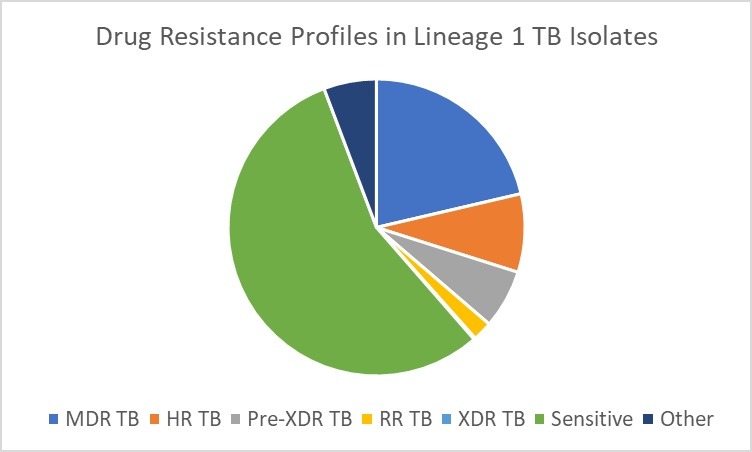
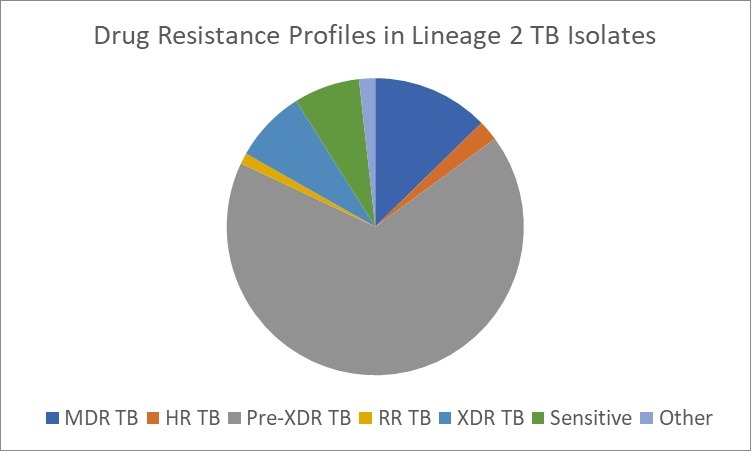
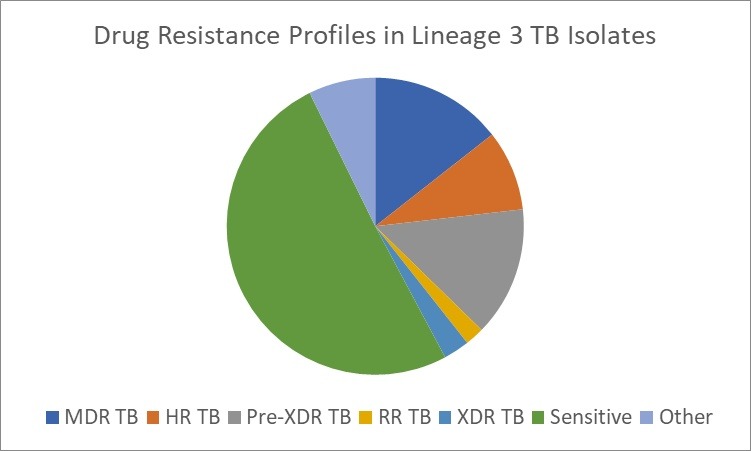
These trends are also reflected in the distribution of tuberculosis throughout the years summarized by Table 1. The earlier decades predominantly featured isolates from the Indo-Oceanic (Lineage 1) lineage, with occasional representation from the Beijing lineage (Lineage 2) and the Central Asian lineage (Lineage 3). However, from 2012 onwards, a marked expansion in both the number and variety of lineages becomes evident. The Beijing lineage began to show consistent prevalence, peaking in 2020 with 347 isolates (χ² = 3.96, p = 0.047). This accounted for approximately 40.8% (95% CI:37.5–44.1%) of all lineage 2 cases across the study period. Among its sub lineages, lineage 2.2.1 emerged as the most frequent, representing over 79% (95% CI:76.3–81.7%) of all Beijing isolates. Other sub-lineages such as 2.2.1.1, 2.2.1.2, 2.2.2, and 2.2 were observed at lower frequencies, suggesting occasional diversification but with limited transmission success. Within this lineage, Pre-XDR TB was the most common resistance profile, present in nearly two thirds of all cases, as seen in Figure 4b. This was followed closely by MDR-TB (12.4% (95% CI:11.3–13.5%) ) and XDR-TB (7% 95% CI:5.29–8.71%) ) which still represented a significant clinical concern, particularly in the peak years of 2019–2021.
