Abstract
Cancer is a multifactorial disease driven by the interaction of genetic susceptibility, epigenetic regulation, and environmental influences within the tumor microenvironment. While advances in targeted therapy and immunotherapy have improved outcomes for select malignancies, therapeutic resistance remains a major obstacle to durable clinical success. This review synthesizes current literature examining how inherited and acquired genetic alterations interact with microenvironmental factors—including immune infiltration, stromal remodeling, chronic inflammation, and hypoxia—to promote tumor progression and resistance to treatment. Evidence from genomic, epigenomic, and translational studies demonstrates that tumor heterogeneity, epigenetic plasticity, and microenvironment-mediated signaling enable cancer cells to adapt under therapeutic pressure. Inflammatory pathways and cancer stem cell dynamics further reinforce resistance across tumor types. Together, these findings highlight the limitations of therapies that target tumor cells in isolation and emphasize the need for integrative strategies that address both tumor-intrinsic alterations and microenvironmental support systems.
Introduction
Cancer encompasses a diverse group of diseases characterized by uncontrolled cellular proliferation, genomic instability, and the capacity for invasion and metastasis1. Despite extensive advances in molecular oncology, cancer remains difficult to treat effectively due to its biological heterogeneity and adaptive capacity. Increasing evidence indicates that tumor behavior cannot be fully explained by genetic alterations alone but instead emerges from dynamic interactions between malignant cells and their surrounding microenvironment2,3.
Genetic susceptibility plays a foundational role in oncogenesis, with inherited and somatic mutations influencing DNA repair, cell cycle regulation, and apoptotic pathways1. However, tumors evolve within complex microenvironments composed of immune cells, fibroblasts, endothelial cells, and extracellular matrix components. These elements actively shape selective pressures, facilitate immune evasion, and promote phenotypic plasticity, contributing to treatment failure even after initial therapeutic response4,5. Although genetic drivers and microenvironmental influences have been studied independently, fewer syntheses focus on how these factors converge to drive therapeutic resistance across cancer types. This gap limits the development of durable treatment strategies capable of preventing adaptation and relapse. This review integrates current evidence on how genetic susceptibility and tumor microenvironmental dynamics interact to promote therapeutic resistance, with an emphasis on mechanistic insight rather than primary clinical trial outcomes.
Literature Search Strategy
A structured literature review was conducted using PubMed, Google Scholar, and Web of Science to identify peer-reviewed articles published between 2000 and 2023. Search terms included combinations of “tumor microenvironment,” “genetic susceptibility,” “epigenetics,” “therapeutic resistance,” “cancer stem cells,” and “inflammation.” Studies were included if they provided mechanistic or translational insight into how genetic or microenvironmental factors influence tumor progression or treatment response. Both primary research articles and high-quality review papers were considered to allow integration of experimental findings with conceptual frameworks6,7. Studies lacking methodological transparency or relying on non-peer-reviewed sources were excluded. Findings were synthesized using a narrative, theme-based approach.
Tumor Microenvironment–Mediated Therapeutic Resistance
Therapeutic resistance is increasingly recognized as an emergent property of interactions between malignant cells and the tumor microenvironment rather than a consequence of tumor-intrinsic mutations alone5,3. The tumor microenvironment consists of immune cells, fibroblasts, endothelial cells, and extracellular matrix components that collectively influence drug response by regulating immune surveillance, nutrient availability, and survival signaling. Immune cell composition plays a critical role in determining therapeutic outcomes, as chronic inflammatory signaling can promote immunosuppressive phenotypes that blunt cytotoxic immune responses and reduce the effectiveness of immunotherapies4,8. Cancer-associated fibroblasts further remodel the extracellular matrix, creating physical barriers that limit drug penetration while secreting growth factors that support tumor survival.
Tumor Microenvironment–Driven Transcriptional Reprogramming
Beyond cellular composition, the tumor microenvironment exerts a powerful influence on cancer progression through transcriptional reprogramming. Microenvironmental stressors such as hypoxia, inflammatory cytokines, nutrient deprivation, and immune-mediated pressure activate coordinated gene expression programs that allow cancer cells to survive under adverse conditions and therapeutic stress9,6. These transcriptional adaptations promote phenotypic plasticity, enabling tumor cells to transiently adopt states associated with immune evasion, altered metabolism, and resistance to apoptosis.
