Abstract
Social learning, once thought to be limited to higher vertebrates, is now emerging as a powerful behavioral strategy even in insects. In this study, we demonstrate that honeybees can improve their maze navigation performance by visually observing the exploratory behavior of conspecifics. Observer bees showed significantly reduced flight times compared to untrained controls, indicating the presence of observational learning. Transcriptomic profiling revealed a marked upregulation of genes associated with synaptic plasticity and memory in observer bees, most notably Amfor. Feeding-based RNA interference targeting Amfor abolished the behavioral advantages gained from observation, confirming its essential role in mediating experience-driven learning. These findings suggest that Amfor is associated with neural and behavioral states that facilitate the encoding of visual experience into improved task performance. Our work highlights a genetically regulated mechanism of social learning in insects and underscores the broader relevance of foraging-related genes in cognitive plasticity across species.
Introduction
Honeybees (Apis mellifera) display a remarkable repertoire of cognitive behaviors, including sophisticated navigation, symbolic communication, associative learning, and long-term memory formation. Traditionally, many of these abilities were regarded as genetically hard-wired instincts shaped by natural selection1,2. However, a growing body of evidence demonstrates that honeybees are capable of flexible learning, experience-dependent behavioral modification, and environmentally driven cognitive plasticity3,4.
Recent studies reveal that insects—including bees—can learn not only through direct experience but also by observing conspecifics5,6. For example, bumblebees have been shown to acquire string-pulling and puzzle-solving behaviors after watching trained demonstrators7. Honeybees can learn route optimization from other foragers under certain conditions8, and observational learning has been proposed as a mechanism underlying social transmission of foraging strategies9. Despite these findings, the neural and molecular mechanisms enabling such observational learning remain almost entirely unexplored.
At the molecular level, one of the most well-characterized genes associated with honeybee behavioral plasticity is Amfor, which encodes a cGMP-dependent protein kinase (PKG)10. Amfor regulates the developmental transition from nursing to foraging behavior and modulates key behavioral traits including phototaxis, reward sensitivity, and exploratory tendencies10,11. In Drosophila, the orthologous foraging (for) gene influences learning, memory, and social behavior12, raising the possibility that Amfor may similarly contribute to higher-order cognitive processes in honeybees. Yet, no prior study has examined whether Amfor expression changes during social or observational learning, nor whether this gene is required for such learning to occur.
Thus, two major knowledge gaps remain unresolved:
(1) whether honeybees can acquire complex spatial information solely through visual observation of conspecifics, and
(2) what molecular pathways—including Amfor and plasticity-related genes—are activated during such observational learning.
To address these gaps, we designed a three-stage investigation:
(1) a behavioral assay testing whether honeybees improve maze navigation after watching demonstrators,
(2) transcriptomic profiling to identify molecular pathways enriched in observer bees, and
(3) RNA interference (RNAi) targeting Amfor to test its causal contribution to observational learning.
This study provides new evidence that honeybees use visual observational learning to optimize spatial navigation and identifies Amfor as a molecular correlate—and functional contributor—to this process.
Methods & Materials
Honeybee Collection and Differentiation
All honeybees used for RNA extraction (n=10 per group) were strictly standardized to control for age, behavioral role, and sex—factors known to influence Amfor expression. To eliminate variation arising from nurse-forager division of labor, we collected only newly emerged worker bees (24–48 hours old). Brood frames containing capped pupae were placed in a controlled incubator (34°C, 60% RH), and bees were collected immediately upon emergence using soft forceps. This procedure ensured that all individuals had not yet engaged in brood care, hive maintenance, or foraging, and thus remained task-naïve.
Only female worker bees were used; no drones or queens were included. After age-matching, bees were randomly assigned to the control (Group A) or observer (Group B) conditions. For RNA extraction, tissue was collected exclusively from these age-matched, behaviorally naïve workers, thereby preventing confounding differences in Amfor expression that are known to occur in older or foraging bees.