Similarly, lineage 3, a historically dominant lineage, portrayed a surge in 2020 (n= 212) and 2021 (n=160), with increasing proportions of HR, MDR, and RR-TB displayed in Figure 4c. When compared across all four major lineages, the distribution of Pre-XDR TB differed significantly by lineage (χ² = 23.42, df = 3, p < 0.0001), indicating a disproportionate burden of drug resistance within Lineage 2. In contrast, the Indo-Oceanic lineage was the most prevalent lineage during the earlier phase of the post-2012 expansion, particularly between 2013 and 2017 (χ² = 174.32, p < 0.0001). Although its relative contribution declined during the peak years as Lineages 2 and 3 expanded more sharply, it still demonstrated one of the largest proportional increases in later years, showing the second-greatest rise between 2019 and 2020, surpassed only by the Euro-American lineage (Lineage 4), which doubled over the same period. This pattern suggests that while the Indo-Oceanic lineage may be less competitively dominant during periods of intense expansion of highly resistant lineages, it remains epidemiologically resilient, continuing to expand alongside, but not at the pace of, more genomically adaptive strains. This is further exemplified by the relatively lower burden of drug resistance in Lineage 1 isolates, as evident in Figure 4a.
Furthermore, Lineage 4 maintained a steady presence across the study period. While not as strongly associated with drug resistance as lineage 2, highlighted in Figure 4d, it contributed significantly to the overall TB burden, particularly through its genetically diverse sub lineages such as 4.2, 4.4, 4.5, 4.8, or 4.9, highlighting the expanding genomic diversity of resistant strains. This diversification, particularly from 2013 onward, aligns with the timeline of increased resistance gene detection and suggests not only a wider pool of circulating strains but also enhanced adaptive potential in response to drug pressure, complicating treatment and control efforts.
Discussion
Rising Drug-Resistant Tuberculosis Trends and Drivers
The findings highlight a concerning shift in the tuberculosis landscape, marked by a substantial rise in drug-resistant forms and a corresponding decline in drug-sensitive cases. This underscores the growing challenge in disease control and treatment efficacy in the country. A particularly alarming finding is the dominance of Pre-XDR TB, which accounted for a considerable proportion of cases, surpassing MDR-TB and indicating widespread resistance to both first and second-line anti-tuberculosis drugs. This result is consistent with multiple published sources containing global analyses. For instance, Goyal et al17, which identified increasing fluoroquinolone resistance in India as an early indicator of growing Pre-XDR TB prevalence. Similarly, the WHO Global TB Reports in 2023 alone, 63,929 MDR/RR-TB cases were reported, including 11,749 Pre-XDR-TB and 114 XDR-TB patients, suggesting both improved diagnostics and rising transmission18.
This sharp escalation can be attributed to a combination of biological, clinical, and programmatic factors that converged during this period. The overuse and misuse of fluoroquinolones, both in TB and non-TB settings (e.g., in general respiratory infections), played a central role in accelerating this shift19. The easy availability and indiscriminate use of these antibiotics for common, non-TB conditions are among the most significant risk factors contributing to the development of resistance to these critical second-line drugs20,21,22,23,24. Studies also report that as short as 13 days of exposure to fluoroquinolones can lead to the development of resistance among treatment naïve and treatment-experienced TB patients25. Case studies further suggest that even short-term monotherapy with fluoroquinolones can lead to the acquisition of resistance in M. tuberculosis26,27. India’s disproportionately large share of the private-sector TB drug market (accounting for 63% of second-line drug consumption) has also contributed substantially to this problem28. Notably, fluoroquinolones dominate this market, making up 96% of second-line drug volume compared to just 1% for injectables. This imbalance suggests they are frequently used either as monotherapy or as inappropriate add-ons to first-line regimens, rather than within structured second-line treatment protocols. Such irrational and inconsistent use of second-line drugs in the private sector has led to inadequate treatment, thereby worsening outcomes and driving the further emergence of resistance29. This further underscores the need to use these drugs judiciously and regulate their availability stringently.
The increasing burden of drug-resistant TB has led to the rapid expansion of Programmatic Management of Drug-Resistant Tuberculosis (PMDT) services30. Endorsed by the WHO in 2002, India adopted the PMDT services in 2007, and complete geographic coverage was achieved in 2013. Since then, fluoroquinolones have formed the backbone of second-line treatment regimens for MDR-TB. While this marked an important step in expanding care, the large-scale and prolonged use of fluoroquinolones created strong selective pressure for the emergence of fluoroquinolone-resistant strains. This contributed to the emergence of strains harboring gyrA and gyrB mutations, which are defining features of Pre-XDR TB. A study analysing 152 MDR TB patients in New Delhi detected 27 mutations in the gyrA genes of 25 strains31.
Traditionally, culture-based methods have been used for determining antimicrobial susceptibility. These procedures usually take several weeks. Thus, a patient possessing a drug-resistant M. tuberculosis strain may continue to spread the infection undetected in the community32. Molecular methods, however, have decreased the time for the detection of drug resistance33. Among these methods, reverse hybridization or line probe assays (LPAs) have been implemented as part of national TB programs in high-burden countries for rapid detection34. Therefore, the introduction of practices, such as universal Drug Susceptibility Testing (DST) under the National TB Elimination Programme (NTEP), significantly improved the detection of fluoroquinolone resistance, revealing its true burden. This coincided with a genuine increase in biological spread, driven by incomplete treatment and poor adherence.