Importantly, these changes are often reversible and context dependent, distinguishing them from permanent genetic mutations. Studies have consistently reported that genes involved in immune modulation, stress response, cell survival, and metabolic regulation display environment-specific expression patterns across tumors1. As a result, tumors with similar genetic drivers may respond differently to therapy depending on microenvironmental conditions, contributing to heterogeneous treatment outcomes observed in clinical practice.
This transcriptional flexibility represents a central mechanism by which tumors adapt to therapeutic pressure without requiring additional mutational events. Understanding how microenvironmental signals reshape gene expression therefore provides critical insight into the emergence of resistance and highlights the limitations of therapies that fail to account for dynamic tumor states.
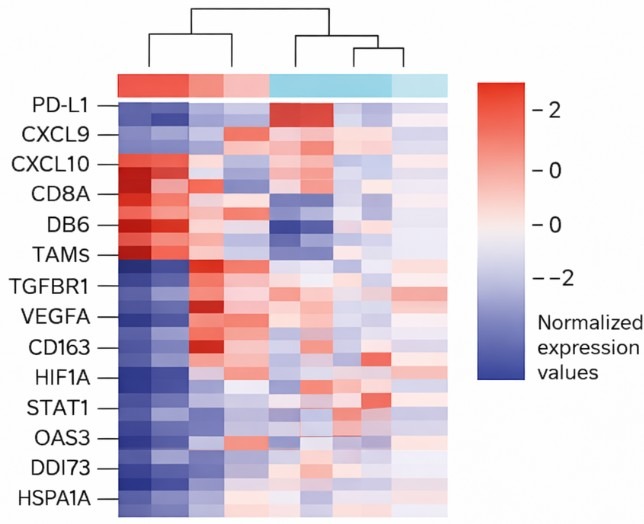
Conceptual illustration of representative gene expression patterns associated with varying tumor microenvironmental conditions. Genes commonly discussed in the literature, including immune- and stress-response–associated markers, are shown to illustrate how transcriptional programs may shift in response to inflammatory and immune signaling. This figure is provided as a schematic summary of published findings and does not represent original experimental data.
Environmental and Lifestyle Factors as Systemic Modulators of Cancer Risk
Environmental and lifestyle exposures act as upstream modulators of cancer risk by interacting with genetic susceptibility and shaping tumor-permissive microenvironments. Among these factors, tobacco smoking remains one of the most extensively studied carcinogenic exposures, with well-established links to malignancies affecting the lung, bladder, gastrointestinal tract, and cardiovascular system4,8. Smoking-related carcinogens induce direct DNA damage while simultaneously promoting chronic inflammation and oxidative stress, accelerating the accumulation of oncogenic alterations.
Crucially, the oncogenic impact of environmental exposures extends beyond localized tissue injury. Persistent exposure to carcinogens can induce systemic inflammation, immune dysregulation, and vascular dysfunction, creating physiological conditions that support tumor initiation and progression across multiple organ systems2,10. These systemic effects help explain why environmental risk factors influence cancer susceptibility in tissues distant from the original exposure site and why lifestyle factors can exacerbate genetically mediated cancer risk.
By linking environmental exposures to both genetic instability and sustained inflammatory signaling, this body of evidence underscores the importance of considering systemic physiological context when evaluating cancer risk and therapeutic resistance.
