Abstract
Antimicrobial resistance (AMR) is a growing and spreading global health threat affecting the ability of current medicine to treat bacterial infections, increasing the risk of widespread health crises. This study aims to investigate the presence of antimicrobial resistance in the Alameda Creek by performing Kirby-Bauer disk-diffusion tests with the commonly used antibiotics tetracycline, erythromycin, ampicillin, and chloramphenicol. A total of 12 bacterial isolates, four each at three sites from upstream to downstream, were tested for antimicrobial resistance to distinguish its presence and preliminary patterns along the creek. The isolates in our small sample displayed similar resistance trends—all showed susceptibility to tetracycline and no detectable inhibition by erythromycin and ampicillin under our conditions. Only one isolate derived from the downstream site showed no inhibition by chloramphenicol under our conditions, while all other isolates showed susceptibility. This suggests the possible presence of chloramphenicol non-susceptible bacteria downstream, but not widespread resistance. Additional experiments compared bacterial response to antibiotics applied immediately versus 24 hours after bacterial inoculation. Results displayed a sharp decrease in antibiotic effectiveness when applied 24 hours after introduction of bacteria, aligning with the idea that quick treatment of bacteria with antibiotics is important. The findings in this study highlight the value of monitoring environmental waterways like Alameda Creek as potential reservoirs of antibiotic resistance, with implications for ecosystem health, regional public safety, and future public health research initiatives.
Keywords: Antimicrobial resistance, environmental microbiology, ampicillin, tetracycline, erythromycin, chloramphenicol, Alameda Creek, disk-diffusion, stream bacteria, antibiotic susceptibility.
Introduction
Antibiotics are incredibly effective against a majority of bacteria, being able to cause bacterial death through various pathways affecting vital bacterial functions1. However, increased use of antibiotics along with the tendency of bacteria to develop resistance mechanisms under the pressure of selection have resulted in a greater number of bacteria being resistant to previously effective antibiotics, a phenomenon coined antimicrobial resistance (AMR)2. As a result, treatment options for some infections have become limited, and many medical procedures have become riskier3. Bacteria may have intrinsic resistance to antibiotics due to structural barriers or naturally developed defense barriers, while many acquire resistance through mechanisms such as enzymatic inactivation, horizontal transfer of resistance genes, and active efflux pumps4,5.
Though the development of AMR is an inevitable evolutionary outcome with any antibiotic use, the overuse and misuse of antibiotics has allowed AMR to surpass pre-21st century levels6,7. Now, it has become a global health threat, with increased resistance leading to reduced treatment options and therefore higher mortality rates6. In 2019 alone, AMR killed over 1.27 million people and is linked to 5 million deaths globally8. For this reason, AMR needs to be monitored worldwide to determine the scope of the problem and find solutions through increasing antibiotic effectiveness and limiting the development of new resistant strains2.
AMR can be evaluated in a variety of methods, including dilution methods, epsilometer testing (Etest), and Kirby-Bauer disk diffusion testing9,10. While diverse methodologies exist, the Kirby-Bauer disk diffusion test was designated as most standardized and is therefore most widely used9. This method utilizes standardized protocols and the use of antibiotic-impregnated discs on inoculated agar to visually determine bacterial susceptibility to the chosen antibiotics. This disk-diffusion method is also widely adopted for its simplicity and ability to test multiple antibiotics simultaneously10,11.
Antibiotic-resistant bacteria can often be found in environmental streams and natural water sources due to the presence of highly diverse bacteria and potential contamination from wastewater12. Past studies have revealed the presence of vertically and horizontally transferred antibiotic resistance genes (ARGs) and therefore widespread resistance to commonly used antibiotics, such as β-lactams, tetracyclines, and aminoglycosides, in environmental water sources13,14. To further explore antimicrobial resistance in environmental water sources, the Alameda Creek, which stretches from the Diablo Range to the San Francisco Bay, was chosen as a point of study. Past investigations in Alameda County and around the San Francisco Bay have seen an increase in prevalent resistant bacteria, highlighting the need for further testing and monitoring in the region15,16.
It was hypothesized that bacterial density and antimicrobial resistance would increase at downstream sites of Alameda Creek compared to upstream due to higher nutrient concentrations, slower water flow, and increased human activity. This study utilizes a modified version of the disk-diffusion test to investigate and compare the presence and patterns of antimicrobial resistance to tetracycline, ampicillin, erythromycin, and chloramphenicol in the California Alameda Creek. Past studies have shown that tetracycline and erythromycin are common in environmental compartments, which leads to a greater likelihood of resistance development17,18. Chloramphenicol and ampicillin have also been detected in aquatic environments, indicating that the presence of resistance to these antibiotics is probable19,20. In addition to general AMR testing, two methodologies were compared: the standard procedure, which introduces antibiotic discs immediately after inoculation, and a modified procedure, which introduces antibiotic discs 24 hours after inoculation of bacteria. This was done to evaluate how bacterial growth phases impact observed zones of inhibition.
Materials and Methods
Water Sampling from Three Sites
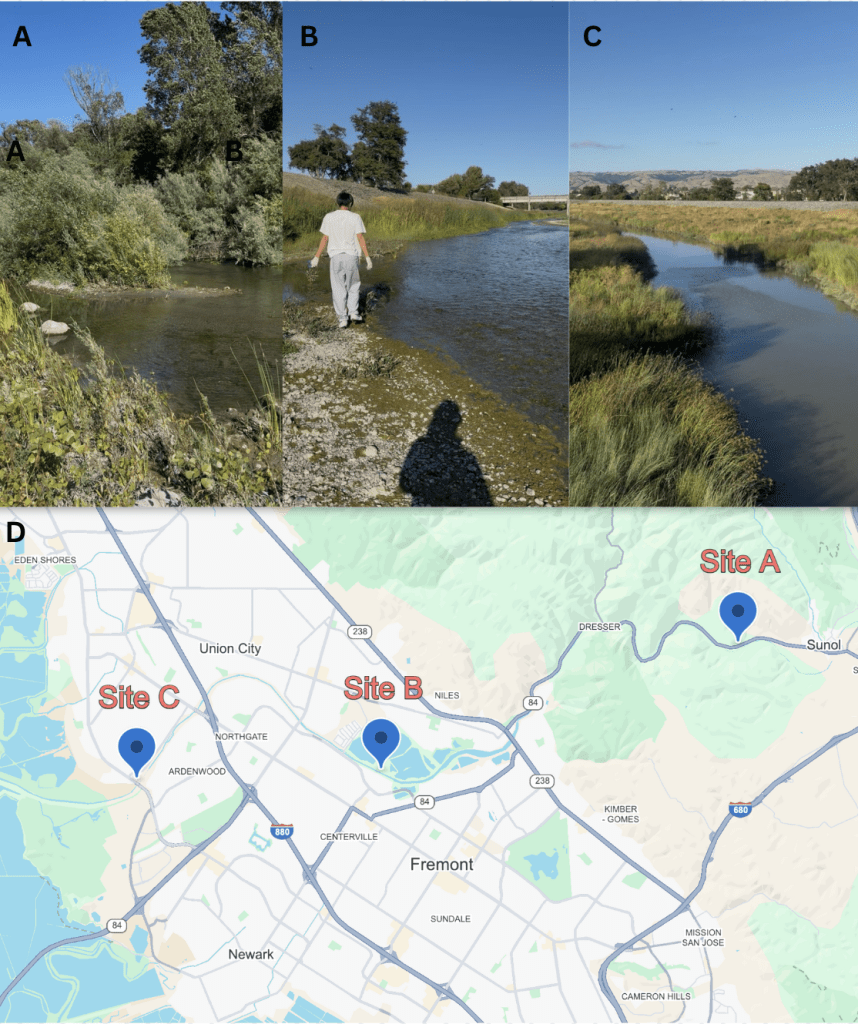
One 50 mL water sample was collected from each of three sites on the Alameda Creek on a single day, July 4th, 2025, and stored in sterile conical tubes. Site A (37.5937539, -121.9103416) was located upstream in Sunol, CA, alongside California State Route 84; Site B (37.5675508, -122.0040036) was located midstream in Fremont, CA near the Quarry Lakes Regional Recreation Area; Site C (37.5656207, -122.0678975) was located downstream in Fremont, CA near Coyote Hills Regional Park (Figure 1). Water samples were transported in a cooler with ice before being stored at 4℃.