While these factors have contributed to rising resistance across the spectrum, from HR-TB to XDR-TB, the disproportionate increase in Pre-XDR TB highlights the growing challenge of second-line drug failure. This form of resistance not only signals treatment escalation but also poses a greater threat to control strategies due to limited therapeutic options and higher transmission potential.
The genomic data in this study also identified katG and rpoB as the most frequently detected resistance mutations. The surge in these mutations is consistent with India’s increased use of molecular diagnostics, which made it easier to identify mutations previously undetectable by phenotypic methods. However, the increasing frequency of these mutations also reflects ongoing transmission of resistant strains within communities, especially in urban regions where treatment interruptions, inadequate regimens, and poor follow-up are common35. For instance, a Gujarat cohort study reported that 19.2% of MDR-TB patients were lost to follow-up, with 87% of them discontinuing treatment within the first six months, underscoring critical gaps in treatment retention36. Most people with TB are cured by a 6-month treatment regimen that is provided to patients with adequate support. Inappropriate or incorrect use of TB drugs, ineffective formulations (such as single drugs, poor quality medicines, or improper storage conditions), and premature treatment interruption can cause drug resistance. This can then be transmitted, especially in crowded settings such as prisons and hospitals37. Furthermore, the emergence of rarer mutations such as ethA, tlyA, gyrB, and thyX, from 2019 onwards, suggests a growing threat posed by complex resistance patterns. This reflects a broadening of the resistance spectrum beyond traditional first- and second-line drugs. This not only makes treatment more difficult but also raises concerns about the potential for these strains to evolve into extensively drug-resistant forms if not effectively managed. This pattern reflects the natural evolutionary response of M. tuberculosis, marked by its high adaptability to ongoing selective pressures and incomplete eradication. As resistant strains continue to circulate and acquire new mutations, the resistance landscape becomes increasingly complex37.
Lineage analysis further contextualizes these trends. The rise of Lineage 2 strains in the dataset adds an important evolutionary layer. The consistent presence of this lineage across the years can be linked to the active transmission in the community. A genomic epidemiology study in Mumbai revealed that three dominant Lineage 2 clusters accounted for nearly 50% of Pre‑XDR and 40% of XDR-TB cases; these clusters exhibited high fluoroquinolone resistance rates (81–91%), and Pre‑XDR/XDR strains showed a 79% clustering rate38. This further highlights the strain’s rapid adaptability, suggesting that it evolves under drug pressure more swiftly than other lineages. Its enhanced survival traits, such as resistance development, immune evasion, and metabolic flexibility, enable it to outcompete other strains, particularly in environments marked by inconsistent treatment or high antibiotic exposure39. Their rise in India also coincides with urbanization, population mobility, and regional outbreaks, particularly in cities like Mumbai and Delhi where resistant TB is highly prevalent. This supports the theory that the Beijing lineage has not only spread due to intrinsic factors but has also been amplified by social and healthcare infrastructure challenges40. Therefore, initiatives like the “TB Free Slums” aim to eradicate TB by focusing on high-risk areas, including slums and unauthorized settlements. This program involves systematic screening and health evaluations to identify and treat TB cases early, addressing the challenges posed by poor living conditions and inadequate healthcare infrastructure41.
While Lineage 1, Lineage 3, and Lineage 4 remained present across the study period, their resistance profiles were less severe and more variable. The increasing representation of genetically diverse sub lineages from several lineages, like lineage 4 and lineage 2, indicates growing genomic diversity, which complicates efforts to generalize treatment regimens and highlights the need for lineage-informed clinical decisions. More importantly, this indicates that India’s TB epidemic is no longer dominated by a single genetic profile, but rather shaped by a broad array of adaptive, resistant strains42, additionally complicating diagnostics and control measures.
National Policy Response and Programmatic Strengthening
Despite the growing difficulty in treating TB, India has introduced targeted policies to address and reduce the emergence of drug-resistant forms of the disease. It has taken a comprehensive approach through the National Strategic Plan (NSP) 2020–2025, which outlines ambitious goals to eliminate TB by 2025. This strategy is designed to accelerate national progress by drawing on successful elements from past public health campaigns, most notably the Polio eradication program43. The plan calls for a national TB elimination campaign, with a strengthened focus on workforce development and expanded engagement with the private sector. It emphasizes a shift from passive community awareness to active participation and ownership, alongside investments in real-time surveillance systems and access to advanced diagnostic tools. The NSP also prioritizes comprehensive patient support throughout treatment, active case-finding efforts in high-risk populations, and the implementation of shorter preventive treatment regimens for vulnerable groups such as household contacts, children, and people living with HIV44.
