Abstract
Skeletal muscle injury leads to severe functional loss, and current treatments remain limited. In this study, we developed an innovative platform to engineer functional muscle patches by mimicking the natural microenvironment. We combined (i) a three-dimensional GelMA lattice scaffold to enhance nutrient diffusion and cell alignment and (ii) fibroblast-derived ECM priming to promote adhesion and differentiation. Under the combined ECM and 3D hydrogel lattice conditions, C2C12 attachment rates significantly increased, and long, parallel multinucleated myotubes formed along the lattice structure. Molecular analysis revealed strong upregulation of MyoD, Myogenin, and MRF4. Functionally, wound healing experiments demonstrated over 80% healing within 24 hours, and calcium imaging revealed synchronized contraction signals. This study presents an advanced in vitro muscle engineering platform that enhances myotube alignment and myogenic differentiation through the integration of a 3D hydrogel lattice scaffold and fibroblast-derived ECM priming. This system provides a foundation for future applications in implantable muscle patches and drug screening.
Keywords: 3D hydrogel lattice, ECM, 3T3, C2C12, muscle cell
Introduction
Skeletal muscle tissue performs functions essential for physical activity and metabolism1,2 but, but it is easily damaged due to trauma, aging3, and pathological conditions such as Duchenne Muscular Dystrophy (DMD)4,5. Such severe muscle damage often exceeds the tissue’s spontaneous regenerative capacity6, leading to functional loss7. Current treatments face limitations like transplant rejection and restricted functional recovery8,9. As an alternative to overcome these problems, research on muscle tissue reconstruction based on tissue engineering has been actively pursued10,11.
Conventional in vitro skeletal muscle models are predominantly based on two-dimensional (2D) planar cultures, which fail to recapitulate the three-dimensional architecture, mechanical constraints, and biochemical complexity of native muscle tissue12. In such environments, myoblast fusion and myotube alignment are often limited, resulting in structurally disorganized and functionally immature muscle constructs13.
More recently, three-dimensional (3D) hydrogel-based culture systems have been introduced to overcome these limitations14. However, many bulk hydrogel models still suffer from insufficient structural guidance for cell alignment and limited integration of biochemical cues that are essential for muscle maturation15. As a result, although these models improve cellular encapsulation, they often fail to generate uniformly aligned and mature myotubes16.
To address these limitations, there is a need for in vitro muscle models that integrate both physical and biochemical microenvironmental cues. Structural guidance that promotes anisotropic alignment, together with biologically relevant extracellular matrix (ECM) signals, is critical for driving efficient myoblast fusion and maturation17. However, platforms that effectively combine these elements in a reproducible and modular manner remain limited18.
This study aims to construct a functional muscle patch by precisely mimicking the in vivo microenvironment of muscle tissue. To this end, we integrated two core strategies. First, we fabricated a 3D GelMA hydrogel scaffold with a lattice architecture to provides structural guidance that promotes myotube alignment and organization, while ECM priming enhances myogenic differentiation. Second, we applied fibroblast-derived extracellular matrix (ECM) priming to provide a biomimetic environment that assists in muscle cell attachment and maturation. This research goes beyond a simple in vitro model, presenting the potential of an innovative, implantable muscle patch and drug screening platform for muscle damage regeneration.
Literature Review
Skeletal Muscle Injury and Limitations of Current Regenerative Approaches
Despite the possibility of some limited intrinsic regeneration of skeletal muscle through satellite cells19 in situations of extreme traumatic injury, degenerative ageing or associated with some pathological conditions such as Duchenne muscular dystrophy (DMD), the regenerative capacity is nevertheless inadequate to replace the volumes of muscle that have been permanently compromised, leading to loss of function. Standard current approaches to skeletal muscle reconstruction including surgical reconstruction, autologous muscle grafts and stem-cell transplantations frequently result in poor prognosis in the clinic due to immune rejection, donor-site morbidity, poor engraftment efficiency and incomplete functional recovery8.
The considerable limitations of skeletal muscle regeneration have led to increased interest in tissue engineered muscle constructs in regenerative medicine. Such constructs aim to replace the cellular component of muscles as well as their architecture and contractile properties. However, the generation of functional mature muscle tissue has been difficult due to the complexity of the muscle microenvironment and the challenges of recapitulating it in culture20.
Limitations of Conventional 2D Muscle Cell Culture Models
Historically, most skeletal muscle studies have been performed in vitro using 2D planar cell culture systems. While these cultures are simple to set up and allow for repeatable experimental results, they fail to mimic the native physiological environment of the skeletal muscle due to their lack of structure and specific biochemical signals. Myoblasts in 2D cultures frequently display aberrant morphology, fusion deficiencies, poor alignment, and generate immature myotubes with sub-optimal cellular functions21.
