Abstract
Antimicrobial resistance (AMR) is a growing global health concern, driven in part by widespread empirical antibiotic use that overlooks individual biological variability. This exploratory pilot study aimed to characterize empirical prescribing practices in outpatient settings and to examine whether community-level antimicrobial responses differ among individuals sharing the same environment. A cross-sectional survey revealed that 97.4% of respondents had received empirically prescribed antibiotics without prior antibiotic susceptibility testing (AST), and 38.5% reported that the prescribed antibiotics were ineffective. To explore the potential biological variability, oral microbiota samples were assessed using community-level disk diffusion assays with five common antibiotics. The assays revealed reproducible interindividual differences in inhibition zone diameters. Notably, one participant consistently exhibited smaller inhibition zone diameters for ampicillin relative to the other participants, whereas streptomycin and chloramphenicol showed relatively uniform inhibition patterns across participants. These preliminary observations suggest that community-level antimicrobial responses may vary among individuals sharing the same environment. Larger studies integrating microbiome characterization and rapid susceptibility methods are needed to determine the mechanisms underlying this variability and to inform more personalized, evidence-based antimicrobial prescribing strategies.
Keywords: Antimicrobial resistance (AMR), Personalized antibiotic prescribing, Oral microbiota, Antibiotic Susceptibility testing (AST), Disk diffusion assay, and Empirical prescribing.
Introduction
Antibiotics have revolutionized modern medicine by dramatically reducing morbidity and mortality associated with bacterial infections. However, the global rise of antimicrobial resistance (AMR) has emerged as a critical public health threat. According to a recent Global Burden of Disease (GBD) study, bacterial AMR was directly responsible for approximately 1.14 million deaths and associated with 4.71 million deaths worldwide in 2021. Without effective intervention, this annual mortality toll is projected to increase sharply to 8.2 million by 20501. This alarming trend underscores the urgent need to better understand the biological and clinical factors contributing to antibiotic treatment failure and heterogeneous treatment outcomes2‘3.
One of the major drivers of AMR is the widespread use of empirical antibiotic prescribing, particularly in outpatient settings where treatment is often initiated without pathogen-specific or patient-specific diagnostic testing4‘5. Although empirical treatment is sometimes necessary in urgent clinical situations, it implicitly assumes that all patients will respond similarly to a given antibiotic regimen. Such an assumption overlooks substantial interindividual biological variability that can influence antimicrobial effectiveness. Host genetic factors represent one source of such variability. For example, genetic polymorphisms affecting drug-metabolizing enzymes, such as those in the cytochrome P450 family, can alter antibiotic pharmacokinetics, including absorption, distribution, and metabolism6‘7‘8. In addition to host genetic factors, host-associated microbiota represents an important yet relatively understudied determinant of interindividual variability in antibiotic response. Microbiome composition is highly personalized and shaped by multiple factors including diet, age, immune history, and environmental exposure9‘10‘11. Importantly, substantial interindividual variation persists even among individuals living in the same household, underscoring the individualized nature of the human oral microbiome12‘13. Large cohort and twin studies have further demonstrated that even genetically identical individuals can maintain distinct, person-specific microbial community signatures across adolescence and adulthood14‘15.
Despite increasing recognition of biological individuality, antibiotic susceptibility testing (AST) is rarely performed prior to prescribing in outpatient care due to time constraints, costs, and limited access to rapid diagnostics16‘17‘18. As a result, physicians often lack patient-specific biological information at the time of prescribing, potentially contributing to suboptimal treatment outcomes and the continued propagation of AMR. Therefore, it is essential to understand how individual-level biological factors shape antimicrobial response to improve prescribing practice and mitigate the progression of AMR.
To explore this gap, we conducted a pilot exploratory study integrating two complementary components: (1) a cross-sectional survey assessing empirical prescribing practices and perceived treatment effectiveness and (2) community-level disk diffusion assays using oral microbiota from the same household. The goal of this study was not to generate clinical conclusions but to provide preliminary, hypothesis-generating evidence of interindividual biological variability in antimicrobial response. Specifically, we sought to determine if personalized oral microbial communities exhibit distinct inhibition patterns, and whether this variability supports the potential need for future research toward more personalized, AST-informed prescribing strategies.
