This study employs convolutional neural networks (CNNs) and the extensive CheXpert chest radiograph dataset (including more than 220,000 images) to diagnose 14 distinct chest, heart, and lung conditions from a single radiograph image. In real-world clinical settings, achieving such a comprehensive diagnosis from one x-ray scan is nearly impossible due to the high costs, lengthy diagnosis times, and inherent uncertainties. Physicians face significant challenges when diagnosing conditions solely from chest x-rays. This project aims to bridge this gap by demonstrating the feasibility and high accuracy of using CNNs for such diagnoses. The dataset comprises diverse radiograph images and associated demographic information, such as age ranges, sex, different radiograph views (frontal/lateral), ensuring broad representation. The model architecture integrates multiple layers of image analysis, including convolutional, pooling, and dense layers. The training process focused on optimizing parameters to enhance model accuracy. My deep convolutional neural network handles multivariate analysis, enabling the classification of a wide range of conditions efficiently. The model significantly reduces the diagnosis time from an average of 2 days to several weeks to around 2-3 minutes post-radiograph upload. The neural network achieved approximately 90% accuracy in diagnosing the 14 conditions, highlighting its potential for rapid and reliable clinical application. The findings suggest that neural network models can not only classify thoracic conditions from chest radiographs with high accuracy, but also offer enhanced efficiency and cost-effectiveness compared to traditional methods. This advancement represents a significant step forward in improving patient care and medical decision-making in pulmonology.
Keywords:
Convolutional neural networks, CheXpert dataset, chest radiographs, thoracic conditions, deep learning, machine learning, AI, medical diagnosis, pulmonology, healthcare
Introduction
In recent years, the medical field has increasingly turned to advanced technologies to enhance diagnostic accuracy and efficiency. Among these technologies, convolutional neural networks (CNNs) have emerged as powerful tools for image analysis, particularly in medical imaging1. Previous research1 has shown promising results in using CNNs for detecting specific conditions in medical images, such as identifying tumors or fractures. However, the challenge of diagnosing multiple conditions simultaneously from a single radiograph remains a significant hurdle2. The CheXpertdataset, comprising over 220,000 chest radiographs, provides an unprecedented opportunity to explore the potential of CNNs in diagnosing a wide range of thoracic conditions3.
Accurately diagnosing 14 different thoracic conditions from a single chest x-ray is a complex task that traditional methods struggle to achieve due to high costs, long processing times, and diagnostic uncertainty.4 This study addresses this critical challenge by leveraging CNNs to develop a model capable of diagnosing multiple conditions with high accuracy. By addressing this gap in the existing research, the study aims to enhance diagnostic processes, reduce time and costs, and improve patient outcomes.
The significance of this research lies in its potential to revolutionize diagnostic practices in pulmonology and radiology. By demonstrating the feasibility of using CNNs for comprehensive diagnosis from a single chest radiograph, this study contributes to the broader understanding of machine learning applications in medical imaging. The findings could lead to practical applications in clinical settings, where rapid and accurate diagnosis is crucial. Additionally, the research may pave the way for future studies exploring the integration of AI in other areas of medical diagnosis, ultimately contributing to more efficient healthcare delivery and better patient care. Rather than waiting days, if not weeks, for diagnosis, this paper aims to provide a fast, cost-effective, and accurate alternative for diagnosis.
The scope includes training a multi-label classification model on both frontal and lateral x-ray views, evaluating its diagnostic accuracy, and assessing its practical utility in reducing diagnosis time. The study emphasizes automated image-based diagnosis and does not extend to clinical integration, real-time deployment in hospitals, or the inclusion of non-imaging clinical data such as lab results or patient history.
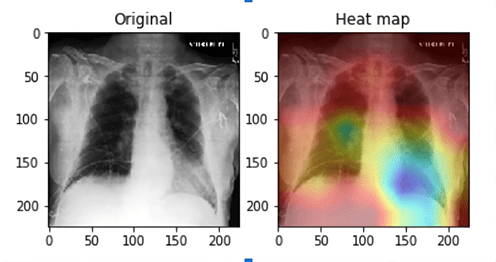
Limitations of the study include potential biases inherent in the CheXpert dataset, such as uneven class distributions, demographic imbalance, and the presence of uncertain or noisy labels. Additionally, resource constraints limited experimentation with multiple model architectures or extensive hyperparameter tuning. The model’s performance is also constrained by its reliance on static images, and it cannot account for temporal disease progression or real-time clinical context. Interpretability remains a challenge, as CNNs function are black-box models, which may hinder clinical trust and adoption.