Preparation of a Low-Contamination Work Area
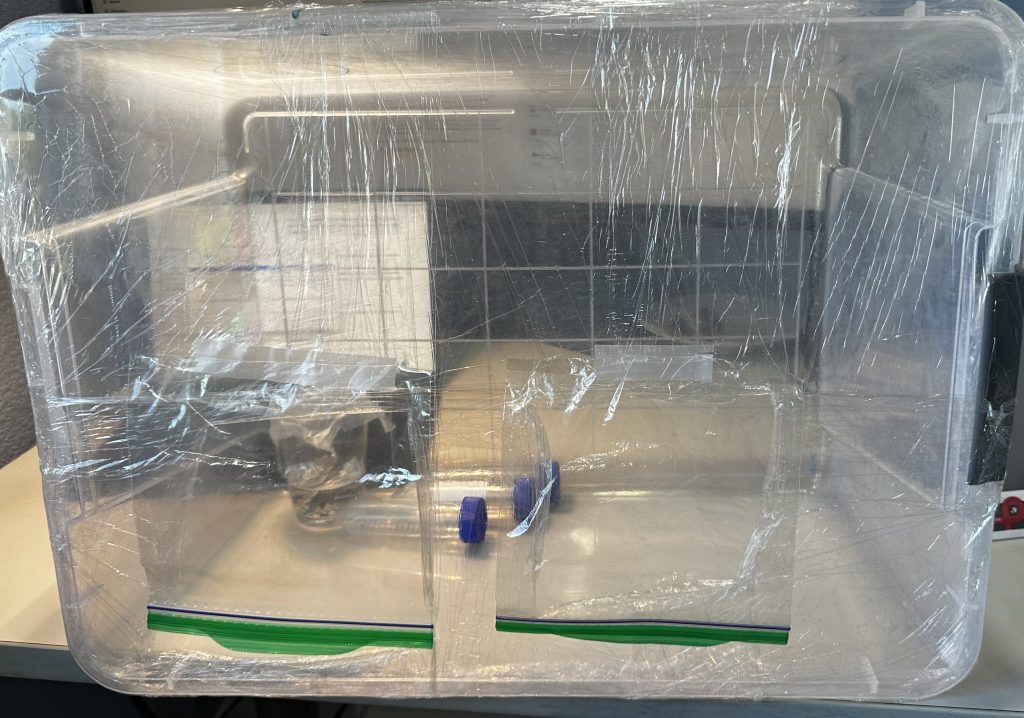
Due to limited access to a standard biosafety cabinet, a still air box was created to maintain a low-contamination workspace for all wet experiments (Figure 2). A transparent plastic box was positioned on its side, and the open face was sealed with multiple layers of plastic wrap to limit air exchange. Hand holes were cut in the plastic wrap using scissors, and plastic bags were taped above the holes to allow handling of materials while blocking out airborne contaminants.
This setup functions similarly to a biosafety cabinet by creating an enclosed, semi-sterile workplace that minimizes exposure to airborne microorganisms. True biosafety cabinets use HEPA-filtered laminar airflow to actively remove contaminants, while our still air box reduces air exchange instead of filtering the air. While the still air box provides sufficient contamination control for the basic microbiological work done in this study, it does not achieve the sterile conditions of a standard biosafety cabinet.
Before and after each experimental session, the interior surfaces of the still air box, as well as gloves and materials to be introduced, were thoroughly sprayed with 70% isopropyl alcohol to disinfect surfaces and reduce microbial load. These precautions were expected to adequately control contamination for the environmental sample processing and antimicrobial resistance testing work done.
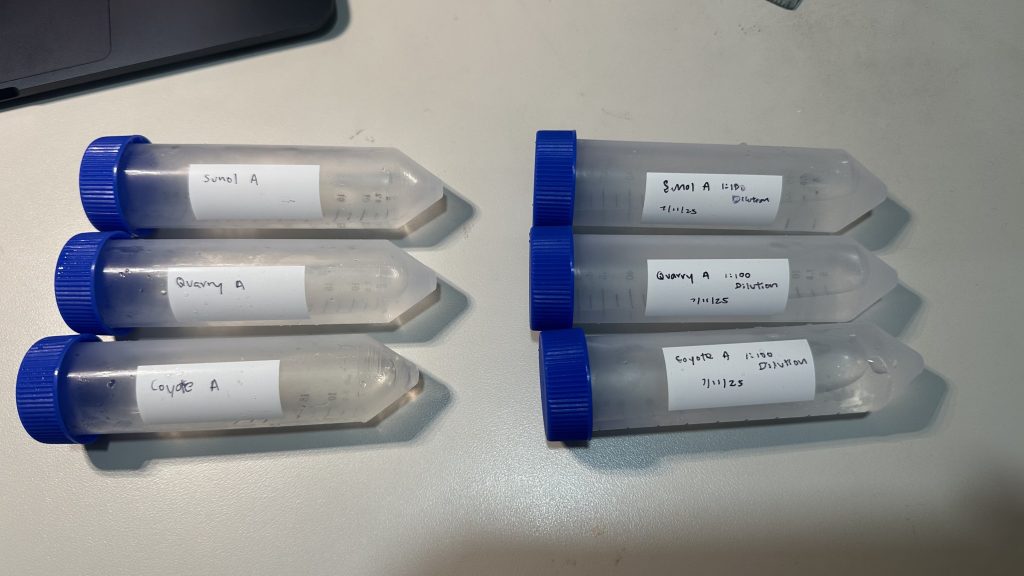
Dilutions
Dilutions of original water samples were conducted to reduce microbial concentration prior to inoculation and incubation of bacteria (Figure 3). A 1:100 dilution of each water sample was created by transferring 0.5 mL of each original water sample to a sterile conical tube with 49.5 mL distilled water.
Inoculation and Incubation of Creek Bacteria
Nutrient agar was chosen as a culture medium for its ability to support a wide variety of bacteria. Nutrient agar plates were sourced from Evviva Sciences (Fremont, CA, USA).
Inside the disinfected still air box, 1 mL of the 1:100 diluted water sample was pipetted onto the surface of an agar plate and gently spread by tilting the plate to distribute the liquid evenly. Streaking for colony isolation was performed using a sterile cotton swab: the swab was drawn across the agar surface in a zig-zag pattern, the plate was rotated 90 degrees, and the streaking was repeated. The sequence was continued until six streaking patterns were completed per plate. The procedure was repeated for three technical plates for each water sample, resulting in nine plates total. All agar plates were sealed with parafilm to minimize moisture loss.
The various bacteria were incubated in an inverted position at room temperature (20°C-24°C) for 38 hours to allow visible colony growth. After inoculation, colony forming units (CFUs) were counted manually. To avoid double counting, each CFU was marked on the plate’s underside with a permanent marker as it was recorded. Confluent or merged areas of growth were recorded as a single CFU. Plates with excessive/overgrown lawns or overlapping colonies were excluded from counts.