Methods
Survey Methodology
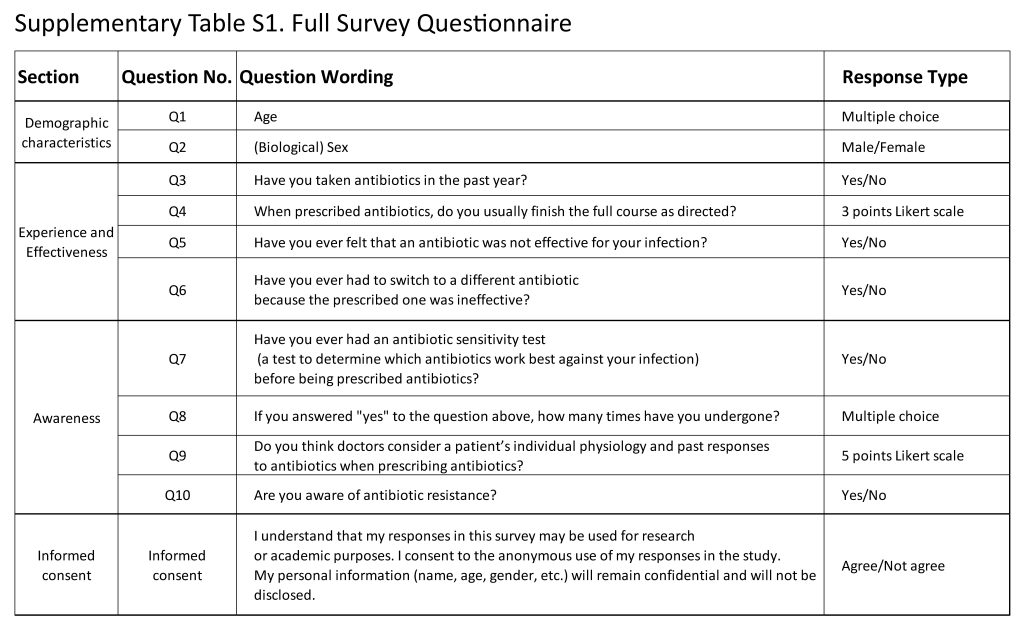
A cross-sectional survey was conducted to assess participants’ perceptions of antibiotic effectiveness and their awareness of antimicrobial susceptibility testing (AST). A total of 39 adult participants aged 21 to 60 years were recruited through workplace networks and extended family contacts using both in-person and online outreach. The survey was administered using a structured Google Forms questionnaire accessed via a QR code distributed either in person or through the KakaoTalk messaging platform. The 10-item questionnaire consisted of three sections: (1) demographic characteristics (age, gender); (2) experiences with antibiotic prescriptions and perceived effectiveness; and (3) awareness and prior experience with AST. All questions were closed-ended, utilized binary (yes/no) or 5-point Likert-scale response formats. Participation was voluntary, and informed consent was obtained from all participants. All survey responses were anonymized prior to analysis. The full questionnaire is provided in Supplementary Table S1.
Oral Microbiota Sample Collection
Oral microbiota samples were collected using a standardized, non-invasive swabbing procedure. Before sample collection, participants rinsed their mouths with 10 mL of sterile normal saline for 10 seconds to remove residual debris. Sterile cotton swabs were used to collect samples from the inner buccal mucosa and dorsal tongue surface using five uniform strokes at each site.
Oral microbiota from three participants (A, B and C) were streaked onto Mueller-Hinton agar (MHA) plates (100 mm diameter, 4 mm thickness; Carolina Biological Supply Company, Burlington, NC, USA) using a three-directional streaking method to promote uniform bacterial distribution. Plates were incubated at 37 ℃ for 24 h in a standard laboratory incubator (no active humidity regulation) under aerobic conditions to generate mixed polymicrobial growth characteristic of each participant.
Preparation of Community-Level Microbial suspensions
After 24 h, all visible colonies from each participant’s plate were gently scraped and resuspended in 500 μL of phosphate-buffered saline (PBS; pH 7.2; Thermo Fisher Scientific, Waltham, MA, USA). Suspensions were vortexed for 10 seconds to homogenize the microbial mixture. Inoculum turbidity was visually adjusted to achieve comparable opacity across participants. Subsequently, 100 μL of each suspension was evenly spread onto fresh MHA plates and allowed to air-dry for 3-5 minutes prior to antimicrobial susceptibility testing.
Antibiotic Susceptibility Testing (Disk Diffusion Assay)
Antimicrobial susceptibility was assessed using the Kirby-Bauer disk diffusion method, following Clinical and Laboratory Standards Institute (CLSI) guidelines where applicable for disk diffusion methodology. Five commonly used antibiotics were tested: ampicillin (10 μg), tetracycline (30 μg), streptomycin (10 μg), penicillin G (10 units), and chloramphenicol (30 μg) (6mm antibiotic disk; Carolina Biological Supply Company, Burlington, NC, USA). Antibiotic disks were placed on the inoculated agar surface in a randomized arrangement using a sterile disk dispenser, with a minimum spacing of 20 mm to prevent overlapping inhibition zones. Plates were incubated at 37 ℃ for 24 h in a standard laboratory incubator (no active humidity regulation) under aerobic conditions. Each antibiotic susceptibility assay was independently performed three times using freshly prepared microbial suspensions, yielding three independent biological replicates per antibiotic and participant (n = 3). For each biological replicate, inhibition zone diameters were measured at four directions around each antibiotic disk and averaged to obtain a single value per plate. Because this assay involved mixed oral microbial communities rather than isolated strains, CLSI interpretive categories (Susceptible/Intermediate/Resistant) were not applied. All measurements were interpreted only as relative differences in inhibition zone diameters.