RNA Extraction and Sequencing
Honeybee brains were dissected on ice and homogenized in TRIzol™ reagent. Total RNA was extracted following the manufacturer’s protocol, including chloroform phase separation and isopropanol precipitation. RNA purity and integrity were confirmed using Nanodrop and Bioanalyzer. RNA-seq libraries (n = 3 pooled biological replicates per group) were sequenced to ~45M paired-end reads. High-quality RNA (RIN >7.0) was used for library construction (NEBNext® Ultra™ II RNA Kit), and sequencing was performed on an Illumina NovaSeq 6000 platform (150 bp paired-end). Reads were aligned to the Apis mellifera reference genome (Amel_HAv3.1), and differentially expressed genes were identified using DESeq2.
dsRNA synthesis
The target region of the Apis mellifera foraging gene (Amfor, NCBI Accession: AF469010) was selected based on sequence uniqueness and absence of off-target similarity. A ~200 bp fragment was amplified by PCR using primers appended with the T7 promoter sequence at the 5′ end of both forward and reverse primers:
- T7-Amfor-Forward: TAATACGACTCACTATAGGG…
- T7-Amfor-Reverse: TAATACGACTCACTATAGGG…
A scrambled sequence of the same length, with no homology to any known A. mellifera transcript, was used as a negative control:
- T7-Scrambled-Forward: TAATACGACTCACTATAGGGGATCGTACGTAGTCGATGCT…
- T7-Scrambled-Reverse: TAATACGACTCACTATAGGGGCTAGCTACGTACGATCGTC…
PCR products were purified using a standard gel extraction kit and used as templates for in vitro transcription following the manufacturer’s instructions for the MEGAscript™ T7 Transcription Kit (Invitrogen). The resulting sense and antisense RNAs were generated according to the kit protocol, annealed by controlled cooling to form double-stranded RNA (dsRNA), ethanol-precipitated, and resuspended in nuclease-free water. This procedure follows established honeybee RNAi methods described in Amdam et al. (2003)13 and Jarosch & Moritz (2011)14.
dsRNA feeding
Observer bees were fed with 50% sucrose solution supplemented with 1 µg/µL of Amfor dsRNA or scrambled dsRNA. Sterile filter paper disks soaked in the dsRNA-sucrose mixture were placed in Petri dishes, and bees were allowed to consume the solution ad libitum for 24 hours in a controlled incubator (33°C, 60% RH).
Gene expression analysis (RT-qPCR)
RT-qPCR was performed using SYBR Green Master Mix (Bio-Rad) with Amfor-specific primers, and AmActin was used as the reference gene. Relative expression levels were calculated using the ΔΔCt method.
| Gene name | Forward (5’ -> 3’) | Reverse (5’ -> 3’) |
| AmFor | ACCGACAATGCTTCCAAACC | AATGTGTCTCCTCTCGCTCC |
| AmCREB | ACGTCAGGAACAACACAGGT | TCAACCACCTGAAGCGTTTG |
| AmSynapsin | CCAGAGGGTTCAGAGCTTGA | TTGCTCCAGTCGGTGTTTTG |
Behavioral Assay (Flight Time, Maze Exploration)
Bees were individually introduced into a custom-built transparent maze arena (25 × 40 × 15 cm) sectioned off into 3 separate parts, with openings such that there was one available path. Bees were monitored under natural daylight conditions.
A single custom-built transparent maze arena was used for all trials to maintain consistency across experiments. After each bee completed a trial, the arena was cleaned with 70% ethanol and dried thoroughly before the next subject was tested.
Flight time was measured as the total duration (in seconds) that the bee remained airborne within the arena over a 5-minute observation period.
Path repetition was defined as the number of times the bee revisited previously explored regions, and stop events were counted when the bee remained immobile for more than 2 seconds. Behavioral recordings were analyzed using manual video tracking.
Gene Ontology (GO) enrichment analysis
GO analysis for Biological Process terms was performed using the clusterProfiler R package (v4.2.0). Differentially expressed genes (DEGs; FDR < 0.05) identified by DESeq2 were subjected to an over-representation analysis using a hypergeometric test. The Apis mellifera genome annotation (Amel_HAv3.1) served as the background gene set. P-values were adjusted using the Benjamini–Hochberg method, and GO terms with adjusted p < 0.05 were considered significantly enriched.
dsRNA Quality Control
The dsRNA products were run on a 1.5% agarose gel to confirm integrity and approximate size (~200 bp). Concentration and purity were measured using a NanoDrop spectrophotometer, and final dsRNA solutions were adjusted to 1 µg/µL for feeding.