This study is grounded in the theoretical framework of deep learning, specifically convolutional neural networks (CNNs), as a tool for AI-based image diagnosis. CNNs operate by hierarchically extracting certain features from images5, making them especially effective for analyzing complex medical imagery such as chest radiographs. The study also draws on the principles of multi-label classification, which allows the model to predict multiple co-occurring thoracic conditions from a single input6. This framework supports a shift from traditional, single-condition diagnostic models toward more holistic and data-driven methods. By applying this lens, the study seeks to demonstrate how CNNs can replicate and potentially augment radiological assessments, ultimately contributing to more efficient and scalable diagnostic workflows.
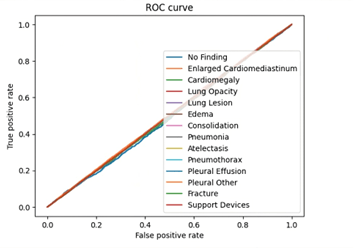
This research employed a quantitative, data-driven approach to train and evaluate a deep learning model for medical image classification. Using the publicly available CheXpert dataset of chest x-rays, the study applied convolutional neural networks to identify 14 thoracic conditions through multi-label classification. The dataset was preprocessed to ensure consistency and improve model generalizability, and a CNN architecture was trained on both frontal and lateral radiographs. Model performance was assessed using common classification metrics such as AUC, precision, and recall. This methodology allows for scalable analysis and lays the foundation for potential clinical applications of AI-assisted diagnosis.
Dataset Description
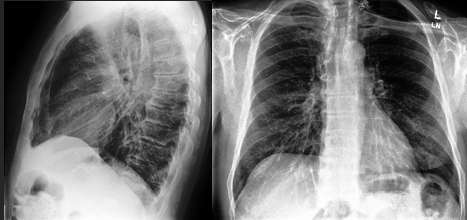
This study accesses and manipulates data from the Stanford CheXpert dataset3. This data came from over 225,000 different chest radiographs. The x-rays were collected from Stanford Health Care between October 2002 and July 2017 in inpatient and outpatient centers. This dataset includes over 200,000 images of frontal and lateral thoracic radiographs. The sheer size of the dataset contributes to the diversity and reach of this study. The dataset contains patients with a multitude of conditions, meaning this dataset would be applicable to many different respiratory or thoracic conditions and diseases. The dataset even accounts for patients with pacemakers or other support devices, making it even more implementable in a clinical setting. Additionally, the radiograph images were taken from a diverse set of patients further ensuring universality and accuracy.
This study includes convolutional neural network based diagnosis methods for the 14 different thoracic conditions included in the CheXpert dataset: enlarged cardiomediastinum, cardiomegaly, lung opacity, lung lesion, edema, consolidation, pneumonia, atelectasis, pneumothorax, pleural effusion, plural other, fracture, and support devices. Along with the condition stated in the data (or no finding, meaning no condition was present), the data’s columns included the path to the associated radiograph image, the sex of the patient, age, the view of the scan (frontal/lateral), and finally the type of radiation wave used for the x-ray (AP/PA). All of the above demographics, along with the type of condition, are crucial pieces of information and depending on each patient’s demographics, there are different approaches in examinations and diagnosis. Below are descriptions of each condition.
- Enlarged Cardiomediastinum: abnormal enlargement of the heart and other parts of the mediastinum (part of the chest that holds heart and other thoracic structures)
- Cardiomegaly: the heart is abnormally thick and/or large, making blood pumping difficult, caused by damage to heart muscle or abnormal speed of heartbeat
- Lung Opacity: opaque/hazy/dark areas shown in lung X-Rays (supposed to be translucent), often caused by liquids, cells, or substances that are in the pockets of the lungs supposed to be filled with air
- Lung Lesion: abnormal spots in the lung tissue such as nodules, masses, infiltrates, cysts, and granulomas caused by cancers, diseases, tumors, infections, and more
- Edema: excessive fluid buildup in the thoracic cavity, caused by heart failure and other conditions
- Consolidation: excessive density of the lung tissue, often caused by the replacement of air in the lungs with fluids like blood, pus, or cells
- Pneumonia: inflammation of the air sacs in the lungs caused by bacterial, viral, and fungal infections
- Atelectasis: complete or partial collapsing of the lung(s) caused by the alveoli failing to inflate the lung adequately (4 types: resorption, compression, contraction, and adhesive atelectasis)
- Pneumothorax: abnormal air accumulation in the pleural space (space between the lungs and chest wall), which often leads to the collapse of a lung(s) (2 types: spontaneous and traumatic pneumothorax)
- Pleural Effusion: abnormal fluid accumulation in the pleural space caused by disease, heart failure, cancer, and more
- Pleural Other: other abnormalities in the pleural space
- Fracture: break or crack in a rib or other bones in the thoracic area
- Support Devices: devices inside the thoracic area such as pacemakers that support the patient’s health
The wide diversity of the dataset was paired with data preprocessing methods and appropriate data splitting in order to ensure that the model was trained as fairly as possible and the results could apply to the largest audience possible. This will allow the methods in this paper to be applied to a wider medical audience rather than a more concentrated group.