In 2021, the Ministry of Health and Family Welfare, in collaboration with development partners, launched the “TB Mukt Bharat Abhiyaan” as a flagship mission under NSP 2020–2025. This initiative emphasizes identifying all TB patients, including those seeking care in the private sector and among undiagnosed high-risk populations, and ensuring they receive timely diagnosis and treatment. As a result of this renewed focus, there has been a significant expansion in the deployment of advanced diagnostic tools across the country. Technologies such as CBNAAT (GeneXpert), TrueNAT, and LPAs have been scaled up nationwide, enabling rapid and accurate detection of drug-resistant strains, including rifampicin and fluoroquinolone resistance45. These tools not only reduce diagnostic delays but also improve the precision of treatment initiation, particularly for patients with multidrug-resistant TB.
Within the context of this study, the nationwide expansion of rapid molecular diagnostics likely contributed to the increased detection of rifampicin and fluoroquinolone-associated mutations observed in later years. The temporal rise in rpoB and gyrA mutations, particularly post-2013, aligns with this diagnostic scale-up, suggesting that part of the apparent surge in resistant isolates reflects improved case ascertainment and resolution of previously undetected second-line resistance rather than solely the emergence of new mutations. This enhanced detection is further reflected in the reclassification of isolates into higher resistance categories, notably Pre-XDR TB, and may also indicate a measurable genomic restructuring of circulating strains coinciding with the wider deployment of second-line therapies.
To support national efforts, WHO-India launched Project GATIMAN to strengthen technical assistance across states, focusing on public-private partnerships, drug-resistant TB management, laboratory capacity, surveillance, and communication. Additionally, 100 high-burden and hard-to-reach districts were identified for intensified intervention46. In fact, a key pillar of India’s TB response is Active Case Finding (ACF), which has been implemented nationwide since 2017 and expanded in 2021 to include high-risk groups such as individuals with chronic illnesses and those who are immunocompromised. During the COVID-19 pandemic, targeted ACF campaigns also screened patients with influenza-like and severe respiratory symptoms47. By the end of 2021, ACF efforts had reached over 22 million people, resulting in the testing of 1.75 million samples and the detection of more than 73,000 additional TB cases. The expansion of ACF may also explain the increased identification of genetically diverse sub lineages in recent years within this dataset, as intensified community screening broadens sampling and reduces under-detection of resistant clusters, rather than reflecting a sudden evolutionary shift.
India has also implemented several initiatives to improve treatment adherence and patient support. These include financial incentives such as ₹1000 for completing treatment in drug-sensitive TB cases, ₹5000 for treatment supporters managing drug-resistant cases, and ₹1000 per notification for private providers reporting TB patients. Additionally, patients in tribal areas receive ₹750 in travel support. Under the Nikshay Poshan Yojana, all notified TB patients are provided ₹500 per month as a direct benefit transfer to support nutritional needs during treatment48. To strengthen surveillance and case management, the government launched Nikshay, a national web-based platform developed by the Central TB Division and the National Informatics Centre. Introduced in 2012, Nikshay enables real-time tracking of TB cases and outcomes, and reporting through the platform is now mandatory for both public and private healthcare providers49. The integration of Nikshay has likely improved linkage between diagnostic confirmation and treatment initiation. In parallel with enabling earlier detection of potential resistance, it limits onward transmission. This claim is consistent with the dataset, where the proportion of drug-sensitive isolates increased from approximately 37% in 2020 to nearly 44% in 2022, indicating strengthened control of resistance.
The progress in India’s TB response is evident in the significant reduction in disease burden over the years. As outlined by WHO Global Tuberculosis Reports, TB incidence per 100,000 population decreased from 322 (95% CI: 110-644) in 2000 to 195 (164-228) in 2023. Similarly, TB-related mortality dropped from 87 (95% CI: 67-109) per 100,000 population in 2000 to 22 (95% CI: 16-30) in 2023 (Figure 5). In absolute terms, annual TB-related deaths have been reduced by two-thirds, from approximately 1 million deaths to one-third of that figure50.
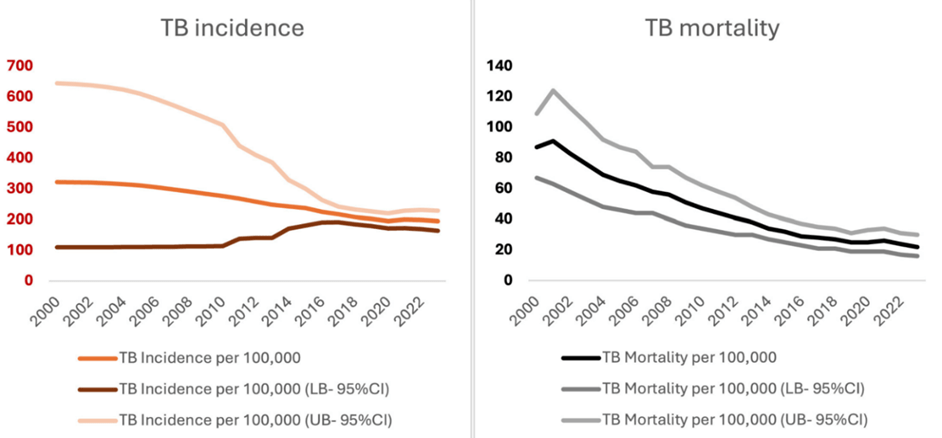
However, while these results are promising, they highlight the need for sustained and consistent implementation. Continued investment in infrastructure, early detection, patient support, and surveillance systems is essential to maintain momentum. As drug-resistant TB remains a major threat, these efforts must not only continue but also adapt to evolving challenges to ensure long-term success in eliminating TB from India.