Statistical Analysis
Data were presented as mean ± standard deviation (SD). Two-group comparisons (e.g., scrambled vs Amfor dsRNA) were performed using unpaired two-tailed Student’s t-tests. Significance was accepted at *p < 0.05, **p < 0.01, or ***p < 0.001.
Results
Honeybees learn through visual observation alone
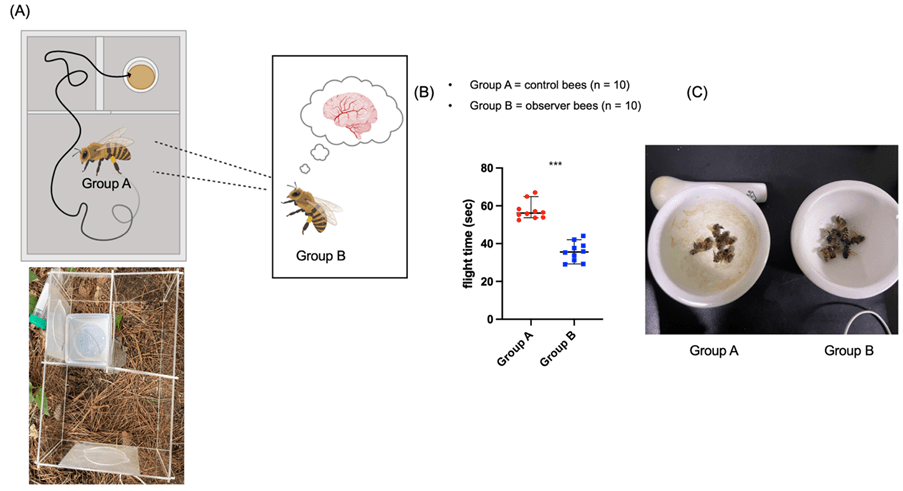
(B) Comparison of flight time between Group A and Group B. Observer bees (Group B) took significantly less time to complete the maze compared to control bees (Group A), demonstrating the effectiveness of visual observational learning. Data are presented as mean ± SEM. *** indicates p<0.001.
(C) RNA extraction from each bee group. Bees are dissected and processed for RNA extraction to be used for subsequent gene expression analysis, including RNA-seq and RT-qPCR.
Our behavioral experiments clearly demonstrated that honeybees can learn by watching others. Compared to the control group(Group A), which navigated the maze firsthand without prior observation, the observer bees (Group B), who watched Group A’s navigation through a transparent barrier, took a significantly shorter time to escape the maze (Figs. 1A and 1B). For observational trials, the demonstrator bee (Group A) was first introduced into the maze. Simultaneously, the observer bee (Group B) was placed in an adjacent compartment separated by a transparent acrylic partition, enabling unobstructed visual observation of the demonstrator’s movements. All experiments were conducted using a single demonstrator–observer pair per trial to prevent group interactions. This indicates that honeybees don’t just learn through trial and error; they can also efficiently acquire spatial information by observing the behavior of others and applying it to their own actions.