Isolation of Bacteria
4 bacteria with visibly different colors were selected from each site’s diluted water plates to capture apparent morphological diversity (total 12 bacteria selected). Due to limited access to a sterilized loop, the wood side of a sterile cotton swab was used to pick up and transfer bacteria from one colony on the diluted creek water plate to a new agar plate. The cotton end of the swab was then submerged in distilled water and was used to spread the transferred bacteria around on the new plate in a zig-zag motion repeated 6 times total, as previously described (Figure 4). Swab use was kept as consistent as possible. This process was repeated 3 times for each of the 4 chosen bacteria at each site (12 plates per site, 36 plates total for all 3 sites). All agar plates were sealed with standard parafilm.

Isolated bacteria samples were allowed to incubate in an inverted position for 24 hours at room temp (20℃-24℃) before antibiotic discs were introduced to each agar plate.
Antibiotic Resistance Testing
To determine comparative antibiotic susceptibility of isolated creek bacteria to various antibiotics, the standard Kirby-Bauer testing procedure was utilized21.
Tetracycline (30 μg), erythromycin (15 μg), ampicillin (10 μg), and chloramphenicol (30 μg) antibiotic discs were sourced from Carolina Biological (Burlington, NC, USA). All antibiotic discs were stored at 4℃ in separate sealed plastic bags with desiccant. A pair of tweezers was disinfected in 70% isopropyl alcohol before and after transferring each of 4 individual antibiotic discs onto the agar plate. The 4 antibiotic discs were distributed evenly across the plate. Each of the 12 isolates was tested against these four antibiotics with 3 technical replicates.
Due to safety restrictions, all plates were incubated at room temperature (20℃-24℃) for 24 hours instead of the standard 35-37℃ outlined in the Kirby-Bauer testing protocol10. For this reason, CLSI/EUCAST breakpoints cannot be applied to our work. Though differing from standard procedure, consistency for all samples allows for a valid comparison of relative susceptibility between distinct bacterial isolates and individual antibiotics under our experimental conditions.
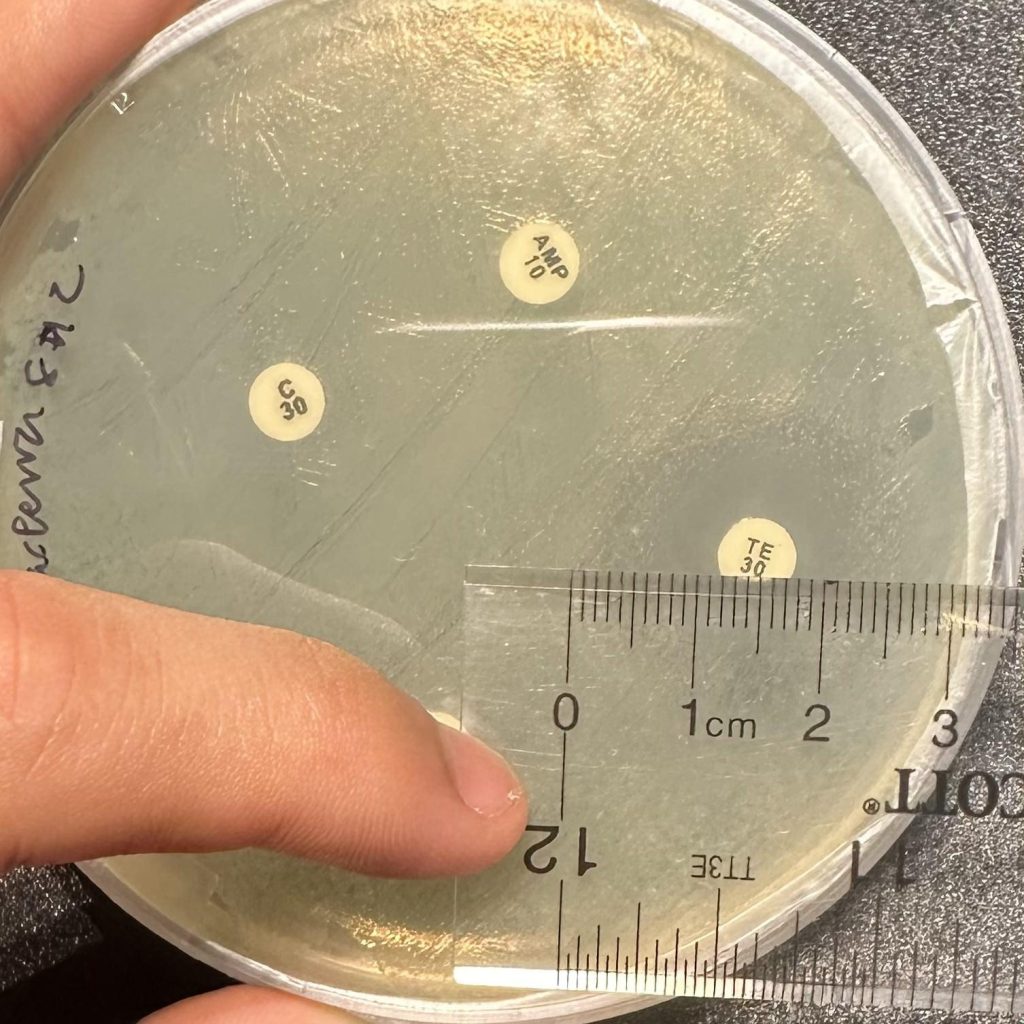
Zones of inhibition were measured in millimeters using a ruler 24 hours after the introduction of antibiotic discs. A measurement of 6 mm indicates no zone of inhibition (the diameter of the antibiotic disc is 6 mm), while measurements above 6 mm indicate some level of susceptibility (Figure 5).
An inhibition-abundance score was developed as an exploratory tool to combine inhibition and abundance metrics, matching our experiment design. We defined this score as the average zone of inhibition resulting from an antibiotic for a given isolate, divided by the recorded CFU/mL at the isolate’s origin site, and then multiplied by 103. Here, a higher number signifies greater susceptibility, while a lower number signifies lower inhibition.
IAS=Z.103/C
Where IAS = Inhibition-Abundance Score; Z = ZOI; C = CFU/mL.
Statistical Testing
A two-way ANOVA was conducted to measure the effect of site (Factor A: 3 levels—Sites A, B, C) and antibiotic (Factor B: 4 levels—tetracycline, erythromycin, ampicillin, chloramphenicol) on zones of inhibition. The statistical analysis incorporated a total of 144 observations: 4 isolates per site × 3 sites × 4 antibiotics × 3 technical replicates. Main effects and site x antibiotic interaction were tested at α = 0.05 significance level.
Methodological Testing
A modified methodology was also tested in comparison to the standard Kirby-Bauer method, which calls for the introduction of antibiotic discs immediately after inoculation10. In this modified methodology, antibiotic discs were only introduced to the bacterial sample 24 hours after inoculation. All isolates used in the standard methodology were tested against all antibiotics using the modified methodology, for a total of 12 isolates x 4 antibiotics x 3 technical replicates. This allows for comparison of antibiotic effectiveness on growing versus established bacteria, highlighting the impact of growth phase on inhibition zones.