Data Collection and Statistical Analysis
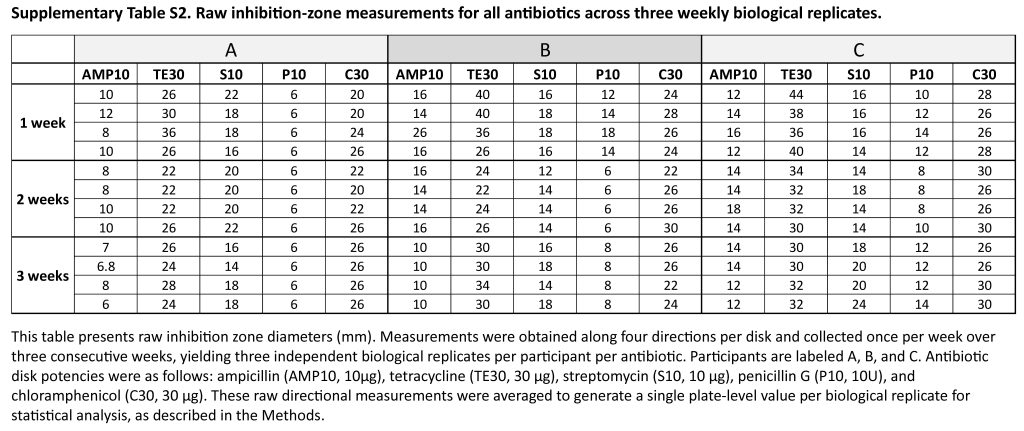
Inhibition zone diameters are represented as mean standard error (SE) calculated from three independent biological replicates (n = 3). Raw directional inhibition zone measurements for each antibiotic and participant are provided in Supplementary Table S2. Due to the pilot and exploratory nature of the study and the limited number of independent biological replicates (n = 3), inferential statistical testing (e.g., ANOVA, t-test) was not performed. Instead, all data are presented descriptively to illustrate the magnitude and reproducibility of interindividual differences observed across independent experiments.
Ethical Statement
This study involved anonymous survey responses and non-invasive oral swab collection, without the collection of identifiable personal or health information. All procedures were conducted in accordance with standard guidelines for minimal-risk human research. Informed consent was obtained from all participants prior to data collection, and all survey and microbiological data were anonymized before analysis. Because the study was exploratory and nonclinical, and involved minimal risk with anonymized data collection, it was considered exempt from formal institutional review. Therefore, formal IRB was not sought.
Results
Survey Findings
A total of 39 adults (aged 21-60 years; mean ± SE: 39.8 ± 1.7) with prior antibiotic use participated in the survey. Participants were recruited through workplace networks and online distribution, as described in the Methods. Among respondents, 97.4 % (n = 38) reported receiving antibiotics without prior antimicrobial susceptibility testing (AST), and 38.5 % (n = 15) reported that the prescribed antibiotic was ineffective, defined as a lack of symptom improvement, and 12.8 % (n = 5) reported requesting a change in medication due to perceived treatment failure. Although 74.4 % of respondents reported awareness of antimicrobial resistance (AMR), only 38.5 % (n = 15) believed that physicians consider individual biological traits when prescribing antibiotics (Table 1). Collectively, these results suggest empirical antibiotic prescribing is nearly universal among respondents and that patients perceive a significant lack of individual biological consideration in outpatient care.
| Variable | Response | n (%) | |
| Participant Characteristics | |||
| Total participants | 39 (100) | ||
| Age (years) | Mean±SE | 39.79 ± 1.73 | |
| Gender | Male | 26 (66.7) | |
| Female | 13 (33.3) | ||
| Antibiotic prescription experiences | |||
| Have you ever taken antibiotics | Yes | 39 (100) | |
| Received antibiotics without prior AST | Yes | 38 (97.4) | |
| Prescribed antibiotic perceived as ineffective | Yes | 15 (38.5) | |
| Requested change in antibiotic due to ineffectiveness | Yes | 5 (12.8) | |
| Had undergone AST prior to antibiotic prescription | Yes | 1 (2.6) | |
| Awareness and perceptions | |||
| Awareness of antimicrobial resistance | Yes | 29 (74.4) | |
| Believed physicians consider individual physiological traits when prescribing antibiotics | Yes | 15 (38.5) | |
*AST, antimicrobial susceptibility test; Ineffective: defined as a lack of symptom improvement.