Study Constraints and Future Research Implications
Although this study offers valuable insights, several limitations must be acknowledged. The dataset, although large, is dependent on publicly available genomes, which may not equally represent all regions or time periods. This reliance may also introduce selection bias, as such repositories are often enriched for drug-resistant isolates and strains originating from referral or research centres rather than routine surveillance. Additionally, isolates subjected to WGS are more likely to represent clinically complex or treatment-refractory cases. While inclusion of both susceptible and resistant isolates mitigates this effect to some extent, the dataset may not fully reflect the true population-level prevalence of resistance in India. Phenotypic confirmation of resistance was not consistently available, which may affect mutation interpretation. Moreover, patient-level clinical data, including treatment history and adherence were unavailable, limiting causal conclusions about how resistance emerged or progressed and restricting a more nuanced understanding of treatment outcomes. As such, the findings should be interpreted in the context of these data constraints with emphasis placed on resistance patterns and evolutionary trends rather than on absolute prevalence estimates.
Future work should explore the longitudinal evolution of resistance within individual hosts and investigate host-pathogen interactions, especially for dominant lineages prevalent in India. Such studies will further clarify how genetic variation influences transmissibility, virulence, and resistance acquisition51. Integrating host genetic and immunological data with pathogen genomics could open new avenues for personalized treatment strategies and vaccine development. Regional and spatial analyses of genomic data could identify TB transmission hotspots, helping public health authorities to prioritize areas for intensified active case finding and treatment adherence programs52. This approach would also help in detecting outbreaks early and understanding the spread of resistant strains in urban versus rural settings. Prospective, real-time genomic surveillance integrated into national TB control programs would enable timely detection of emerging resistance mutations and treatment failures, allowing rapid adjustments to therapy guidelines53. Such systems could leverage portable sequencing technologies and advanced bioinformatics pipelines to bring genomic epidemiology closer to the point of care54.
Conclusion
This study offers a comprehensive view of the growing complexity of drug-resistant tuberculosis in India, with a particular focus on the rising prevalence of Pre-XDR TB, the dominant role of Lineage 2, and the emergence of diverse resistance mutations across multiple lineages. These findings underscore how biological adaptability, combined with programmatic gaps such as inconsistent treatment and limited private sector regulation, has accelerated resistance. India has been actively involved in TB control activities for more than 50 years now and has made commendable progress. The country is now better prepared to tackle TB than before However, while these efforts have contributed to measurable reductions in TB incidence and mortality, the ongoing transmission of highly resistant strains, especially in urban and underserved areas, signals the need for sustained and adaptive action. Continued investment in real-time genomic surveillance, stronger treatment adherence mechanisms, and region-specific interventions will be crucial to achieve long-term TB control and meet elimination goals. As drug resistance continues to evolve, India’s success will depend not just on the scale of its response, but on its ability to anticipate and outpace the pathogen’s adaptive potential.
Acknowledgements
I would like to express my deepest gratitude to my mentor, Xiaoliang Ba, for his invaluable guidance, encouragement, and constructive feedback throughout this project. Their expertise and support were instrumental in helping me develop a deeper understanding of tuberculosis and the global efforts to combat it. Additionally, I extend my heartfelt thanks to the Cambridge Centre for International Research (CCIR) team for their invaluable assistance in reviewing my paper and guiding me through the publication process. Finally, I would also like to thank my family for their unwavering encouragement, patience, and commitment throughout this journey. Their belief in me, even during moments of doubt and difficulty, has been my greatest source of strength and motivation, and I dedicate this achievement to them with deep appreciation.