Methodology
Data preprocessing
This data came from over 225,000 different chest radiographs. The x-rays were collected from Stanford Health Care between October 2002 and July 2017 in inpatient and outpatient centers. The sheer size of the dataset contributes to the diversity and reach of this study.
Along with exploring the massive dataset and its attributes, I implemented methods of data augmentation. Data augmentation methods include resizing the data, or iterating through each image and ensuring that they have the same size and dimensions7. I also employed other image preprocessing methods such as rotating the image, resizing it, and blurring it to reduce the amount of computational power necessary, considering the large amount of image data. This optimizes the model training while reducing the computing strain.
In addition to image augmentation, I used data generator functions such as flow_from_dataframe from libraries like ImageDataGenerator8. These functions read the image data from the CSV data file and convert them to images, and then randomly generate them for training usage. The data generators also are important to ensure the model is being trained in the right ways. For example, flow_from_dataframe includes important parameters such as x_col, y_col, target_size, batch_size, and class_mode. The x_col and y_col columns act as the input and output datasets. The x_col is the path of the image, which the model is taking as input. The y_col is the feature_string column which I created earlier, which includes all the “results” or outputs that the model is expected to output based on the input image data.
Finally, I iterated through the data and created a new column in the CSV file called ‘feature_string’ that creates a string of all the conditions that are present in the data. This way, rather than iterating through each individual condition column in the data and checking for the binary value (condition present or not present), the model can use a singular “results” column to compare back to the image given.
Model architecture
After completely preparing the data, I started creating the CNN model. After researching the different layers of CNN models, I learned that CNNs needed convolutional, pooling, and connected layers9.
- Convolutional Layer: This is the foundation of CNNs, and is where most of the computation is being done. The layer uses filters or kernels to extract features or key parts of the image to determine how to classify it. For example, for a dog some of its features could be its ears, soft fur, or four legs. CNNs are composed of multiple convolutional layers, with the first layers being the most general and the later ones being the most detailed.
- Pooling Layer: Pooling is used to decrease the number of pixels in the image, therefore these layers are used to increase efficiency. Although it may be thought that the more pixels or the more clear an image, the better a model would be trained, this unfortunately is not the case. If a model is trained on “non-pooled” images, it can take an enormous amount of time, memory, and storage to run, when it isn’t truly necessary. Therefore, the pooling stage can be thought of as decreasing the resolution of the image or making it more blurry to allow the model to better visualize the image without excessive computation.
- Fully Connected Layer: This is the most important layer in a CNN, as it is the layer that classifies the images based on the features extracted from the convolutional layers. These layers take in many different parameters adding to model density and makes it difficult to run due to GPU usage and time. Using too many fully connected layers may also risk overfitting, which would not make the model compatible in a clinical setting.