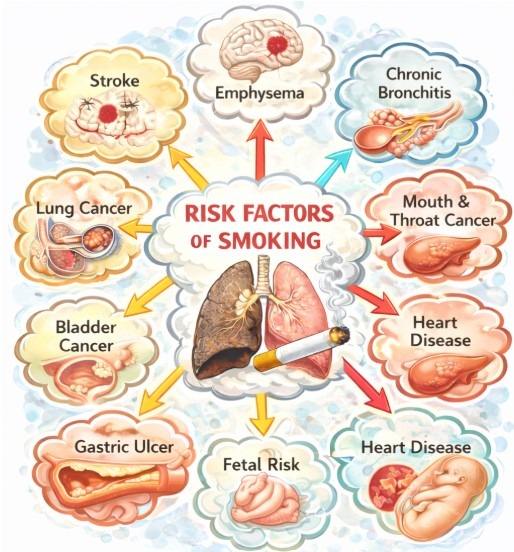
Schematic overview of systemic health effects and cancer risk factors associated with tobacco smoking. Smoking-related carcinogens contribute to DNA damage, chronic inflammation and immune dysregulation across multiple organ systems. This figure serves as a contextual illustration of well-established epidemiological relationships rather than as primary scientific evidence.
Histopathological Consequences of Malignant Transformation
The cumulative effects of genetic alterations and microenvironmental stress are reflected in distinct histopathological changes that characterize malignant tissue. Across epithelial cancers, malignant transformation is associated with increased cellular density, loss of organized tissue architecture, nuclear atypia, and impaired differentiation11,5. These morphological features arise from dysregulated proliferation, defective apoptotic signaling, and altered interactions between tumor cells and their surrounding stroma.
In lung cancer, malignant tissue frequently exhibits dense, disorganized cellular arrangements that contrast sharply with the thin, structured alveolar architecture of healthy lung tissue. Such alterations compromise normal gas exchange and contribute directly to clinical symptoms, while also providing diagnostic cues used in histopathological evaluation4,9. These structural changes are not merely descriptive but reflect underlying molecular and microenvironmental processes driving tumor progression.
Histopathology therefore serves as an important bridge between molecular mechanisms and clinical manifestation, linking genetic susceptibility and microenvironmental influence to observable disease outcomes.
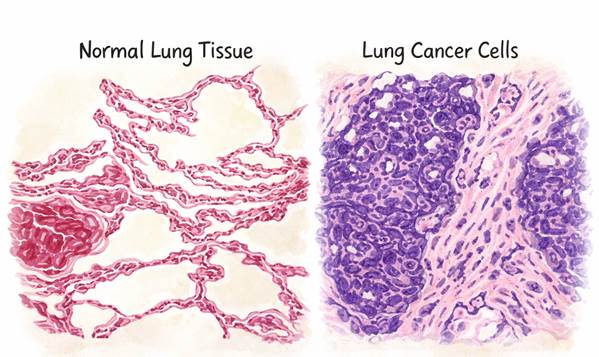
Representative histological comparison between normal lung tissue and lung cancer tissue, highlighting differences in tissue organization, cellular density, and nuclear morphology commonly described in pathological studies. The image illustrates structural changes associated with malignant transformation and is intended for educational context.
Carcinogenesis and Oncogenesis
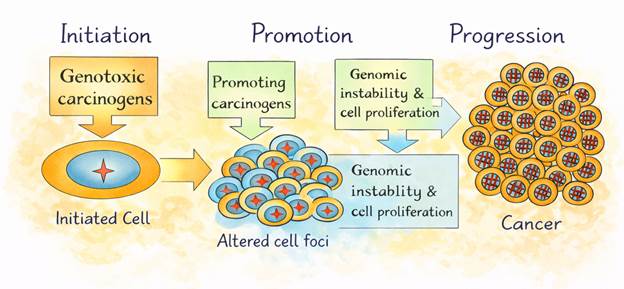
Carcinogenesis is a multistep process through which normal cells acquire malignant properties via the accumulation of genetic and epigenetic alterations. Classical models describe this progression in terms of initiation, promotion, and progression, with environmental exposures often acting as initiating or promoting agents1. During initiation, genotoxic stress induces mutations that disrupt normal regulatory pathways governing cell growth and survival. Promotion involves selective expansion of altered cells under favorable microenvironmental conditions, while progression reflects the accumulation of additional changes that enable invasion and metastasis.