Interindividual Differences in Cultured Oral Microbiota
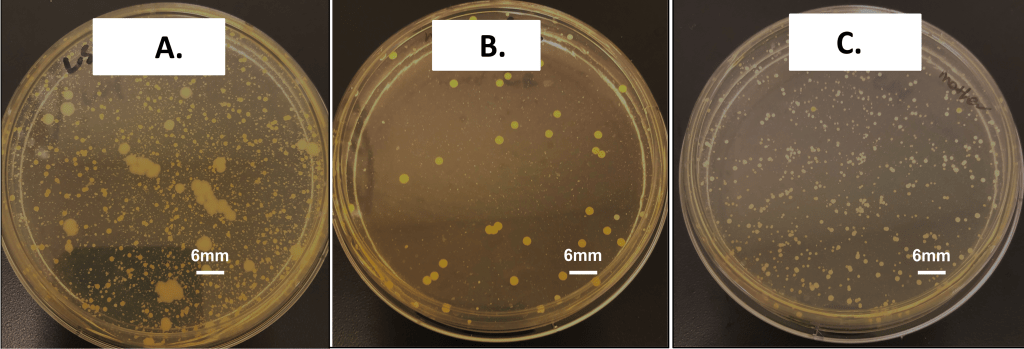
Oral microbiota cultured on Mueller-Hinton agar revealed visually distinct growth patterns across samples A, B, and C despite shared genetic and household environments (Figure 1). Sample A showed dense bacterial growth with highly heterogeneous colony morphologies, including several large mucoid-like colonies. Sample B showed fewer colonies with relatively uniform morphologies, whereas sample C displayed numerous small and evenly distributed colonies. Collectively, these distinct macroscopic characteristics indicate substantial person-specific heterogeneity in oral microbiota composition, even among individuals sharing genetics and the same household environment.
Interindividual variation in Antimicrobial Susceptibility
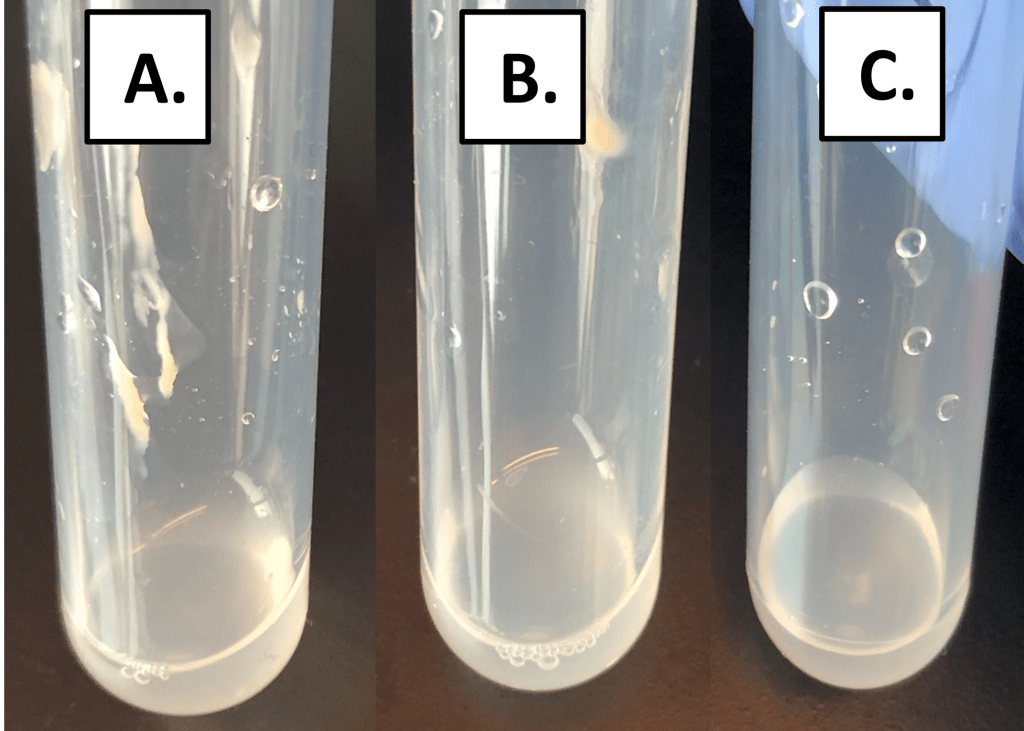
Visually standardized community-level microbial suspensions were prepared from each participant’s oral microbiota prior to susceptibility testing (Figure 2).