References
- Centers for Disease Control and Prevention. Clinical overview of tuberculosis disease. Atlanta: CDC; (2022). [↩]
- A. Natarajan, P.M. Beena, A. V. Devnikar, S. Mali. A systemic review on tuberculosis. Indian Journal of Tuberculosis. 67, 295-311 (2020). [↩]
- World Health Organization. Global tuberculosis report.Geneva: WHO (2024). [↩]
- Government of India, Ministry of Health and Family Welfare. National Strategic Plan 2016–2023. New Delhi: Government of India(2017). [↩]
- Government of India, Ministry of Health and Family Welfare. National Strategic Plan for Tuberculosis Elimination 2017–2025. New Delhi: Government of India; (2017). [↩]
- S. Mohan , V.Pujari , P.Meshram . Drug resistance patterns and treatment outcomes in DR-TB patients at a tertiary care centre in Mumbai. Indian Journal of Tuberculosis. 71, 510-514 (2024). [↩]
- D. Vishwakarma, A. Gaidhane, S. Sahu, A.S. Rathod. Multi-drug resistance tuberculosis (MDR-TB) challenges in India: A review. DMIHER School of Epidemiology and Public Health. Cureus (2023). [↩]
- S.M. Batt, D.E. Minnikin, G.S. Besra; The thick waxy coat of mycobacteria, a protective layer against antibiotics and the host’s immune system. Biochemical Journal. 477,1983–2006 (2020). [↩]
- S. Shukla, N. Bhardwaj, A. Singh. Drug resistance in Mycobacterium tuberculosis: An evolutionary perspective and its adaptation to the lung microenvironment. The Microbe. 5 (2024). [↩]
- R. Singh, S.P. Dwivedi, U.S. Gaharwar, R. Meena, P. Rajamani, T. Prasad. Recent updates on drug resistance in Mycobacterium tuberculosis. Applied Journal of Microbiology. 128, 1547-1567 (2019). [↩]
- World Health Organization. WHO announces updated definitions of extensively drug-resistant tuberculosis. Geneva: World Health Organization (2021). [↩]
- T. M. Walker, T.A. Kohl, S.V Omar, J. Hedge, C.D. Elias, P. Bradley, Z. Iqbal, S. Feuerriegel, K.E. Niehaus, D. J Wilson, D.A Clifton, G. Kapatai, C.L.C Ip, R. Bowden, F.A. Drobniewski, C. Allix-Béguec, C. Gaudin, J. Parkhill, R. Diel, P. Supply, D.W.Crook, E Grace Smith, A Sarah Walker, N. Ismail, S. Niemann, T.E.A Peto,Whole-genome sequencing for prediction of Mycobacterium tuberculosis drug susceptibility and resistance: a retrospective cohort study.The Lancet Infectious Diseases. 15, 1193-1202 (2019). [↩]
- V. Nikolayevskyy, S. Niemann, R. Anthony, D. van Soolingen, E. Tagliani, C. Ködmön, M.J. van der Werf, D.M. Cirillo.Role and value of whole genome sequencing in studying tuberculosis transmission. Clinical Microbiology and Infection. 25, 1377-1382 (2019). [↩]
- J. E. Phelan, D. M. O’Sullivan, D. Machado. Integrating informatics tools and portable sequencing technology for rapid detection of resistance to anti-tuberculous drugs. Genome Medicine. 11, 41. (2019). [↩]
- F. Coll, R. McNerney, M. D. Preston, J. A. Guerra-Assunção, A. Warry, G. Hill-Cawthorne, K. Mallard, M. Nair, A. Miranda, A. Alves, J. Perdigão, M. Viveiros, I. Portugal, Z. Hasan, R. Hasan, J. R. Glynn, N. Martin, A. Pain, T. G. Clark. Rapid determination of anti-tuberculosis drug resistance from whole-genome sequences. Genome Medicine. 7, 51 (2015). [↩]
- E. Afgan, D. Baker, B. Batut, M. van den Beek, D. Bouvier, M. Čech, J. Chilton, D. Clements, N. Coraor, B. A. Grüning, A. Guerler, J. Hillman-Jackson, S. Hiltemann, V. Jalili, H. Rasche, N. Soranzo, J. Goecks, J. Taylor, A. Nekrutenko, D. Blankenberg. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: Nucleic Acids Research. 46, 537–544 (2018). [↩]
- V. Goyal, V. Kadam, P. Narang, V. Singh. Prevalence of drug-resistant pulmonary tuberculosis in India: systematic review and meta-analysis. BMC Public Health. 17, 817 (2017). [↩]
- Central TB Division, Ministry of Health and Family Welfare. National Guidelines for Management of Drug-Resistant Tuberculosis (DR-TB). New Delhi: Government of India; (2025). [↩]
- N. Selvakumar, V. Kumar, S. Balaji, S. Prabuseenivasan, R. Radhakrishnan, G. Sekar, V. Chandrasekaran, T. Kannan, A. Thomas, S. Arunagiri, P. Dewan, S. Swaminathan. High rates of ofloxacin resistance in Mycobacterium tuberculosis among both new and previously treated patients in Tamil Nadu, South India. PLoS One.10 (2015). [↩]