Observational Learning Alters Gene Expression in the Brain
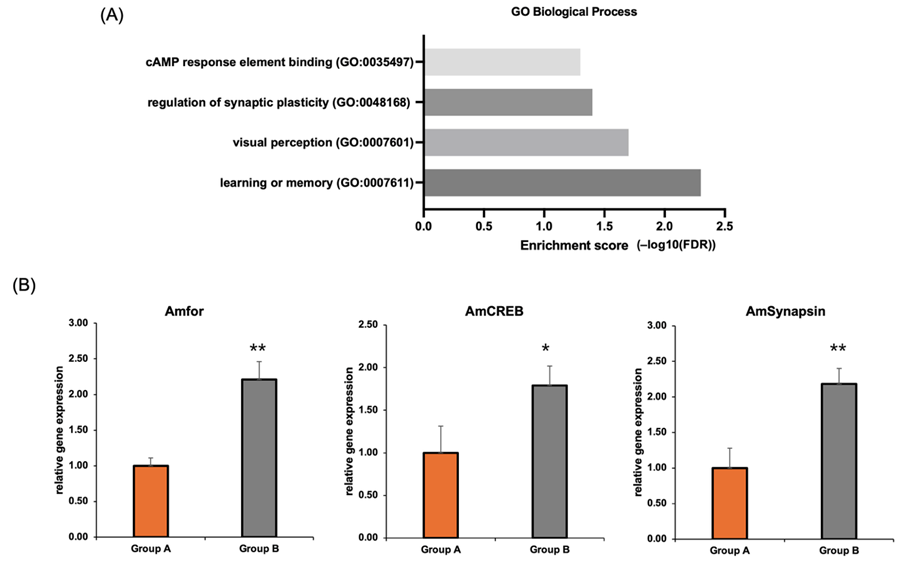
(B) Bar graphs showing the relative expression of key genes. The expression levels of AmCREB, Amfor, and AmSynapsin were significantly higher in observer bees (Group B) compared to control bees (Group A). Data are presented as mean ± SEM. * indicates p<0.05, ** indicates p<0.01.
To find the cause of this behavioral difference, we performed an RNA sequencing analysis on the honeybees. Surprisingly, the observer group’s brains showed a significant upregulation of genes related to memory formation and synaptic plasticity (Figs. 1c and 2A). To validate the RNA-seq results, we performed independent RT-qPCR measurements of three key genes—Amfor, AmCREB, and AmSynapsin. Consistent with the transcriptomic data, RT-qPCR confirmed significantly higher expression levels in observer bees compared to controls (Fig. 2B). This upregulation of AmCREB and AmSynapsin further supports the involvement of experience-dependent neural plasticity. CREB is a central transcription factor for long-term memory consolidation, while Synapsin regulates synaptic remodeling and vesicle dynamics. Their increased expression, together with elevated Amfor, suggests coordinated activation of molecular pathways underlying learning and memory during observational learning.
Amfor is a gene in honeybees (Apis mellifera) and is an ortholog of the foraging gene (for) in fruit flies (Drosophila melanogaster)10,15. This gene encodes a cGMP-dependent protein kinase (PKG) and is known to play a key role in regulating behavioral development throughout the honeybee’s life cycle, particularly the transition from indoor roles (foraging within the hive) to outdoor foraging behavior16.
Previous studies indicate that Amfor gene expression increases with the age of the honeybee and is more highly expressed in the brains of bees engaged in external activities including corpse removal, foraging. This increased Amfor expression induces the early expression of ‘positive phototaxis’, the behavior of moving toward light10. This suggests that Amfor does not merely direct foraging behavior itself, but rather plays a role in preparing the sensory and motor capabilities essential for foraging at the molecular level.
Amfor is a Key Regulator of Observational Learning
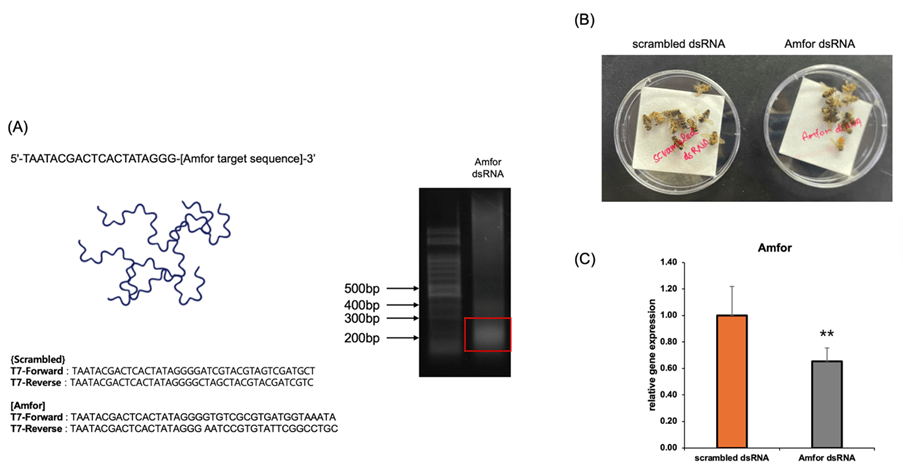
(B) Photo of the feeding-based RNAi delivery. The synthesized dsRNA was mixed with a sugar solution (50% sucrose) and fed to the bees for 24 hours. A scrambled dsRNA was used as a control.