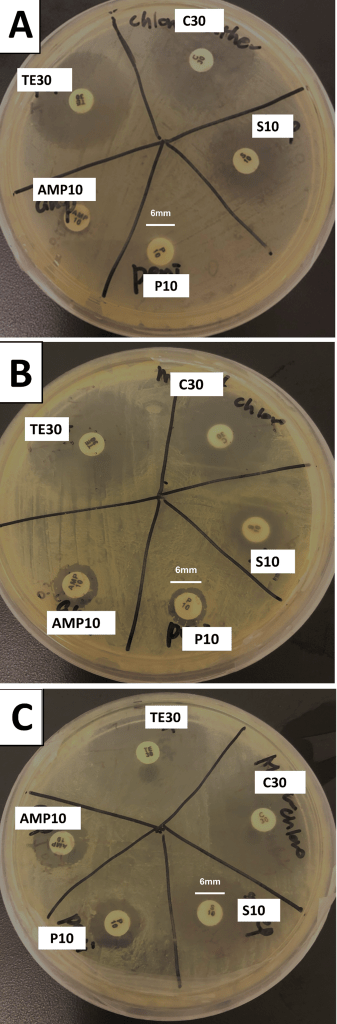
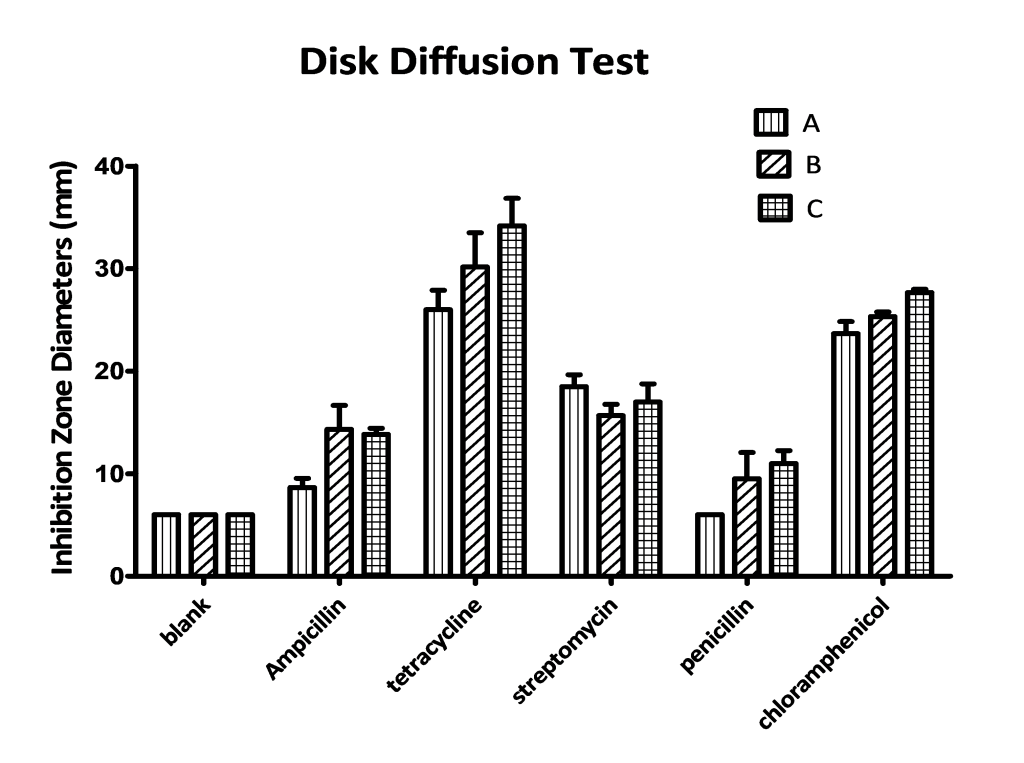
Disk diffusion assays revealed reproducible interindividual differences in inhibition zone diameters for ampicillin, penicillin G, and tetracycline (Table 2, Figure 3 and 4). Sample A showed consistently smaller inhibition zones for ampicillin (8.7 0.9 mm) relative to sample B (14.3 2.3 mm) and C (13.8 0.6 mm). Similar patterns were observed for penicillin G (A: 6.0 0.0 mm; B: 9.5 2.6 mm; C: 11.0 1.3 mm) and tetracycline (A: 26.0 1.9 mm; B: 30.2 3.3 mm; C: 34.2 2.7 mm). In contrast, streptomycin and chloramphenicol exhibited relatively uniform inhibition zone across all three samples.