- S.P. Rai, D. Bhattacharyya, M. Kashyap. Pattern of initial drug resistance and its impact on short course chemotherapy of pulmonary tuberculosis. Lung India.24, 51–53 (2007) [↩]
- C. Porwal, A. Kaushik, N. Makkar, J. N. Banavaliker, M. Hanif, R. Singla, A. K. Bhatnagar, D. Behera, J. N. Pande, U. B. Singh. Incidence and risk factors for extensively drug-resistant tuberculosis in Delhi region. PLoS One. 8 (2013 [↩]
- A. Jain, P. Dixit, R. Pre-XDR & XDR in MDR and Ofloxacin and Kanamycin resistance in non-MDR Mycobacterium tuberculosis isolates. Tuberculosis.92, 404–406 (2012) [↩]
- N. Hemvani, D. S. Chitnis, G. C. Bhatia, N. Sharma. Drug resistance among tubercle bacilli from pulmonary tuberculosis cases in central India. Indian Journal of Medical Sciences. 55,382–392 (2001 [↩]
- C.N. Paramasivan, F. Rehman, F. Wares, N. Sundar Mohan, S. Sundar, S. Devi. First- and second-line drug resistance patterns among previously treated tuberculosis patients in India. The International Journal of Tuberculosis and Lung Disease. 14, 243–246 (2010). [↩]
- A. S. Insburg, S. C. Woolwine, N. Hooper, W. H. Benjamin Jr, W. R. Bishai, S. E. Dorman. The rapid development of fluoroquinolone resistance in M. tuberculosis. New England Journal of Medicine.349, 1977–1978 (2003). [↩]
- S. Rajasekaran, C. Chandrasekar, A. Mahilmaran, K. Kanakaraj, D. S. Karthikeyan, J. Suriakumar. HIV coinfection among multidrug resistant and extensively drug resistant tuberculosis patients–a trend. Journal of Indian Medical Association. 107, 281–282 (2009 [↩]
- C. K. Mlambo, R. M. Warren, X. Poswa, T. C. Victor, A. G. Duse, E. Marais. Genotypic diversity of extensively drug-resistant tuberculosis (XDR-TB) in South Africa. International Journal of Tuberculosis and Lung Disease.12, 99–104 (2008). [↩]
- W. A. Wells, C. F. Ge, N. Patel, T. Oh, E. Gardiner, M. E. Kimerling. Size and usage patterns of private TB drug markets in the high burden countries. PLoS.6 (2011). [↩]
- D. Agrawal, Z. F. Udwadia, C. Rodriguez, A. Mehta. Increasing incidence of fluoroquinolone-resistant Mycobacterium tuberculosis in Mumbai. India. International Journal of Tuberculosis and Lung Disease.13, 79–83 (2009). [↩]
- A. Khanna, R. Saha, N. Ahmad.National TB elimination programme – What has changed. Indian Journal Medical Microbiology. 42, 103-107 (2023). [↩]
- R. Singhal, P. R. Reynolds, J. L. Marola, L. E. Epperson, J. Arora, R. Sarin, V. P. Myneedu, M. Strong, M. Salfinger. Sequence Analysis of Fluoroquinolone Resistance-Associated Genes gyrA and gyrB in Clinical Mycobacterium tuberculosis Isolates from Patients Suspected of Having Multidrug-Resistant Tuberculosis in New Delhi, India. Journal of Clinical Microbiology. 54 (2019). [↩]
- K. Jabeen, S. Shakoor, R. Hasan. Fluoroquinolone-resistant tuberculosis: implications in settings with weak healthcare systems. International Journal of Infectious Diseases.32, 118–123 (2015). [↩]
- D. I. Ling, A. A. Zwerling, M. Pai. GenoType MTBDR assays for the diagnosis of multidrug-resistant tuberculosis: a meta-analysis.European Respiratory Journal.32, 1165–1174 (2008). [↩]
- R. Singhal, V. P. Myneedu, J. Arora, N. Singh, G. C. Sah, R. Sarin. Detection of multi-drug resistance and characterization of mutations in Mycobacterium tuberculosis isolates from North-Eastern states of India using GenoType MTBDRplus assay. Indian Journal of Medical Research. 140, 501–506 (2014). [↩]
- World Health Organization. Tuberculosis: Multidrug-resistant (MDR-TB) or rifampicin-resistant TB (RR-TB). Geneva: World Health Organization (2024). [↩]
- K.S. Shringarpure, P. Isaakidis, K.D. Sagili, R.K. Baxi. Loss‑to‑follow‑up on multidrug‑resistant tuberculosis treatment in Gujarat, India: the WHEN and WHO of it. PLoS ONE.10 (2015). [↩]
- R. Singh, S. P. Dwivedi, U. S. Gaharwar, R. Meena, P. Rajamani, T. Prasad. Recent updates on drug resistance in Mycobacterium tuberculosis. Journal of Applied Microbiology. 128, 1229-1241 (2019). [↩] [↩]
- V. Dreyer, A. Mandal, P. Dev, S. M. Gagneux, R. Diel, S. Niemann, J. E. Phelan, T. Wirth. High fluoroquinolone resistance proportions among multidrug-resistant tuberculosis driven by dominant L2 Mycobacterium tuberculosis clones in the Mumbai Metropolitan Region. Genome Medicine. 14, 95 (2022). [↩]