(C) Validation of Amfor knockdown efficiency. RT-qPCR analysis confirmed a significant decrease in Amfor mRNA expression in the knockdown group, indicating the successful suppression of the target gene. Data are presented as mean ± SEM. ** indicates p<0.01.
To confirm whether the increased Amfor gene expression in observer bees was directly linked to improved spatial learning, we performed a feeding-based RNAi knockdown experiment. We selected a 200 bp target sequence from the Apis mellifera Amfor mRNA and synthesized double-stranded RNA (dsRNA) using primers with T7 promoter sequences (TAATACGACTCACTATAGGG) attached to both ends of the target sequence for in vitro transcription (Fig. 3A). The synthesized dsRNA was then mixed into a sugar solution (50% sucrose) and provided to the honeybees ad libitum for 24 hours. As a control, bees were fed scrambled dsRNA under identical conditions, and both groups were housed in petri dishes in the same environment (Fig. 3B). RT-qPCR confirmed that Amfor mRNA levels were reduced as expected in knockdown bees (Fig. 3C).
Amfor is required for the behavioral benefits of observational learning
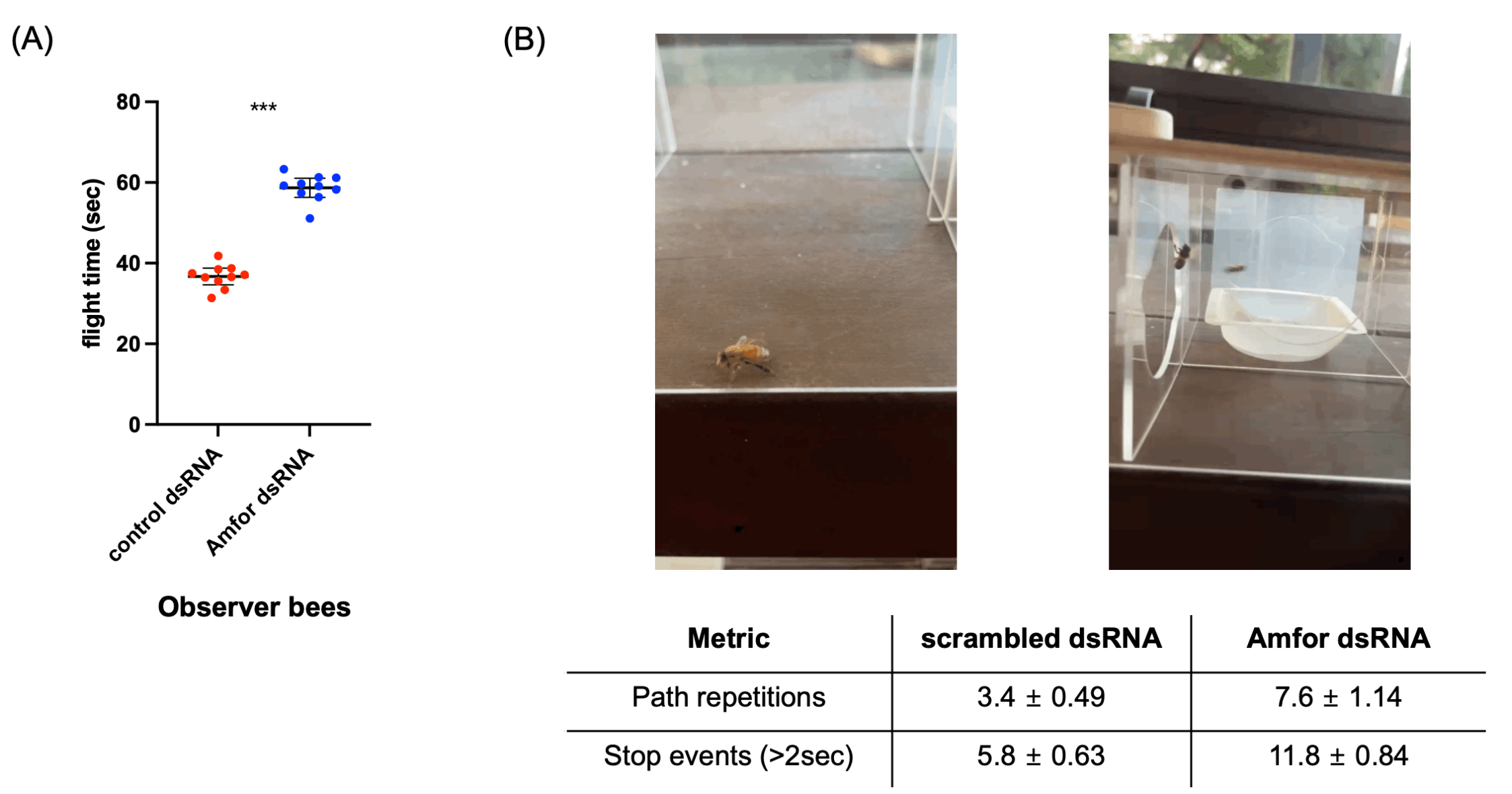

(B) Representative maze exploration videos comparing behavioral patterns of Amfor knockdown bees. Knockdown bees exhibited increased path repetition (left) more frequent stop events (>2 s immobility) (right), and less efficient maze progression compared to controls. These video examples illustrate the qualitative behavioral differences quantified in panel A.
To confirm whether the Amfor gene functionally contributes to the enhanced spatial navigation behavior gained through observer bees’ visual learning, we analyzed the behavior of observer bees. When observer bees were fed Amfor dsRNA, their maze exploration time (flight time) significantly increased back to control levels (Fig. 4A). This indicates that Amfor contributes to memory formation and path optimization through observation. Furthermore, knockdown bees showed decreased path efficiency and increased hesitation, with more path repetitions and stop events (Fig. 4B). However, we acknowledge that our current experimental design cannot fully exclude simpler mechanisms such as local enhancement, heightened arousal, or responses to general movement behind the barrier. While our results strongly support the presence of observational learning–like effects, additional control conditions (e.g., naïve demonstrators, moving-object controls, ghost controls) will be essential in future work to conclusively demonstrate true observational learning.