This framework highlights how genetic susceptibility and environmental context cooperate over time to drive malignant transformation. Importantly, the tumor microenvironment influences each stage of carcinogenesis by shaping selective pressures that determine which cellular clones expand and persist. As a result, oncogenesis is best understood not as a linear genetic process but as an adaptive evolutionary trajectory influenced by both intrinsic and extrinsic factors.
Schematic representation of the classical multistage model of carcinogenesis, including initiation, promotion, and progression. The figure illustrates how genetic mutations and environmental exposures cooperate over time to drive malignant transformation.
Hallmarks of Cancer as a Framework for Therapeutic Resistance
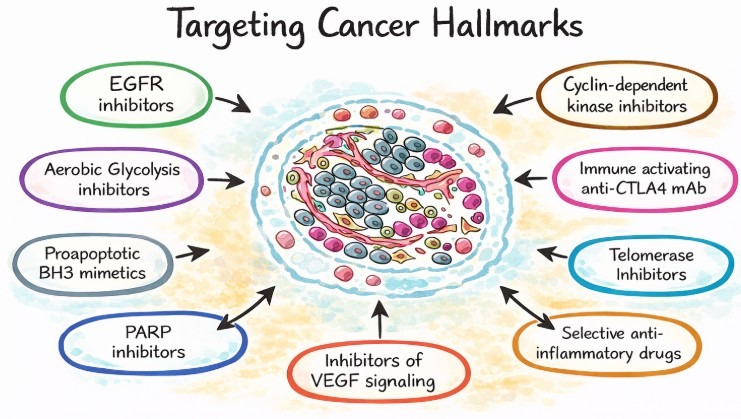
The Hallmarks of Cancer framework provides a unifying model for understanding how genetic and microenvironmental factors collectively drive tumor progression and resistance to therapy5. Hallmarks such as sustained proliferative signaling, immune evasion, resistance to cell death, angiogenesis, and tumor-promoting inflammation are not independent processes but are reinforced by reciprocal interactions between cancer cells and their surrounding microenvironment.
Overview of the Hallmarks of Cancer and representative therapeutic strategies targeting these biological capabilities. The figure summarizes how tumor-intrinsic and microenvironment-mediated processes contribute to resistance and treatment failure.
Epigenetic Plasticity and Cancer Stem Cell Dynamics
Epigenetic regulation provides cancer cells with a reversible mechanism for adapting to environmental stress without altering DNA sequence12. DNA methylation and histone modification can silence tumor suppressor genes or activate oncogenic pathways, enabling rapid phenotypic shifts in response to therapy. These changes contribute to tumor heterogeneity and undermine long-term treatment efficacy.
Cancer stem cell populations further exacerbate resistance by maintaining self-renewal capacity and surviving therapies that target rapidly dividing cells13. Microenvironmental cues such as hypoxia and inflammation support the maintenance of these populations, reinforcing resistance and contributing to disease recurrence.
Emerging Technologies as Investigative Tools
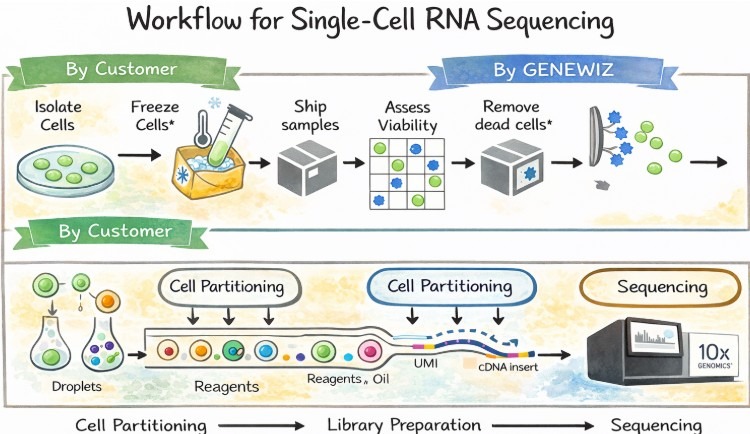
Advances in experimental and computational technologies have expanded understanding of tumor heterogeneity and resistance mechanisms. Single-cell RNA sequencing has enabled high-resolution characterization of transcriptional diversity within tumors and their microenvironments, revealing rare cell populations and dynamic cellular states associated with therapeutic resistance14,7. These approaches have reshaped understanding of intratumoral heterogeneity and highlighted the role of microenvironment-dependent adaptation.