Results
Bacterial Abundance Across Sites
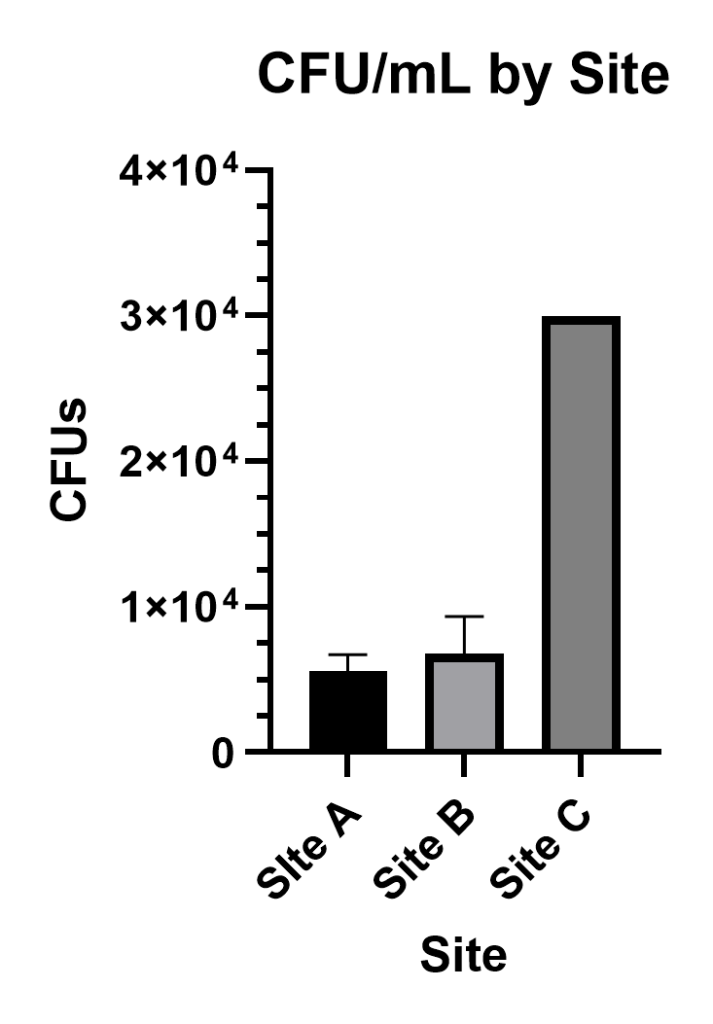
To quantify bacterial abundance across all three sites, plates derived from 1:100 diluted creek water were counted for colony forming units (CFUs). Values are reported as mean ± SD. Averaging across the three technical replicates made for each site’s sample, Site A displayed 56 ± 12 CFU/mL (n=3), Site B displayed 68 ± 26 CFU/mL (n=3), and Site C’s plates were too numerous to count (TNTC) and therefore reported as >300 CFU/mL (n=3) per plate. Taking the 10-2 dilution factor into account, bacterial density by site can be reported as (5.6 ± 1.2) ⨉ 103 CFU/mL (n=3) for Site A, (6.8 ± 2.6) ⨉ 103 CFU/mL (n=3) for site B, and TNTC, reported as >3.0 ⨉ 104 CFU/mL (n=3), for Site C (Figure 6).
Antibiotic Susceptibility of Isolates
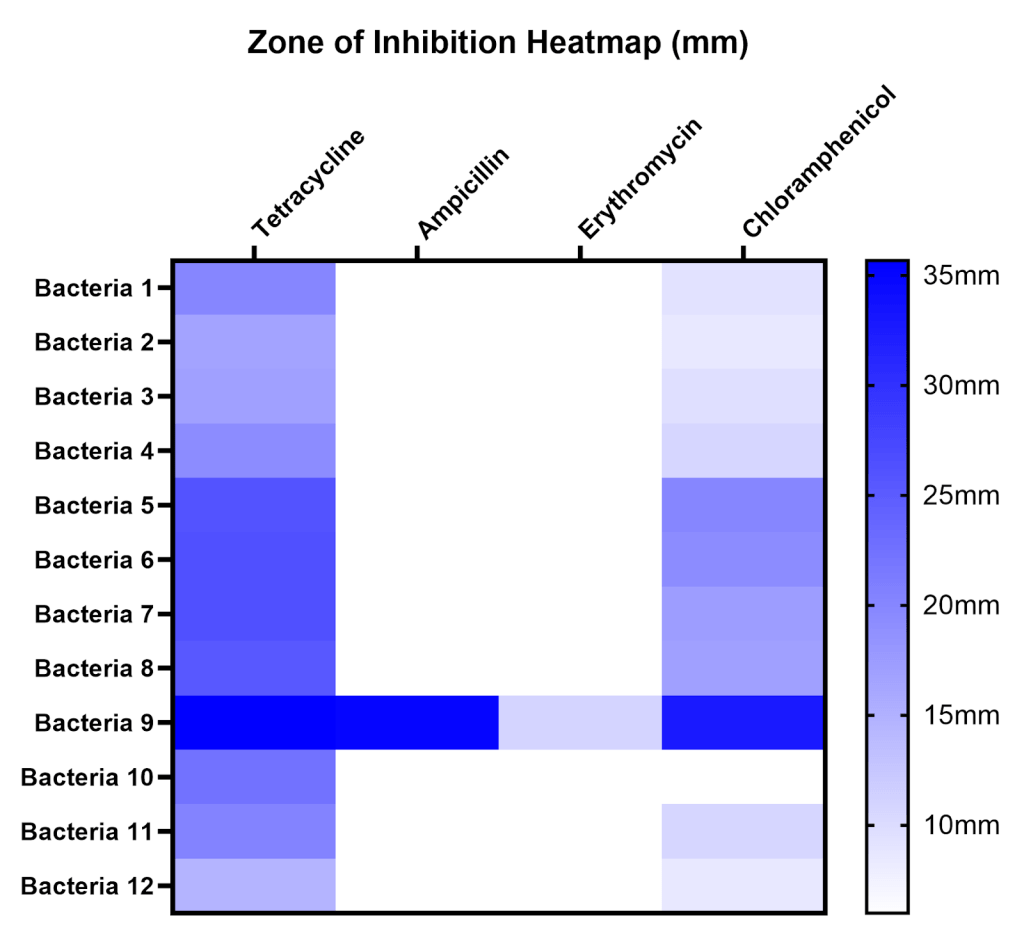
To compare susceptibility to different antibiotics and across different sites, zones of inhibition (ZOI) were measured for all antibiotics on all isolates, the mean and standard deviation of ZOI are displayed in a table, and average ZOI was summarized on a heatmap (Table 1, Figure 7). A ZOI of 6 mm (white) indicates no inhibition under our conditions. Bacteria 1-4 were isolated from Site A, bacteria 5-8 from Site B, and bacteria 9-12 from Site C.
| Tetracycline | Ampicillin | Erythromycin | Chloramphenicol | |
| Bacterium 1 | 20 ± 2 | 6 ± 0 | 6 ± 0 | 9.33 ± 0.58 |
| Bacterium 2 | 19.33 ± 4.16 | 6 ± 0 | 6 ± 0 | 10.67 ± 0.58 |
| Bacterium 3 | 16.67 ± 1.15 | 6 ± 0 | 6 ± 0 | 8.67 ± 1.15 |
| Bacterium 4 | 17 ± 2 | 6 ± 0 | 6 ± 0 | 9.67 ± 0.58 |
| Bacterium 5 | 26 ± 2.65 | 6 ± 0 | 6 ± 0 | 20 ± 1.73 |
| Bacterium 6 | 26.33 ± 0.58 | 6 ± 0 | 6 ± 0 | 19.33 ± 1.15 |
| Bacterium 7 | 26.33 ± 2.31 | 6 ± 0 | 6 ± 0 | 17.33 ± 1.53 |
| Bacterium 8 | 25.33 ± 0.58 | 6 ± 0 | 6 ± 0 | 17 ± 0 |
| Bacterium 9 | 35.67 ± 2.08 | 35 ± 1 | 11 ± 1 | 32.67 ± 3.06 |
| Bacterium 10 | 22.33 ± 3.06 | 6 ± 0 | 6 ± 0 | 6 ± 0 |
| Bacterium 11 | 20.33 ± 0.58 | 6 ± 0 | 6 ± 0 | 10.67 ± 1.15 |
| Bacterium 12 | 14.67 ± 3.51 | 6 ± 0 | 6 ± 0 | 8.67 ± 1.15 |
A two-way ANOVA was conducted to evaluate the significance of site and antibiotic on the resulting zones of inhibition (Table 2). Antibiotic type had a significant effect on ZOI (F3,132=58.19, p<0.0001), and the site was also found to have a significant effect on ZOI (F2,132=9.60, p=0.0001). The site x antibiotic interaction was also found to be significant (F6,132=3.40, p=0.0037). These results suggest that under our experimental conditions, both the choice of antibiotics and sites, as well as their combined effect, influenced the observed patterns of inhibition.