| Antibiotic | Inhibition zone (mm ± SE) | ||
| A | B | C | |
| Ampicillin (AMP10, 10 μg) | 8.7 ± 0.9 | 14.3 ± 2.3 | 13.8 ± 0.6 |
| Tetracycline (T30, 30 μg) | 26.0 ± 1.9 | 30.2 ± 3.3 | 34.2 ± 2.7 |
| Streptomycin (S10, 10 μg) | 18.5 ± 1.2 | 15.7 ± 1.1 | 17.0 ± 1.8 |
| Penicillin (P10, 10 U) | 6.0 ± 0.0 | 9.5 ± 2.6 | 11.0 ± 1.3 |
| Chloramphenicol (C30, 30 μg) | 23.7 ± 1.2 | 25.3 ± 0.4 | 27.7 ± 0.3 |
| Blank Disk (Control) | 6 ± 0 | 6 ± 0 | 6 ± 0 |
*Values represent mean ± standard error (SE) from three independent weekly biological replicates (n = 3). Assays were conducted using mixed oral microbial communities, not isolated strains; therefore, CLSI susceptible/intermediate/resistant interpretive breakpoints do not apply. Inhibition zones reflect relative, community-level antimicrobial response under standardized aerobic incubation conditions (37 ℃, 24 h)
These inhibition patterns were consistent across three independent weekly biological replicates, indicating that the observed differences are likely due to participant-specific microbial profiles rather than week-to-week variability. Because these assays were conducted using mixed oral microbial communities rather than isolated strains, the inhibition zones represent relative community-level response, and CLSI clinical interpretive categories (Susceptible/Intermediate/Resistant) were not applied. Together, these results provide exploratory evidence of interindividual variability in antimicrobial response at the community level.
Discussion
This pilot exploratory study suggests that interindividual differences in community-level antimicrobial response may be observed even among individuals living in the same household. Although the experimental sample size (N = 3) was small, the reproducible inhibition-zone patterns observed across three weekly biological replicates suggest that the differences are unlikely to be solely attributable to procedural variation and likely reflex underlying biological variability. These observations are consistent with prior microbiome research demonstrating that oral microbiota communities are highly personalized and shaped by multiple factors, including diet, immune history, age, environmental exposures, and host genotype9‘10‘19. Furthermore, substantial person-specific variation is known to persist even among individuals sharing similar household environments12‘13‘14. Collectively, these findings indicate that individual microbial ecology may contribute to measurable differences in community-level antimicrobial responsiveness.
The cross-sectional survey provides complementary real-world context by characterizing how empirical prescribing is practiced in outpatient settings, whereas the experimental findings suggest potential biological variability. The majority of participants (97.4 %) reported receiving empirical antibiotic treatment without prior AST, and 38.5% perceived their prescribed antibiotic as ineffective. Although these perceptions are self-reported and cannot be clinically validated in this study, they are consistent with national reports demonstrating widespread empirical prescribing and frequent inappropriate antibiotic use in outpatient settings3‘4. Collectively, these observations underscore that patient-specific biological variability may not be routinely considered at the time of treatment selection.
The disk diffusion assays revealed reproducible participant-specific inhibition zone patterns, particularly for ampicillin and penicillin G, across three independent weekly biological replicates, while streptomycin and chloramphenicol exhibited relatively uniform inhibition zones across participants. This observed variability aligns with previous studies demonstrating that microbial community composition influence antimicrobial efficacy20, that metabolic interactions can modulate antibiotic tolerance within mixed-species communities21, and that resistant subpopulations can provide protective buffering to susceptible taxa during antibiotic exposure22. Collectively, these established mechanisms offer a plausible biological explanation for the interindividual variability in community-level antimicrobial responses observed in this pilot study.
Several methodological limitations should be considered when interpreting these findings. First, the disk diffusion assays were performed on mixed microbial community suspensions rather than isolated strains; therefore, the measured inhibition zones represent relative community-level responsiveness and cannot be interpreted using CLSI-defined susceptibility categories. Second, the inoculum turbidity was visually standardized rather than calibrated to a McFarland standard, which may introduce variability in bacterial density. Third, all assays were conducted under aerobic conditions, limiting the detection of obligate anaerobic taxa that comprise a substantial portion of the oral microbiota. Finally, molecular analyses such as rRNA sequencing or shotgun metagenomics were not performed to identify specific taxa or resistance genes, further constraining mechanistic interpretation.