Discussion
This study demonstrates that the Amfor gene plays a critical role not only in classical foraging behavior, as previously known10, but also in a novel context of visual observational learning in honeybees. Our RNA-seq analysis revealed that genes associated with memory formation and synaptic plasticity were significantly upregulated in the brains of observer bees, among which Amfor exhibited more than a twofold increase in expression compared to control bees.
To functionally validate its role, we performed feeding-based RNA interference (RNAi) using double-stranded RNA targeting Amfor. Upon knockdown, observer bees lost the behavioral advantages gained from prior observation, including reduced maze exploration time, enhanced path efficiency, and decreased hesitation. These results indicate that Amfor contributes to experience-dependent behavioral optimization. Additionally, Amfor expression appears correlated with neural states that facilitate learning and plasticity.
Our findings support a causal relationship in which visual observation elevates Amfor expression in the brain, thereby facilitating memory formation and leading to behaviorally optimized navigation. This implies that a form of observational learning previously underexplored in honeybees enables bees to adapt to complex environments through enhanced learning and memory encoding.
The Amfor gene in honeybees shares significant homology with the foraging (for) gene in Drosophila melanogaster, which has also been implicated in behavioral plasticity, learning, and memory17. Previous studies in Drosophila have shown that for is expressed in specific brain clusters such as the central complex and optic lobes18,19, raising the possibility that Amfor may similarly exert region-specific functions in the honeybee brain. So, the loss of observational learning–derived behavioral optimization in Amfor knockdown bees is consistent with current understanding that Amfor/PKG activity facilitates experience-dependent plasticity20,21, sensory integration, and behavioral maturation in honeybees.