| Source | DF | SS | MS | F | P-value |
| Factor A | 2 | 617.5417 | 308.7708 | 9.6040 | 0.0001 |
| Factor B | 3 | 5612.2500 | 1870.7500 | 58.1877 | <0.0001 |
| Interaction (AxB) | 6 | 656.1250 | 109.3542 | 3.4013 | 0.0037 |
| Error | 132 | 4243.8333 | 32.1503 | – | – |
| Total | 143 | 11129.7500 | – | – | – |
Methodological Comparison Results
Plates that were immediately introduced to antibiotics (standard methodology) after inoculation displayed visual zones of inhibition 24 hours after antibiotic disc introduction, while those that were only introduced to antibiotics 24 hours after inoculation (modified methodology) mostly showed no zones of inhibition 24 hours after disc introduction (Figure 8, 9). For example, when testing all 12 isolates against tetracycline with 3 technical replicates per isolate, plates using the standard methodology had an average ZOI of 22.5 ± 5.95 mm (n=36), while plates using the modified methodology had an average ZOI of 2.14 ± 3.53 mm (n=36). Similar patterns were observed for chloramphenicol, while ampicillin and erythromycin mostly showed low levels of inhibition under both conditions (Figure 9). Values are reported as mean ± SD.
In some plates using modified methodology, damp regions were initially recorded as potential zones of inhibition. However, these were later determined to be artifacts caused by disc placement on an established lawn rather than true inhibition.
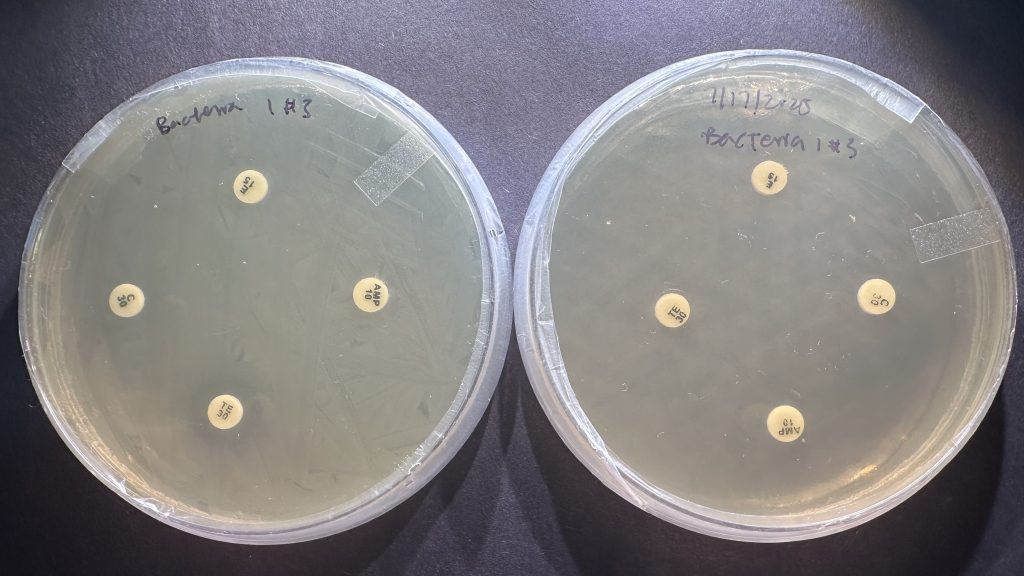
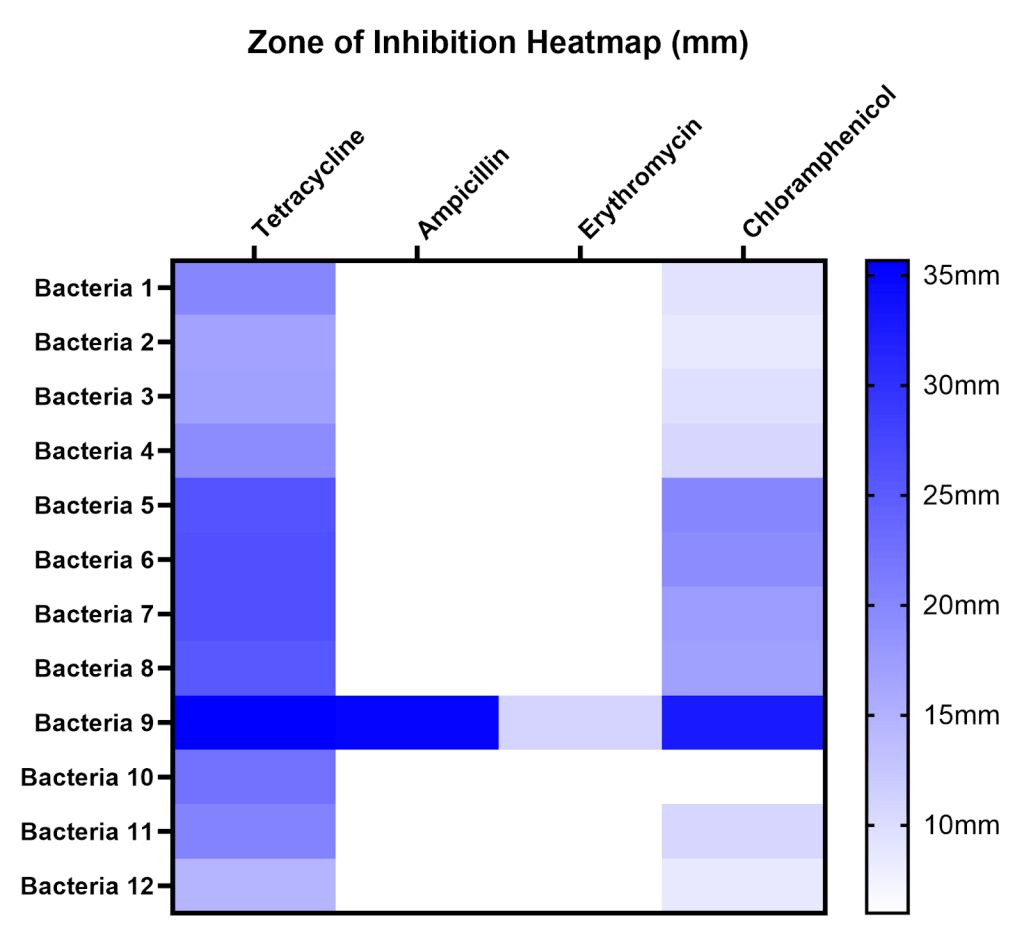
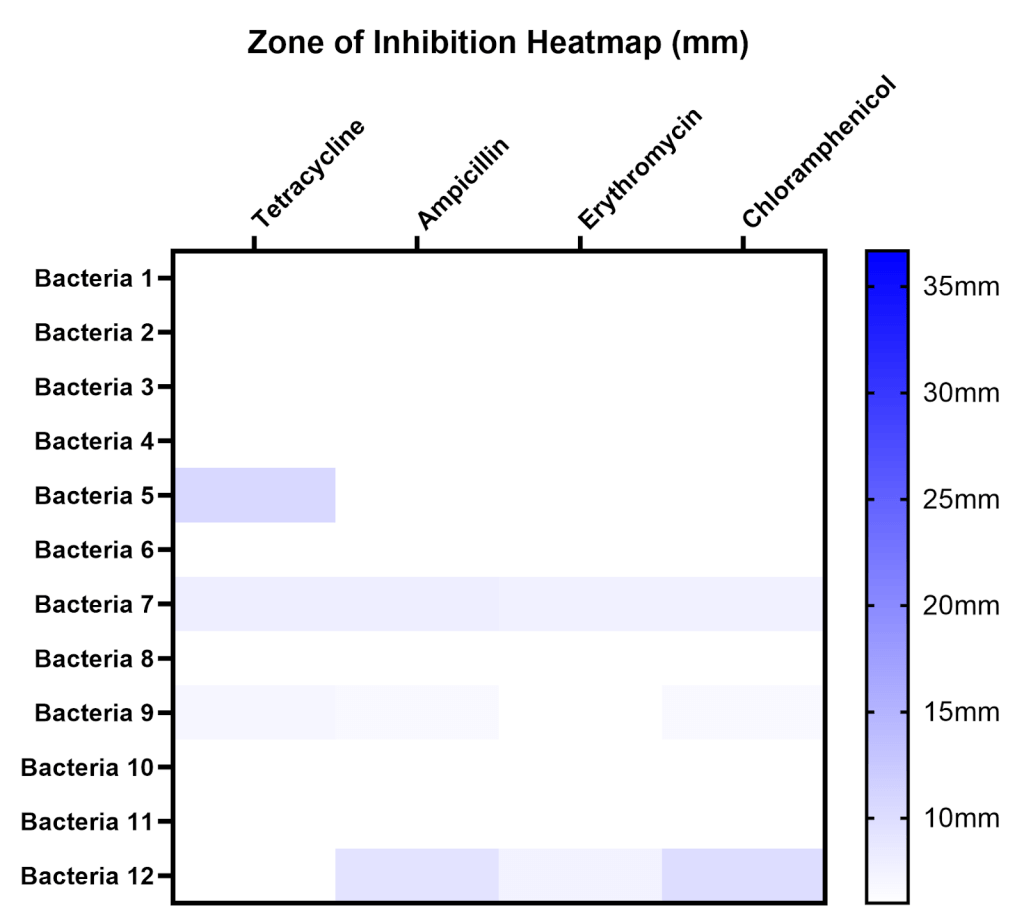
Figure 9 | ZOIs resulting from antibiotics are much more apparent using the standard methodology (left) in comparison to the modified methodology (right). Most ZOIs resulting from the modified methodology show no detectable inhibition (6 mm).
This data can be put into the context of our Inhibition-Abundance Score. The IAS of each antibiotic at each site was calculated using an average zone of inhibition for each antibiotic, and the IASs of all trials of all antibiotics were averaged for each bacterium. IASs for each bacterium were averaged to determine an average IAS per site. Site A and B have similar IASs (Site A = 17.93, Site B = 19.79), while Site C’s IAS (4.85) is significantly lower than previous sites (Table 3).
| Average ZOI for all isolates at given site against all antibiotics | CFU/mL | Inhibition-Abundance Score | |
| Site A | 9.98 | 55.67 | 17.93 |
| Site B | 13.48 | 68 | 19.82 |
| Site C | 14.56 | 300 (TNTC) | 4.85 |
Discussion
Bacterial Density Patterns and Environmental Factors
The measurement of CFU/mL for each site showed a slight increase in bacterial density from Site A to Site B and a notable jump from Site B to Site C. When applied to the IAS, CFU increase causes it to decrease dramatically from Site B to Site C. One possible explanation for this is an increase in the availability of nutrients vital for bacterial life, such as nitrogen and phosphorus, potentially derived from municipal and residential wastewater or urban runoff from the nearby Brookvale, Cabrillo, Ardenwood, Northgate, Union City, and Lakes and Birds communities. Supporting these hypotheses would require tests to measure specific nutrients and wastewater indicators.
Visually, the creek at Site C was also wider with deeper and more stagnant water. Another possible explanation is that slower water may be exposed to more direct sunlight, which could cause an increase in the creek’s temperature and the bacteria’s metabolic and reproduction rate22 Conversely, the cooler water and faster flow rate found in the upstream sites, A and B, may flush bacteria constantly and slow metabolic rate, potentially preventing large populations from forming. These hypotheses about water flow, temperature, and sunlight exposure would require direct measurement to confirm.