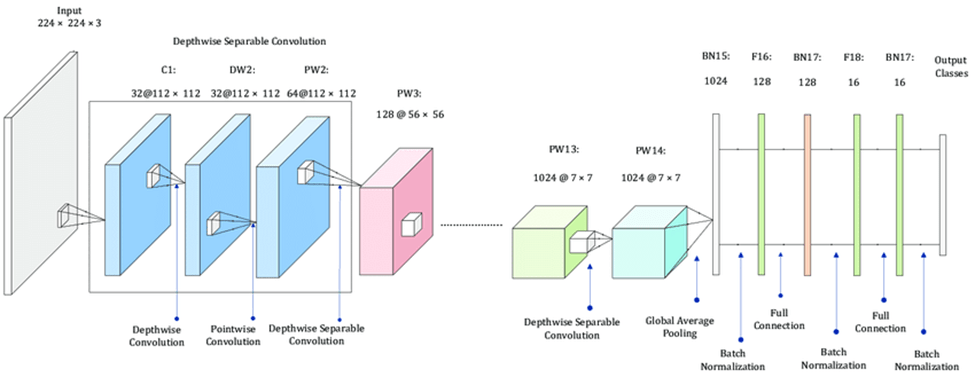
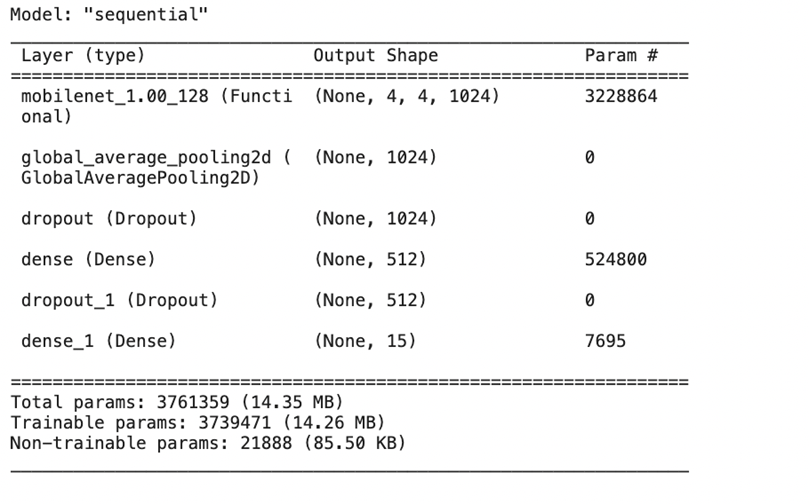
After researching each type of layer in a CNN model, I searched for existing convolutional, pooling, and fully connected layers that fit my requirements and special details of the dataset. I found a special type of layer called MobileNet that is used for creating deep neural networks, which was the type of neural network I was interested in creating. MobileNet10 uses several steps to break down the 3-dimensional image input into small groups of 2-dimensional data, and then break those groups down into 1-dimensional layers of data which are then processed in the feature layers. The feature layers recognize patterns in the image data that help them classify validation data. These patterns and learnings are then consolidated into the output classes which can predict based on the given input image data. I then added the Global Average Pooling 2D pooling layer11 to consolidate the image data. In between, I added dropout and dense layers to further optimize accuracy. After finding the best layers for the model, I put the whole model together.
Prior to training, every CNN model requires some baseline hyperparameters: an optimizer and a loss function12. Optimizers are algorithms used to adjust the weights and biases of a neural network to minimize the loss function during training. They play a crucial role in determining how the model learns from the data by controlling the direction and step size in the weight space. The goal of an optimizer is to find the set of parameters (weights) that minimize the loss function, thereby improving the model’s accuracy. This model implements the Adam optimizer13, a common optimizer for CNNs, because of its efficiency and ability to fit large datasets.
A loss function is a key component in training a neural network. It measures how well the model’s predictions match the actual target values. The loss function quantifies the difference between the predicted output (from the neural network) and the true labels (from the dataset). During training, the optimizer uses the gradients of the loss function to update the model’s weights, aiming to minimize the loss. For the model, I used the categorical cross-entropy loss function14, which is the most commonly used loss function for multiclass image classification tasks. It is particularly well-suited for problems where each input image is associated with a single class label out of multiple possible classes.
Implementing these two base hyperparameters ensures the best weights before training to ensure the best accuracy possible.
Model Training
Now that I had completed data collection, data preprocessing, building the custom CNN model architecture, and adding hyperparameters, I moved to training the model. Training any CNN model includes the above mentioned data preprocessing, along with data splitting. I employed a 80-20 split in the training vs testing data to ensure that the model was trained fairly and to prevent overfitting, where the model is tested on the same data it was trained upon, leading to false positive results and a false, higher accuracy rate. In order to create this split, I used Scikit learn’s train_test_split function15 to generate a fair split between training and validation data.
Results/Discussion
My deep learning model, trained on the CheXpert dataset, successfully diagnosed 14 thoracic conditions from chest radiographs with an accuracy of approximately 90%. Unlike traditional diagnostic methods that typically take 2 days to several weeks for radiological analysis and interpretation16, the model delivers results in a matter of seconds following radiograph upload. These findings highlight the capability of convolutional neural networks to perform multi-label classification tasks in complex medical imaging scenarios with both speed and precision.