Prolonged culture in 2D will lead to partial myogenic differentiation; however, 2D culture systems produce myotubes that lack intrinsic fiber-type specification and maturity compared with muscle fibers in vivo22. In addition, 2D culture systems provide limited cell-cell and matrix-cell interactions23. Both of these extracellular interactions have been reported to be important in myogenic regulation24; therefore, there is a strong effort to develop 3D culture systems (i.e., culture systems that mimic in vivo muscle tissues) to study myogenesis.
Advances and Challenges in 3D Hydrogel-Based Muscle Tissue Engineering
Three-dimensional (3D) hydrogel scaffolds with tunable mechanical properties, high water content and biocompatibility have been tested in the field of skeletal muscle tissue engineering25. The hydrogels collagens, fibrin, alginates and gelatin methacryloyl (GelMA) have been utilized for encapsulation of myoblasts to foster their muscle tissue differentiation16. Of these, GelMA is particularly attractive as, besides containing the cell adhesive motif of gelatin, the stiffness and architecture can be easily manipulated by photopolymerization26.
However, conventional bulk hydrogels suffer from some limitations: O2 and nutrient diffusion is limited in dense hydrogels, resulting in low oxygen tension and low cell viability, especially in thicker constructs27. Furthermore, bulk hydrogels do not provide spatial cues for aligned myotubes, resulting in randomized orientation of muscle fibers and poor contractile function of the muscle construct28. To address this limitation, recent studies have incorporated structured or patterned hydrogels (microchannels, aligned fibers, or lattice structures) to promote diffusion and preferential cell orientation within the matrix29.
Role of Extracellular Matrix in Myogenic Differentiation
Components of ECM (collagen, fibronectin, laminin etc.) are also involved in ECM-mediated signaling pathways that may define cell behavior by regulating myoblast adhesion, proliferation, and differentiation during development, regeneration, and maturation of skeletal muscle30. These factors are secreted by fibroblasts present in muscle tissue, which also represent the major cells responsible for the maintenance of the regenerative niche in skeletal muscle.
Previous reports have shown that fibroblast-synthesized ECM can initiate myogenic differentiation without MRF expression, indicating that cell matrix interactions are important determinants of myogenesis31. ECM proteins assist in focal adhesion assembly and cytoskeletal organization and may also exert their effects by modulating downstream signaling pathways that affect myotube fusion and maturation. However, many heterotypic in vitro muscle models use simplified or artificial ECM coatings that do not accurately recapitulate the complex composition of fibroblast-derived ECM32.
Although ECM priming strategies that pre-coat hydrogels with endogenous ECM secreted by fibroblasts have been shown to improve myoblast attachment and differentiation33, these strategies have mostly been adopted in 2D cultures and therefore may not accurately recapitulate the ECM architecture of the native 3D microenvironment of skeletal muscle.
Need for Integrated Structural and Biochemical Muscle Engineering Platforms
However, many of the most successful approaches to 3D muscle tissue engineering still rely on either mechanical or biochemical cues34, with structured scaffolds maintaining the alignment of muscle cells, but lacking biochemical factors35. Likewise, ECM-based strategies enable improved differentiation but lack spatial organization36. Therefore, approaches that combine structural guidance and biologically relevant ECM signaling on platforms are still needed.
While engineered 3D structures and cell-derived ECM have been used as independent in vitro models, more recent works suggest that the combination of both may better recapitulate the native muscle microenvironment and improve nutrient diffusion, cell alignment, myogenic differentiation, and functional maturation. However, the systematic evaluation of these combined strategies is limited37.
Rationale for the Present Study
To overcome these limitations, we posit the use of a 3D GelMA lattice scaffold using fibroblast-derived ECM priming to develop a muscle tissue engineering platform. By utilizing a physically aligned, diffusive hydrogel lattice, combined with biologically active ECM components, the microenvironment of our muscle tissue engineering platform can better recapitulate the physiological conditions seen in vivo and thus can be applied to muscle development and regeneration. This allows the generation of fully functional and mature muscle patches for use in regenerative medicine and drug testing applications.
Materials and Methods
Preparation of 3D GelMA hydrogel lattice scaffold
Gelatin methacryloyl (GelMA, 10%, w/v) was synthesized following standard protocols and dissolved in DPBS containing 0.05% (w/v) lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) as a photoinitiator. A GelMA concentration of 10% (w/v) was used throughout the study. This concentration was chosen to ensure sufficient mechanical integrity for stable lattice fabrication while maintaining a stiffness range reported to be permissive for myogenic differentiation.