Intriguingly, for homologs are conserved in mammals and have been shown to influence reward-related behaviors and exploratory activity19,22. This suggests that the molecular mechanisms regulating behavioral plasticity in insects may be evolutionarily conserved and relevant to higher-order cognitive functions in vertebrates. As such, insect models can serve as powerful systems to study the gene–behavior–environment interface, offering fundamental insights into how experience is translated into lasting neural and behavioral changes. We refer to Amfor as a potential regulatory hub of experience-dependent neural plasticity because (i) its expression increases specifically in response to visual observational experience, (ii) it coincides with upregulation of additional plasticity-related genes such as AmCREB and AmSynapsin, and (iii) suppression of Amfor disrupts multiple components of learning-driven behavioral optimization rather than a single isolated behavior. These convergent lines of evidence suggest that Amfor modulates a broader plasticity network rather than acting on a single behavioral endpoint.
Despite these key findings, our study raises important questions regarding the neuroanatomical localization of Amfor activity during observational learning. Future work using in situ hybridization or immunostaining approaches will be essential to map its expression to specific brain regions and neural circuits.
Moreover, the ability of honeybees to learn by observation may offer an adaptive advantage in rapidly changing environments, allowing colonies to efficiently locate food sources and communicate valuable information23,24. This has implications for honeybee resilience in the face of modern ecological challenges, including pesticide exposure and climate change. It is also noteworthy that cognitive performance in bees has been shown to correlate with nutritional status25, suggesting that environmental factors can modulate gene expression and learning ability.
Taken together, our findings highlight Amfor as a potential convergence point for genetic and environmental signals that shape experience-dependent behavioral plasticity. Exploring the interaction between Amfor expression and nutritional status may uncover new insights into how environmental cues influence learning capacity at the molecular level.
References
- Beer, C. Niko Tinbergen and questions of instinct. Animal Behaviour 164, 261-265, doi:https://doi.org/10.1016/j.anbehav.2019.08.005 (2020). [↩]
- Hibbitt, C. Review: The Dancing Bees: Karl von Frisch and the Discovery of the Honeybee Language, by Tania Munz. The American Biology Teacher 81, 592-592, doi:10.1525/abt.2019.81.8.592 (2019). [↩]
- Arenas, A., Ramírez, G., Balbuena, M. S. & Farina, W. M. Behavioral and neural plasticity caused by early social experiences: the case of the honeybee. Frontiers in Physiology Volume 4 – 2013, doi:10.3389/fphys.2013.00041 (2013). [↩]
- Cabirol, A., Brooks, R., Groh, C., Barron, A. B. & Devaud, J. M. Experience during early adulthood shapes the learning capacities and the number of synaptic boutons in the mushroom bodies of honey bees (Apis mellifera). Learn Mem 24, 557-562, doi:10.1101/lm.045492.117 (2017). [↩]
- Giurfa, M. Social learning in insects: a higher-order capacity? Frontiers in Behavioral Neuroscience Volume 6 – 2012, doi:10.3389/fnbeh.2012.00057 (2012). [↩]
- Alem, S. et al. Associative Mechanisms Allow for Social Learning and Cultural Transmission of String Pulling in an Insect. PLOS Biology 14, e1002564, doi:10.1371/journal.pbio.1002564 (2016). [↩]
- Loukola, O. J., Solvi, C., Coscos, L. & Chittka, L. Bumblebees show cognitive flexibility by improving on an observed complex behavior. Science 355, 833-836, doi:doi:10.1126/science.aag2360 (2017). [↩]
- Dormagen, D. M., Wild, B., Wario, F. & Landgraf, T. Machine learning reveals the waggle drift’s role in the honey bee dance communication system. PNAS Nexus 2, doi:10.1093/pnasnexus/pgad275 (2023). [↩]