Despite these limitations, the reproducibility of participant-specific inhibition patterns across independent replicates suggests that individualized microbial community features may contribute heterogeneous antimicrobial responses. Future studies incorporating CFU normalization, controlled anaerobic and aerobic culture conditions, sequencing-based microbial profiling, and host pharmacogenomic characterization will be essential for elucidating the microbial and host determinants of individual antimicrobial responses1823‘24. As rapid diagnostic technologies become increasingly accessible, such approaches may ultimately support the development of more personalized, evidence-based antimicrobial stewardship strategies that better align treatment decisions with patient-specific biological variability.
References
- GBD 2021 Antimicrobial Resistance Collaborators. Global burden of bacterial antimicrobial resistance 1990-2021: a systematic analysis with forecasts to 2050. Lancet (London, England). Vol. 404, pg. 1199–1226, 2024 https://doi.org/10.1016/S0140-6736(24)01867-1 [↩]
- C. L. Ventola. The antibiotic resistance crisis: part 1: causes and threats. P & T: A Peer-Reviewed Journal for Formulary Management. Vol. 40, pg. 277–283, 2015 [↩]
- C. Llor, L. Bjerrum. Antimicrobial resistance: risk associated with antibiotic overuse and initiatives to reduce the problem. Therapeutic Advances in Drug Safety. Vol. 5, pg. 229–241, 2014 https://doi.org/10.1177/2042098614554919 [↩] [↩]
- K. E. Fleming-Dutra, A. L. Hersh, D. J. Shapiro, M. Bartoces, E. A. Enns, T. M. File, J. A. Finkelstein, J. S. Gerber, D. Y. Hyun, J. A. Linder, R. Lynfield, D. J. Margolis, L. S. May, D. Merenstein, J. P. Metlay, J. G. Newland, J. F. Piccirillo, R. M. Roberts, G. V. Sanchez, K. J. Suda, A. Thomas, T. M. Woo, R. M. Zetts, L. A. Hicks. Prevalence of inappropriate antibiotic prescriptions among us ambulatory care visits, 2010-2011. JAMA. Vol. 315, pg. 1864–1873, 2016 https://doi.org/10.1001/jama.2016.4151 [↩] [↩]
- L. A. Hicks, M. G. Bartoces, R. M. Roberts, K. J. Suda, R. J. Hunkler, T. H. Taylor, S. J. Schrag. US outpatient antibiotic prescribing variation according to geography, patient population, and provider specialty in 2011. Clinical Infectious Diseases: An Official Publication of the Infectious Diseases Society of America. Vol. 60, pg. 1308–1316, 2015 https://doi.org/10.1093/cid/civ076 [↩]
- U. M. Zanger, M. Schwab. Cytochrome p450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacology & Therapeutics. Vol. 138, pg. 103–141, 2013 https://doi.org/10.1016/j.pharmthera.2012.12.007 [↩]
- G. R. Wilkinson. Drug metabolism and variability among patients in drug response. The New England Journal of Medicine. Vol. 352, pg. 2211–2221, 2005 https://doi.org/10.1056/NEJMra032424 [↩]
- S. Ahmed, Z. Zhou, J. Zhou, S.-Q. Chen. Pharmacogenomics of drug metabolizing enzymes and transporters: relevance to precision medicine. Genomics, Proteomics & Bioinformatics. Vol. 14, pg. 298–313, 2016 https://doi.org/10.1016/j.gpb.2016.03.008 [↩]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. Vol. 486, pg. 207–214, 2012 https://doi.org/10.1038/nature11234 [↩] [↩]
- B. P. Willing, S. L. Russell, B. B. Finlay. Shifting the balance: antibiotic effects on host-microbiota mutualism. Nature Reviews. Microbiology. Vol. 9, pg. 233–243, 2011 https://doi.org/10.1038/nrmicro2536 [↩] [↩]
- E. K. Costello, K. Stagaman, L. Dethlefsen, B. J. M. Bohannan, D. A. Relman. The application of ecological theory toward an understanding of the human microbiome. Science (New York, N.Y.). Vol. 336, pg. 1255–1262, 2012 https://doi.org/10.1126/science.1224203 [↩]
- Y. Zhou, K. A. Mihindukulasuriya, H. Gao, P. S. La Rosa, K. M. Wylie, J. C. Martin, K. Kota, W. D. Shannon, M. Mitreva, E. Sodergren, G. M. Weinstock. Exploration of bacterial community classes in major human habitats. Genome Biology. Vol. 15, pg. R66, 2014 https://doi.org/10.1186/gb-2014-15-5-r66 [↩] [↩]