Elastic molds were CAD-designed and fabricated using 3D printing (Formlabs, USA) with dimensions compatible with a 6-well plate. Each mold contained lattice patterns with a strut width of 1.0 mm, spacing of 2.0 mm, and thickness of 1.5 mm. GelMA solution was poured into the molds and photo-crosslinked under UV light (405 nm, 10 mW/cm![]() ) for 30 s. The cured hydrogel lattices were carefully detached from the molds and stored in sterile DPBS until use.
) for 30 s. The cured hydrogel lattices were carefully detached from the molds and stored in sterile DPBS until use.
To distinguish the effects of scaffold geometry from GelMA chemistry, flat GelMA and bulk GelMA controls were included. Flat GelMA substrates were prepared by casting a thin layer of GelMA on tissue culture plates followed by photocrosslinking. Bulk GelMA constructs were fabricated as non-patterned, continuous hydrogels using the same GelMA formulation and crosslinking conditions as the lattice scaffolds. All GelMA-based conditions shared identical polymer concentration and photoinitiator composition.
Cell culture and ECM priming
C2C12 murine myoblasts (ATCC, CRL-1772) and NIH3T3 fibroblasts (ATCC, CRL-1658) were cultured in Dulbecco’s Modified Eagle Medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (FBS, Gibco) and 1% penicillin/streptomycin at 37 ![]() C with 5% CO
C with 5% CO![]() .
.
For ECM priming, NIH3T3 fibroblasts were seeded onto either 2D culture surfaces or 3D GelMA lattices and cultured for 5 days to allow deposition of ECM proteins (e.g., collagen I, fibronectin). To prevent fibroblast overgrowth, cells were treated with mitomycin-C (10 ![]() g/mL, 2 h) prior to C2C12 seeding.
g/mL, 2 h) prior to C2C12 seeding.
Myoblast seeding and differentiation
C2C12 myoblasts were seeded at a density of ![]() cells per well (6-well) for flat controls or
cells per well (6-well) for flat controls or ![]() cells per lattice scaffold. After 24 h in growth medium, cultures were switched to differentiation medium (DMEM supplemented with 2% horse serum) to induce myogenic differentiation. Medium was replaced daily for up to 4 days.
cells per lattice scaffold. After 24 h in growth medium, cultures were switched to differentiation medium (DMEM supplemented with 2% horse serum) to induce myogenic differentiation. Medium was replaced daily for up to 4 days.
Electrical stimulation was applied at 1 Hz with a field strength of 5 V/cm and a pulse duration of 2 ms for 1 h per day. These parameters were selected based on previous reports showing that low-frequency electrical stimulation supports myogenic differentiation and calcium responsiveness while minimizing cellular stress and cytotoxic effects in C2C12 myotubes.
Immunofluorescence staining
Cells were fixed in 4% paraformaldehyde for 15 min, permeabilized with 0.1% Triton X-100, and blocked with 5% BSA. Samples were incubated overnight with primary antibodies against Myosin Heavy Chain (MHC, MF20, DSHB, 1:200) and ![]() -actinin (Sigma, 1:200), followed by Alexa Fluor-conjugated secondary antibodies (Invitrogen, 1:500). Nuclei were counterstained with DAPI (1
-actinin (Sigma, 1:200), followed by Alexa Fluor-conjugated secondary antibodies (Invitrogen, 1:500). Nuclei were counterstained with DAPI (1 ![]() g/mL). Images were acquired using a confocal microscope (Zeiss LSM800) and analyzed with ImageJ. Fusion Index (FI, % of nuclei within MHC
g/mL). Images were acquired using a confocal microscope (Zeiss LSM800) and analyzed with ImageJ. Fusion Index (FI, % of nuclei within MHC![]() myotubes with
myotubes with ![]() 2 nuclei/total nuclei) and Orientation Order Parameter (OOP, 0–1 scale) were quantified from
2 nuclei/total nuclei) and Orientation Order Parameter (OOP, 0–1 scale) were quantified from ![]() 5 random fields.
5 random fields.
Gene expression analysis (RT-qPCR)
Total RNA was extracted using TRIzol (Invitrogen), and cDNA was synthesized using a reverse transcription kit (Thermo Scientific). Quantitative PCR was performed using SYBR Green Master Mix (Bio-Rad) with primers specific for MyoD, Myogenin, MRF4, and GAPDH as an internal control. Relative expression was calculated using the ![]() method.
method.
| Forward (5’->3’) | Reverse (5’->3’) | |
| MyoD | GATGCACCGGTTATTTGGGG | ATTCCCTGTAGCACCACACA |
| Myogenin | AAGAAGGGGAGAGGAACAGC | GCAACTTCAGCACAGGAGAC |
| MRF4 (Myf6) | GTGGAGGAAGTGGTGGAGAA | GCATGCACAACAGATTTCGC |
Table 1 | Primer sequences of RT-qPCR
Western blotting
Proteins were extracted with RIPA buffer containing protease inhibitors, separated by SDS-PAGE, and transferred to PVDF membranes. Membranes were blocked with 5% skim milk and probed with primary antibodies against MyoD(Santa Cruz; sc-32758), Myogenin(Santa Cruz; sc-12732), MRF4(Invitrogen; PA5-97990), and β-actin (Santa Cruz; sc-8432). HRP-conjugated secondary antibodies were used, and bands were visualized by ECL detection (Bio-Rad). Densitometry analysis was performed using ImageJ.