Conceptual workflow of single-cell RNA sequencing illustrating major analytical stages commonly reported in cancer research literature. The figure is included to support discussion of technological advances and does not represent original experimental methods used in this review.
Conclusion
Cancer development and therapeutic resistance arise from a complex interplay between genetic susceptibility, epigenetic regulation, environmental exposures, and tumor microenvironmental dynamics. This review synthesizes evidence demonstrating that resistance to treatment is rarely driven by tumor-intrinsic mutations alone but instead emerges through continuous interaction between malignant cells and their surrounding cellular and biochemical environment. Immune modulation, stromal remodeling, inflammatory signaling, and epigenetic plasticity collectively enable tumors to adapt under therapeutic pressure and evade sustained control.
Several key insights emerge from the literature. Tumor heterogeneity and phenotypic plasticity, shaped by microenvironmental cues, undermine the long-term effectiveness of therapies that target single molecular pathways. Epigenetic mechanisms and cancer stem cell populations provide reversible and adaptive routes to resistance, contributing to disease recurrence even after an initial therapeutic response. Chronic inflammation further bridges environmental exposures and genetic instability, reinforcing resistance-promoting conditions across cancer types.
Despite significant advances in cancer biology, current therapeutic strategies often fail to fully account for the dynamic and reciprocal nature of tumor–microenvironment interactions. Treatments that focus exclusively on tumor cell–intrinsic targets may therefore be insufficient to prevent adaptation and relapse. Integrative approaches that simultaneously address genetic alterations and microenvironmental support systems represent a promising direction for improving therapeutic durability. Continued progress will depend on systems-level investigations, longitudinal studies of tumor evolution, and the responsible integration of emerging analytical technologies to advance precision oncology.
References
- Vogelstein, B. & Kinzler, K. W. 2015. Cancer genes and the pathways they control. Nature Medicine. [↩] [↩] [↩] [↩]
- Hanahan, D. & Weinberg, R. A. 2000. The hallmarks of cancer. Cell. [↩] [↩]
- McAllister, S. S. & Weinberg, R. A. 2014. Tumor–host interactions: A far-reaching relationship. Journal of Clinical Investigation. [↩] [↩]
- Coussens, L. M. & Werb, Z. 2002. Inflammation and cancer. Nature. [↩] [↩] [↩] [↩]
- Hanahan, D. & Weinberg, R. A. 2011. Hallmarks of cancer: The next generation. Cell. [↩] [↩] [↩] [↩]
- Madhusoodanan, R., et al. 2020. Systems biology of cancer: A review. Journal of Molecular Oncology. [↩] [↩]
- Sun, M., et al. 2022. The role of omics technologies in cancer research: Trends and challenges. Cancer Genomics & Proteomics. [↩] [↩]
- Karin, M. & Greten, F. R. 2005. NF-κB: Linking inflammation and immunity to cancer development and progression. Nature Reviews Immunology. [↩] [↩]
- Feinberg, A. P., Koldobskiy, M. A. & Jirtle, R. L. 2006. Epigenetic regulation of the genome: Implications for cancer. Annual Review of Genomics and Human Genetics. [↩] [↩]
- Begg, C. B., et al. 2018. The future of personalized medicine in oncology. JAMA Oncology. [↩]
- Vogelstein, B. & Kinzler, K. W. 2015. Cancer genes and the pathways they control. Nature Medicine. [↩]
- Feinberg, A. P., Koldobskiy, M. A. & Jirtle, R. L. 2006. Epigenetic regulation of the genome: Implications for cancer. Annual Review of Genomics and Human Genetics. [↩]
- Al-Hajj, M., et al. 2003. Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences. [↩]
- Navin, N. E. 2015. The first five years of single-cell cancer genomics. Nature Reviews Cancer. [↩]