- S. K. Shanmugam, N. Kumar, T. Sembulingam, S. B. Ramalingam, A. Selvaraj, U. Rajendhiran, S. Solaiyappan, S. P. Tripathy, M. Natrajan, P. Chandrasekaran, S. Swaminathan, J. Parkhill, S. J. Peacock, U. D. K. Ranganathan. Mycobacterium tuberculosis Lineages Associated with Mutations and Drug Resistance in Isolates from India. Microbiol Spectrum.10 (2022). [↩]
- M. Merker, C. Blin, S. Mona, N. Duforet-Frebourg, S. Lecher, E. Willery, M. G. Blum, S. Rüsch-Gerdes, I. Mokrousov, E. Aleksic, C. Allix-Béguec, A. Antierens, E. Augustynowicz-Kopeć, M. Ballif, F. Barletta, H. P. Beck, C. E. Barry III, M. Bonnet, E. Borroni, I. Campos-Herrero, D. Cirillo, H. Cox, S. Crowe, V. Crudu, R. Diel, F. Drobniewski, M. Fauville-Dufaux, S. Gagneux, S. Ghebremichael, M. Hanekom, S. Hoffner, W. W. Jiao, S. Kalon, T. A. Kohl, I. Kontsevaya, T. Lillebæk, S. Maeda, V. Nikolayevskyy, M. Rasmussen, N. Rastogi, S. Samper, E. Sanchez-Padilla, B. Savic, I. C. Shamputa, A. Shen, L. H. Sng, P. Stakenas, K. Toit, F. Varaine, D. Vukovic, C. Wahl, R. Warren, P. Supply, S. Niemann, T. Wirth. . Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nature Genetics. 47, 242-249 (2015). [↩]
- A. Jaiswal. City goes all out to eradicate TB, focus on slums. Times of India. New Delhi; Times of India (2025). [↩]
- S. K. Shanmugam, N. Kumar, T. Sembulingam, S. B. Ramalingam, A. Selvaraj, U. Rajendhiran, S. Solaiyappan, S. P. Tripathy, M. Natrajan, P. Chandrasekaran, S. Swaminathan, J. Parkhill, S. J. Peacock, U. D. K. Ranganathan,Mycobacterium tuberculosis Lineages Associated with Mutations and Drug Resistance in Isolates from India. Microbiology Spectrum. 1o (2022). [↩]
- E.N.Bureau. City goes all out to eradicate TB, focus on slums. Express Healthcare (2018). [↩]
- M. Pai, S. Swaminathan. India is well placed to scale innovations in tuberculosis diagnostics. Indian Journal of Medical Research.157,114-117 (2023). [↩]
- Government of India, Ministry of Health and Family Welfare. National Strategic Plan 2016–2023. New Delhi: Government of India (2017). [↩]
- World Health Organization, India. States and UTs accelerate action to end TB by 2025. New Delhi: WHO India (2021). [↩]
- Central TB Division, Ministry of Health and Family Welfare, Government of India. India TB Report 2022. New Delhi: Government of India (2022). [↩]
- K. Jeyashree, P. Shanmugasundaram, D. Shanmugasundaram, S. L. P. G, J. W. V. Thangaraj, S. T. S, S. Pandey, S. Ramasamy, R. Sharma, S. Arunachalam, V. Shah, V. Janagaraj, S. S. Sundari, J. Chadwick, H. D. Shewade, A. Chowdhury, S. Iyer, R. Rao, S. K. Mattoo, M. V. Murhekar.Direct benefit transfer for nutritional support of patients with TB in India—analysis of national TB program data of 3.7 million patients, 2018–2022. BMC Public Health 24, 299 (2024). [↩]
- RNTCP. Guidance for TB Notification in India. Central TB Division, Directorate General of Health Services (2012). [↩]
- U.B. Singh, K. Rade, R. Rao, N. Kumar, S.K. Mattoo, S. Nair, A. Zumla, S. Sahu. Lessons and updates from India’s National Tuberculosis Elimination Program – bold decisions and innovative ways of fast-tracking progress toward ending tuberculosis. IJID Regions. 14 (2025). [↩] [↩]
- S. Niemann, M. Merker, T. Kohl, P. Supply. Impact of Genetic Diversity on the Biology of Mycobacterium tuberculosis Complex Strains. Microbiology Spectrum.4 (2016). [↩]
- Y. Lan, I. Rancu, M.H. Chitwood, B. Sobkowiak, K. Nyhan, H.H. Lin, C.Y. Wu, B. Mathema, T.S. Brown, C. Colijn, J.L. Warren, T. Cohen. Integrating genomic and spatial analyses to describe tuberculosis transmission: a scoping review. Lancet Microbe. 6, 101094 (2025). [↩]
- M.J. Struelens, C. Ludden, G. Werner, V. Sintchenko, P. Jokelainen, M. Ip; ESCMID Study Group on Public Health Microbiology and ESCMID Study Group on Epidemiological Markers. Real‑time genomic surveillance for enhanced control of infectious diseases and antimicrobial resistance. Frontiers in Science.2, 1298248 (2024). [↩]
- P.J. Gómez-González, S. Campino, J.E. Phelan, T.G. Clark. Portable sequencing of Mycobacterium tuberculosis for clinical and epidemiological applications. Briefings in Bioinformatics. 23 (2022). [↩]