Antibiotic Susceptibility and Resistance Patterns
The two-way ANOVA results support the pattern that both the antibiotic and site influenced observed zones of inhibition. The significant interaction term (p=0.0037) suggests that the effect of antibiotic type on inhibition varied depending on the sampling site. However, given our small sample of 12 isolates and non-standard testing conditions, these statistical findings should be interpreted as preliminary patterns in this limited sample rather than antimicrobial resistance patterns along the Alameda Creek system.
In our small sample, all isolates showed susceptibility to tetracycline, and almost all isolates, except Bacterium 10 from Site C, showed susceptibility to chloramphenicol under our conditions. These results suggest the absence of tetracycline resistance in our sample. The possible presence of chloramphenicol non-susceptibility at Site C may reflect either intrinsic resistance of a naturally resistant species or acquired resistance via mobile genetic elements, which would have greater public health implications. Without bacterial identification via Gram staining, 16S rRNA sequencing, or PCR, we cannot distinguish between the two possibilities. If Bacterium 10 is an ordinarily susceptible species, chloramphenicol non-susceptibility would be noteworthy and potentially indicates selective pressure from environmental contamination. However, if it were to belong to an intrinsically resistant species such as Pseudomonas aeruginosa, this observation may reflect natural resistance patterns23.
Ampicillin and erythromycin showed no detectable inhibition against almost all of our isolates under our conditions, excluding bacterium 9 from Site C. Without Gram staining or molecular identification (16S rRNA/PCR), we cannot distinguish whether non-susceptibility represents intrinsic or acquired resistance. Many Gram-negative bacteria intrinsically resist ampicillin due to beta-lactamase production and erythromycin due to their impermeable outer membranes, and bacteria found in environmental freshwater are predominantly Gram-negative24,25,26. Therefore, the observed non-susceptibility is consistent with expected intrinsic resistance of Gram-negative environmental bacteria.
Tetracycline, erythromycin, and ampicillin are all commonly used clinically to treat both animals and humans—lack of inhibition by these antibiotics suggests pollution from agricultural runoff, human sewage, or hospital waste27,28,29. Chloramphenicol was previously used widely, but its resistance persists in some bacterial populations, which could explain the chloramphenicol non-susceptible isolate observed at Site C30.
Exploring site differences using the IAS (Inhibition-Abundance Score), Site C had a much lower IAS than that of Sites A and B. This decrease reflects the substantially greater bacterial population found at Site C, which reduced the score regardless of actual resistance level of bacteria. Therefore, the lower IAS at Site C should not be interpreted as indicating stronger resistance, but rather as a reflection of the combined effect of higher bacterial density and susceptibility patterns. Here, the IAS focuses on the consideration of both bacterial density and susceptibility patterns in providing a more complete picture of environmental antimicrobial patterns at each site.