- Farina, W. M., Grüter, C. & Díaz, P. C. Social learning of floral odours inside the honeybee hive. Proceedings of the Royal Society B: Biological Sciences 272, 1923-1928, doi:doi:10.1098/rspb.2005.3172 (2005). [↩]
- Ben-Shahar, Y., Leung, H.-T., Pak, W. L., Sokolowski, M. B. & Robinson, G. E. cGMP-dependent changes in phototaxis: a possible role for the foraging gene in honey bee division of labor. Journal of Experimental Biology 206, 2507-2515, doi:10.1242/jeb.00442 (2003). [↩] [↩] [↩] [↩] [↩]
- Anreiter, I. & Sokolowski, M. B. The foraging Gene and Its Behavioral Effects: Pleiotropy and Plasticity. Annual Review of Genetics 53, 373-392, doi:https://doi.org/10.1146/annurev-genet-112618-043536 (2019). [↩]
- Osborne, K. A. et al. Natural Behavior Polymorphism Due to a cGMP-Dependent Protein Kinase of <i>Drosophila</i>. Science 277, 834-836, doi:doi:10.1126/science.277.5327.834 (1997). [↩]
- Amdam, G. V., Simões, Z. L. P., Guidugli, K. R., Norberg, K. & Omholt, S. W. Disruption of vitellogenin gene function in adult honeybees by intra-abdominal injection of double-stranded RNA. BMC Biotechnology 3, 1, doi:10.1186/1472-6750-3-1 (2003). [↩]
- Jarosch, A., Stolle, E., Crewe, R. M. & Moritz, R. F. Alternative splicing of a single transcription factor drives selfish reproductive behavior in honeybee workers (Apis mellifera). Proc Natl Acad Sci U S A 108, 15282-15287, doi:10.1073/pnas.1109343108 (2011). [↩]
- Heylen, K. et al. Amfor expression in the honeybee brain: a trigger mechanism for nurse-forager transition. J Insect Physiol 54, 1400-1403, doi:10.1016/j.jinsphys.2008.07.015 (2008). [↩]
- Ben-Shahar, Y. The foraging gene, behavioral plasticity, and honeybee division of labor. Journal of comparative physiology. A, Neuroethology, sensory, neural, and behavioral physiology 191, 987-994, doi:10.1007/s00359-005-0025-1 (2005). [↩]
- Mery, F., Belay, A. T., So, A. K.-C., Sokolowski, M. B. & Kawecki, T. J. Natural polymorphism affecting learning and memory in <i>Drosophila</i>. Proceedings of the National Academy of Sciences 104, 13051-13055, doi:doi:10.1073/pnas.0702923104 (2007). [↩]
- Sprecher, S. G., Reichert, H. & Hartenstein, V. Gene expression patterns in primary neuronal clusters of the Drosophila embryonic brain. Gene Expr Patterns 7, 584-595, doi:10.1016/j.modgep.2007.01.004 (2007). [↩]
- Miao, H., Li, W., Huang, Y. & Kim, W. J. The foraging gene coordinates brain and heart networks to modulate socially cued interval timing in Drosophila. PLoS Genet 21, e1011752, doi:10.1371/journal.pgen.1011752 (2025). [↩] [↩]
- Ben-Shahar, Y., Robichon, A., Sokolowski, M. B. & Robinson, G. E. Influence of Gene Action Across Different Time Scales on Behavior. Science 296, 741-744, doi:doi:10.1126/science.1069911 (2002). [↩]
- Elekonich, M. M. & Roberts, S. P. Honey bees as a model for understanding mechanisms of life history transitions. Comparative Biochemistry and Physiology Part A: Molecular & Integrative Physiology 141, 362-371, doi:https://doi.org/10.1016/j.cbpb.2005.04.014 (2005). [↩]
- Belay, A. T. et al. The foraging gene of Drosophila melanogaster: Spatial-expression analysis and sucrose responsiveness. Journal of Comparative Neurology 504, 570-582, doi:https://doi.org/10.1002/cne.21466 (2007). [↩]
- Dawson, Erika H., Avarguès-Weber, A., Chittka, L. & Leadbeater, E. Learning by Observation Emerges from Simple Associations in an Insect Model. Current Biology 23, 727-730, doi:https://doi.org/10.1016/j.cub.2013.03.035 (2013). [↩]
- Bidari, S., Peleg, O. & Kilpatrick, Z. P. Social inhibition maintains adaptivity and consensus of honeybees foraging in dynamic environments. Royal Society Open Science 6, 191681, doi:doi:10.1098/rsos.191681 (2019). [↩]
- Arien, Y., Dag, A., Zarchin, S., Masci, T. & Shafir, S. Omega-3 deficiency impairs honey bee learning. Proc Natl Acad Sci U S A 112, 15761-15766, doi:10.1073/pnas.1517375112 (2015). [↩]