- A. Spor, O. Koren, R. Ley. Unravelling the effects of the environment and host genotype on the gut microbiome. Nature Reviews. Microbiology. Vol. 9, pg. 279–290, 2011 https://doi.org/10.1038/nrmicro2540 [↩] [↩]
- S. S. Stahringer, J. C. Clemente, R. P. Corley, J. Hewitt, D. Knights, W. A. Walters, R. Knight, K. S. Krauter. Nurture trumps nature in a longitudinal survey of salivary bacterial communities in twins from early adolescence to early adulthood. Genome Research. Vol. 22, pg. 2146–2152, 2012 https://doi.org/10.1101/gr.140608.112 [↩] [↩]
- P. J. Turnbaugh, M. Hamady, T. Yatsunenko, B. L. Cantarel, A. Duncan, R. E. Ley, M. L. Sogin, W. J. Jones, B. A. Roe, J. P. Affourtit, M. Egholm, B. Henrissat, A. C. Heath, R. Knight, J. I. Gordon. A core gut microbiome in obese and lean twins. Nature. Vol. 457, pg. 480–484, 2009 https://doi.org/10.1038/nature07540 [↩]
- S. Hattab, A. H. Ma, Z. Tariq, I. Vega Prado, I. Drobish, R. Lee, R. Yee. Rapid phenotypic and genotypic antimicrobial susceptibility testing approaches for use in the clinical laboratory. Antibiotics (Basel, Switzerland). Vol. 13, pg. 786, 2024 https://doi.org/10.3390/antibiotics13080786 [↩]
- D. Trevas, A. M. Caliendo, K. Hanson, J. Levy, C. C. Ginocchio. Diagnostic tests can stem the threat of antimicrobial resistance: infectious disease professionals can help. Clinical Infectious Diseases: An Official Publication of the Infectious Diseases Society of America. Vol. 72, pg. e893–e900, 2021 https://doi.org/10.1093/cid/ciaa1527 [↩]
- D. Yamin, V. Uskoković, A. M. Wakil, M. D. Goni, S. H. Shamsuddin, F. H. Mustafa, W. A. Alfouzan, M. Alissa, A. Alshengeti, R. H. Almaghrabi, M. A. A. Fares, M. Garout, N. A. Al Kaabi, A. A. Alshehri, H. M. Ali, A. A. Rabaan, F. A. Aldubisi, C. Y. Yean, N. Y. Yusof. Current and future technologies for the detection of antibiotic-resistant bacteria. Diagnostics (Basel, Switzerland). Vol. 13, pg. 3246, 2023 https://doi.org/10.3390/diagnostics13203246 [↩] [↩]
- D. V. Patangia, C. Anthony Ryan, E. Dempsey, R. Paul Ross, C. Stanton. Impact of antibiotics on the human microbiome and consequences for host health. MicrobiologyOpen. Vol. 11, pg. e1260, 2022 https://doi.org/10.1002/mbo3.1260 [↩]
- C. G. Buffie, V. Bucci, R. R. Stein, P. T. McKenney, L. Ling, A. Gobourne, D. No, H. Liu, M. Kinnebrew, A. Viale, E. Littmann, M. R. M. van den Brink, R. R. Jenq, Y. Taur, C. Sander, J. R. Cross, N. C. Toussaint, J. B. Xavier, E. G. Pamer. Precision microbiome reconstitution restores bile acid mediated resistance to clostridium difficile. Nature. Vol. 517, pg. 205–208, 2015 https://doi.org/10.1038/nature13828 [↩]
- E. M. Adamowicz, J. Flynn, R. C. Hunter, W. R. Harcombe. Cross-feeding modulates antibiotic tolerance in bacterial communities. The ISME Journal. Vol. 12, pg. 2723–2735, 2018 https://doi.org/10.1038/s41396-018-0212-z [↩]
- E. A. Yurtsev, H. X. Chao, M. S. Datta, T. Artemova, J. Gore. Bacterial cheating drives the population dynamics of cooperative antibiotic resistance plasmids. Molecular Systems Biology. Vol. 9, pg. 683, 2013 https://doi.org/10.1038/msb.2013.39 [↩]
- A. van Belkum, T. T. Bachmann, G. Lüdke, J. G. Lisby, G. Kahlmeter, A. Mohess, K. Becker, J. P. Hays, N. Woodford, K. Mitsakakis, J. Moran-Gilad, J. Vila, H. Peter, J. H. Rex, W. M. Dunne, JPIAMR AMR-RDT Working Group on Antimicrobial Resistance and Rapid Diagnostic Testing. Developmental roadmap for antimicrobial susceptibility testing systems. Nature Reviews. Microbiology. Vol. 17, pg. 51–62, 2019 https://doi.org/10.1038/s41579-018-0098-9 [↩]
- L. Radlinski, B. P. Conlon. Antibiotic efficacy in the complex infection environment. Current Opinion in Microbiology. Vol. 42, pg. 19–24, 2018 https://doi.org/10.1016/j.mib.2017.09.007 [↩]