Methodological Comparison and Implications
Comparing zones of inhibition observed following the Kirby-Bauer vs. modified methodology, zones were much greater (>2-3x) following the standard timing. Our results therefore suggest that antibiotics are more effective before dense bacterial growth is established. Such a trend could have importance in public health—our simplified agar experiment does not mimic the complexity of infections in the human body, but it is consistent with the general principle that early antibiotic treatment of bacteria is beneficial.
Limitations and Future Directions
Though these disk-diffusion tests identified the effectiveness of certain antibiotics on the sampled bacteria population, there are important limitations to consider:
- Lack of bacterial identification. With the current resources available, we were unable to determine bacterial species through 16S rRNA, PCR, or classify using Gram-staining or other classification methods. This makes it difficult to understand the bacterial population and does not allow us to differentiate natural vs acquired resistances (many environmental Gram-negative bacteria would be intrinsically resistant to ampicillin)31.
- Deviation from standard methodology. This study was limited to using sterile wooden swabs for bacterial isolation. Therefore, inoculum size may vary between isolates. A room temperature of 20-24℃ was used for incubation, deviating from the standard Kirby-Bauer procedure10. As a result, zones of inhibition cannot be compared to CLSI/EUCAST breakpoints, and susceptibility can only be compared with other data in this experiment.
- Selection bias. 4 bacteria from each site were chosen to be isolated based on apparent color differences, which could introduce bias by oversampling certain morphotypes that are more visually distinct while missing rarer or less pigmented species.
- Lack of standard quality control reference strains. No reference strains, such as E. coli ATCC 25922, were tested to verify that antibiotic discs and the testing protocol yielded results within expected ranges. This makes it difficult to confirm that the discs and procedure performed as intended.
- Low sample size. This study collected water samples on a single day (July 4th, 2025) and isolated only 4 bacterial colonies per site on the Alameda Creek, which may allow for non-susceptible bacteria to go undetected and limits statistical power in detecting true trends in antimicrobial susceptibility patterns. A single non-susceptible isolate at one site, such as the chloramphenicol non-susceptible bacterium at Site C, could be due to chance selection as opposed to a site-level pattern.
- Low IAS scope. The Inhibition-Abundance Score developed in this study is matched to the parameters of this investigation (zones of inhibition and bacterial abundance), but it is not meant to be a standard resistance metric and may not be fit for use in other studies. However, the use of some index variant can be helpful in summarizing resistance profiles at a given site.
- Imprecise CFU counts at high-density sites. Site C plates at 1:100 dilution were too numerous to count (TNTC, >300 CFU/plate), not allowing for precise quantification of bacterial density and therefore potentially inaccurate density difference comparisons to upstream sites.
- Uncontrolled plate differences in methodological comparison. Plates were prepared for both methodologies using the same bacterial isolate, but plate-to-plate differences in lawn density and moisture were not controlled. While the large and consistent differences observed suggest timing is the primary factor, we cannot rule out that conditional differences may have contributed to ZOI differences.
Future work for this study could include more independent water samples per site and ≥20 isolates per site to capture spatial variation and strengthen statistical confidence. At high-density sites like Site C, higher dilutions (10-3 to 10-4) should be used to ensure CFU counts fall within 30-300 CFUs/plate, allowing for more accurate quantification and comparison to other sites. Multiple sampling dates in different seasons could be tested to analyze temporal variation. Numbering initial bacterial colonies and using a random number generator or sampling from different streak sectors could capture bacterial diversity more accurately.
Future studies could test antibiotic discs against a standard quality control reference strain, such as E. coli ATCC 25922, and compare results to published ranges to ensure that discs and methods are functioning appropriately. Gram-staining of bacteria and bacterial identification should be performed in the future to firmly classify observed resistances as intrinsic vs. acquired resistance—this is particularly important for Bacterium 10 from Site C, the only chloramphenicol non-susceptible isolate. Future work should also maintain standard Kirby-Bauer conditions, including Mueller-Hinton agar and 35-37℃ incubation, to allow for comparison with clinical breakpoints and standard resistance classification. Other commonly used antibiotics, such as vancomycin, gentamicin, or ciprofloxacin, could also be tested to scope out antibiotic resistance. Nutrient testing of Alameda Creek waters could also provide insight into evolutionary pressure in the specific area.
Future, more detailed studies could also investigate the ecological and public health implications of antibiotic resistance in stream bacteria—by assessing connections between resistance patterns, human activity, and water quality, we could gain a clearer idea of community risk in the local ecosystem.
Conclusion
This study observed susceptibility to tetracycline and lack of inhibition by chloramphenicol, ampicillin, and erythromycin in the Alameda Creek through sampling of bacterial isolates at three points along the creek. Under our conditions, all sampled isolates showed no detectable inhibition by erythromycin and ampicillin, while one isolate at Site C showed no inhibition by chloramphenicol. Our preliminary data is consistent with the idea that while tetracycline and chloramphenicol non-susceptibility are absent or uncommon in our sample, intrinsic or acquired non-susceptibility to other antibiotics in the suite is more common. Our current data is unable to directly separate these results between intrinsic and acquired resistance due to lack of Gram staining and molecular identification tools. Under our conditions, bacterial density appeared to increase further downstream, while levels of inhibition did not show a similar increase.
An Inhibition-Abundance Score (IAS) was created as an exploratory index that provides a basic profile of resistance in a system, such as Alameda Creek—the IAS accounts for both the degree of susceptibility and bacterial abundance. The inclusion of bacterial density allows for a more complete understanding of a microbiological system.
The methodological comparison in this showed differing results when introducing antibiotic discs 24 hours versus immediately after inoculation. Plates with antibiotics introduced immediately showed larger zones compared to plates with antibiotics introduced 24 hours later, suggesting that faster antibiotic introduction correlates with greater effectiveness in treatment.
Building on previous studies in the area, this study provides preliminary evidence for the presence of bacteria with reduced susceptibility to certain antibiotics in the Alameda Creek system and underscores the need for ongoing investigations. Using standard conditions, quality control reference strains, Gram-staining, commonly applied antibiotics, and a larger sample size, among other potential additions, future studies can be conducted to gain a better understanding of the bacterial population and definitive antimicrobial resistance classifications. Nutrient testing of creek waters and surveys of surrounding land can be beneficial in finding key sources of pollution and antimicrobial resistance.