The results of this study underscore the transformative potential of neural networks in medical diagnostics, particularly in radiology and pulmonology. By significantly accelerating the diagnostic process while maintaining high accuracy, my approach can reduce bottlenecks in clinical workflows, alleviate radiologist workload, and improve patient outcomes through earlier detection and treatment. The integration of such models into existing healthcare systems could lead to more scalable, cost-effective diagnostic solutions, especially in resource-limited settings. Theoretically, the study also validates the effectiveness of deep learning architectures for multi-label classification, contributing to the growing body of knowledge around AI applications in healthcare.
This study met its primary objectives: to implement advanced AI techniques for faster and more accurate diagnosis of thoracic conditions and to provide a cost-effective alternative to conventional diagnostic methods. By demonstrating that my CNN model could diagnose multiple conditions in seconds with high accuracy, the project successfully addressed both the time and accuracy challenges outlined in the problem statement. These results also validate my theoretical framework and justify the scope of the study.
Future research should explore the integration of non-imaging data, such as patient history or lab results, to enhance model performance and clinical relevance. Additionally, further validation across diverse patient populations and hospital settings is recommended to ensure the model’s robustness and generalizability. Comparative studies evaluating different neural network architectures, or combining them with techniques like attention mechanisms or transformers, may further improve interpretability and performance. Finally, collaboration with healthcare institutions could pave the way for clinical trials and real-world deployment.
The CheXpert dataset, while extensive, presents class imbalance and label uncertainty, which may have impacted model learning. The study was also constrained to a single model architecture due to limited computational resources, preventing experimentation with ensemble methods or extensive hyperparameter tuning. Moreover, the black-box nature of CNNs poses a challenge to clinical interpretability, potentially hindering adoption in real-world settings. The model also cannot account for disease progression over time due to its reliance on static images.
As AI continues to reshape the medical landscape, this study offers a glimpse into a future where diagnostics are not only faster and more accurate but also more accessible. While challenges remain, the integration of deep learning into diagnostic pipelines holds great promise for advancing patient care and democratizing access to high-quality healthcare worldwide.
References
- D. S. Boiselle, B. J. Bankier, S. M. Boiselle. Chest radiographs: Understanding opacity patterns. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC10103673/ (2023). [↩] [↩]
- D. J. Brenner, E. J. Hall. Computed tomography—An increasing source of radiation exposure. New Engl J Med. 357, 2277–2284 (2007). https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4954299/ (2007). [↩]
- AIMI Center. CheXpert: A large chest radiograph dataset for classification of thoracic diseases. https://aimi.stanford.edu/datasets/chexpert-chest-x-rays (2019). [↩] [↩]
- C. M. Wood, M. E. Murchison. Chest X-rays and other imaging tests. https://www.cancerresearchuk.org/about-cancer/cancer-unknown-primary-cup/getting-diagnosed/tests/x-rays (2023). [↩]
- MathWorks. Convolutional neural networks (CNN). https://www.mathworks.com/discovery/convolutional-neural-network.html (2023). [↩]
- A. O. Akintibu. Multilabel classification with CNN. https://aoakintibu.medium.com/multilabel-classification-with-cnn-278702d98c5b (2021). [↩]
- AI Multiple. Data augmentation techniques for deep learning. https://research.aimultiple.com/data-augmentation-techniques/ (2023). [↩]
- TensorFlow. ImageDataGenerator class. https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator (2024). [↩]
- S. Amidi. CS 230: Cheatsheet — Convolutional neural networks. https://stanford.edu/~shervine/teaching/cs-230/cheatsheet-convolutional-neural-networks (2019). [↩]
- Keras Documentation. MobileNet applications. https://keras.io/api/applications/mobilenet/ (2024). [↩]
- Keras Documentation. GlobalAveragePooling2D layer. https://keras.io/api/layers/pooling_layers/global_average_pooling2d/ (2024). [↩]
- Analytics Vidhya. Optimizer & loss functions in neural networks. https://medium.com/analytics-vidhya/optimizer-loss-functions-in-neural-network-2520c244cc22 (2020). [↩]
- PyTorch Documentation. torch.optim.Adam. https://docs.pytorch.org/docs/stable/generated/torch.optim.Adam.html (2024). [↩]
- PyTorch Documentation. torch.nn.CrossEntropyLoss. https://docs.pytorch.org/docs/stable/generated/torch.nn.CrossEntropyLoss.html (2024). [↩]
- Scikit-learn. train_test_split. https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html (2024). [↩]
- Mayo Clinic. Thoracic outlet syndrome: Diagnosis and treatment. https://www.mayoclinic.org/diseases-conditions/thoracic-outlet-syndrome/diagnosis-treatment/drc-20353994 (2023). [↩]