References
- G. Kapoor, S. Saigal, A. Elongavan, Action and resistance mechanisms of antibiotics: A guide for clinicians, Journal of Anaesthesiology Clinical Pharmacology, Vol. 33, pg. 300-305, 2017, https://doi.org/10.4103/joacp.JOACP_349_15. [↩]
- M. A. Salam, M. Y. Al-Amin, M. T. Salam, J. S. Pawar, N. Akhter, A. A. Rabaan, M. A. A. Alqumber, Antimicrobial resistance: A growing serious threat for global public health, Healthcare. Vol 11, 1946 (2023), https://doi.org/10.3390/healthcare11131946. [↩] [↩]
- World Health Organization. Antimicrobial resistance. 2023, https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance. [↩]
- Z. Li, J. Tang, X. Wang, X. Ma, H. Yuan, C. Gao, Q. Guo, X. Guo, J. Wan, C. Dagot. The environmental lifecycle of antibiotics and resistance genes: transmission mechanisms, challenges, and control strategies. Microorganisms. Vol. 13, pg. 2113, 2025, https://doi.org/10.3390/microorganisms13092113. [↩]
- M. E. Elshobary, N. K. Badawy, Y. Ashraf, A. A. Zatioun, H. H. Masriya, M. M. Ammar, N. A. Mohamed, S. Mourad, A. M. Assy. Combating antibiotic resistance: mechanisms, multidrug-resistant pathogens, and novel therapeutic approaches: an updated review. Pharmaceuticals. Vol. 18, pg. 402, 2025, https://doi.org/10.3390/ph18030402. [↩]
- M. A. Salam, M. Y. Al-Amin, M. T. Salam, J. S. Pawar, N. Akhter, A. A. Rabaan, M. A. A. Alqumber, Antimicrobial resistance: A growing serious threat for global public health, Healthcare. Vol 11, 1946 (2023), https://doi.org/10.3390/healthcare11131946. [↩] [↩]
- J. Davies, D. Davies. Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev. Vol. 74, pg. 417-433, 2010, https://doi.org/10.1128/MMBR.00016-10. [↩]
- Centers for Disease Control and Prevention, Antimicrobial resistance facts and stats, Centers for Disease Control and Prevention (2025), https://www.cdc.gov/antimicrobial-resistance/data-research/facts-stats/index.html. [↩]
- Z. A. Khan, M. F. Siddiqui, S. Park. Current and emerging methods of antibiotic susceptibility testing. Diagnostics (Basel). Vol. 9, pg. 49, 2019, https://doi.org/10.3390/diagnostics9020049. [↩] [↩]
- J. Hudzicki, Kirby-Bauer disk diffusion susceptibility test protocol, American Society for Microbiology (2016), https://asm.org/protocols/kirby-bauer-disk-diffusion-susceptibility-test-pro. [↩] [↩] [↩] [↩] [↩]
- I. Gajic, J. Kabic, D. Kekic, M. Jovicevic, M. Milenkovic, D. Mitic Culafic, A. Trudic, L. Ranin, N. Opavski, Antimicrobial susceptibility testing: A comprehensive review of currently used methods, Antibiotics, Vol. 11, pg. 427, 2022, https://doi.org/10.3390/antibiotics11040427. [↩]
- D. G. J. Larsson, C. F. Flach, Antibiotic resistance in the environment, Nature Reviews Microbiology, Vol. 20, pg. 257-269, 2022, https://doi.org/10.1038/s41579-021-00649-x. [↩]
- X. X. Zhang, T. Zhang, H. H. P. Fang. Antibiotic resistance genes in water environment. Appl Microbiol Biotechnol. Vol. 82, pg. 397-414, 2009, https://doi.org/10.1007/s00253-008-1829-z. [↩]
- P. T. Biyela, J. Lin, C. C. Bezuidenhout. The role of aquatic ecosystems as reservoirs of antibiotic resistant bacteria and antibiotic resistance genes. Water Sci Technol. Vol. 50, pg. 45-50, 2004, https://doi.org/10.2166/wst.2004.0014. [↩]
- R. Marusinec, M. Shemsu, T. Lloyd, B. M. Kober, D. T. Heaton, J. A. Herrera, M. Gregory, V. Varghese, J. Nadle, K. K. Trivedi, Epidemiology of carbapenem-resistant organisms in Alameda County, California, 2019-2021, Antimicrobial Stewardship & Healthcare Epidemiology, Vol. 4, e64, 2024, https://doi.org/10.1017/ash.2024.33. [↩]
- C. F. Moritz, R. E. Snyder, L. W. Riley, D. W. Immke, B. K. Greenfield. Antimicrobial drug-resistant gram-negative saprophytic bacteria isolated from ambient, near-shore sediments of an urbanized estuary: Absence of β-lactamase drug-resistance genes. Antibiotics. Vol. 9, pg. 400, 2020, https://doi.org/10.3390/antibiotics9070400. [↩]
- R. Daghrir, P. Drogui. Tetracycline antibiotics in the environment: a review. Environ Chem Lett. Vol. 11, pg. 209-227, 2013, https://doi.org/10.1007/s10311-013-0404-8. [↩]
- B. H. Schafhauser, L. A. Kristofco, C. M. Ribas de Oliveira, B. W. Brooks. Global review and analysis of erythromycin in the environment: Occurrence, bioaccumulation and antibiotic resistance hazards. Environmental Pollution. Vol. 238, pg. 440-451, 2018, https://doi.org/10.1016/j.envpol.2018.03.052. [↩]
- L. M. Nguyen, N. T. T. Nguyen, T. T. T. Nguyen, et al. Occurrence, toxicity and adsorptive removal of the chloramphenicol antibiotic in water: a review. Environ Chem Lett. Vol. 20, pg. 1929-1963, 2022, https://doi.org/10.1007/s10311-022-01416-x. [↩]
- R. Hirsch, T. Ternes, K. Haberer, K.-L. Kratz. Occurrence of antibiotics in the aquatic environment. Science of The Total Environment. Vol. 225, Issues 1-2, pg. 109-118, 1999, https://doi.org/10.1016/S0048-9697(98)00337-4. [↩]
- J. Hudzicki, Kirby-Bauer disk diffusion susceptibility test protocol, American Society for Microbiology (2016), https://asm.org/protocols/kirby-bauer-disk-diffusion-susceptibility-test-pro. [↩]
- Y. Qiu, Y. Zhou, Y. Chang, X. Liang, H. Zhang, X. Lin, K. Qing, X. Zhou, Z. Luo, The effects of ventilation, humidity, and temperature on bacterial growth and bacterial genera distribution, International Journal of Environmental Research and Public Health, Vol. 19, pg. 15345, 2022,
https://doi.org/10.3390/ijerph192215345. [↩]
- M. Kono, K. O’Hara. Mechanism of chloramphenicol‑resistance mediated by kR102 factor in Pseudomonas aeruginosa. J Antibiot (Tokyo). Vol. 29, pg. 176-180, 1976, https://doi.org/10.7164/antibiotics.29.176 [↩]
- K. Bush, P. A. Bradford, Epidemiology of β-lactamase-producing pathogens, Clinical Microbiology Reviews, Vol. 33, e00047-19, 2020, https://doi/org/10.1128/CMR.00047-19. [↩]
- J. C. Mao, M. Putterman, Accumulation in gram-positive and gram-negative bacteria as a mechanism of resistance to erythromycin, Journal of Bacteriology, Vol. 95, pg. 1111-1117, 1968, https://doi.org/10.1128/jb.95.3.1111-1117.1968. [↩]
- A. Yannarell, A. Kent, Bacteria, distribution and community structure, in Encyclopedia of Inland Waters, Vol. 2, pg. 820-831, 2009, https://www.d.umn.edu/biology/documents/Sterner1_000.pdf. [↩]
- I. Chopra, M. Roberts, Tetracycline antibiotics: Mode of action, applications, molecular biology, and epidemiology of bacterial resistance, Microbiology and Molecular Biology Reviews, Vol. 65, pg. 232-260, 2001, https://doi.org/10.1128/MMBR.65.2.232-260.2001. [↩]
- K. Farzam, T. A. Nessel, J. Quick, Erythromycin, in StatPearls [Internet], StatPearls Publishing, Treasure Island, FL, 2023, https://www.ncbi.nlm.nih.gov/books/NBK532249/. [↩]
- B. Pawłowska, M. Sysa, A. Godela, R. Biczak, Antibiotics amoxicillin, ampicillin and their mixture—impact on bacteria, fungi, ostracods and plants, Molecules, Vol. 29, pg. 4301, 2024, https://doi.org/10.3390/molecules29184301. [↩]
- J. C. Hanekamp, A. Bast, Antibiotics exposure and health risks: Chloramphenicol, Environmental Toxicology and Pharmacology, Vol. 39, pg. 213-220, 2015, https://doi/org/10.1016/j.etap.2014.11.016. [↩]
- Z. Breijyeh, B. Jubeh, R. Karaman, Resistance of gram-negative bacteria to current antibacterial agents and approaches to resolve it, Molecules, Vol. 25, pg. 1340, 2020, https://doi.org/10.3390/molecules25061340. [↩]