Wound healing assay
A scratch wound was generated using a sterile 200 µL pipette tip across confluent C2C12 cultures (flat, 2D ECM primed, 3D lattice ECM primed). Images were captured at 0 h and 24 h, and wound area was quantified using ImageJ. Relative wound closure (%) was calculated as:
![]()
Calcium imaging
Calcium imaging was performed to compare relative calcium signal intensity across myotubes under different culture conditions. Cells were loaded with Fluo-4 AM, and fluorescence images were acquired at a single time point during electrical stimulation. Mean fluorescence intensity was quantified using ImageJ to assess calcium responsiveness. Non-stimulated control samples were prepared for each condition. However, in the absence of electrical stimulation, calcium fluorescence signals remained near baseline levels and were close to the detection limit. Therefore, quantitative comparisons of calcium responses were performed under electrically stimulated conditions.
Statistical analysis
All experiments were independently repeated at least three times. For all quantitative analyses, n denotes the number of independent biological replicates. Data are presented as mean ± standard deviation (SD). Statistical significance was determined using one-way ANOVA followed by Tukey’s post hoc test. A value of p < 0.05 was considered significant. All statistical summary tables used in this study are provided in the Supplementary Data.
Results
Fabrication and Characterization of 3D GelMA Hydrogel Lattice Scaffolds
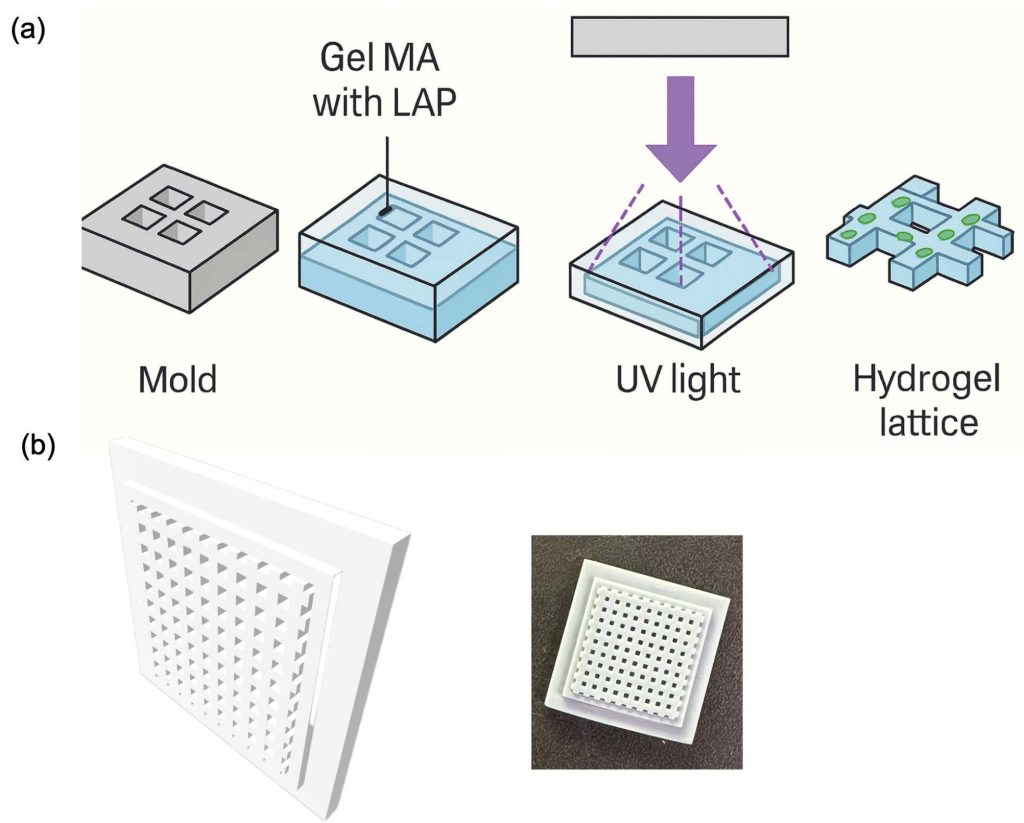
A CAD-designed elastic mold was 3D printed, and a release coating was applied to its surface to facilitate easy demolding of the GelMA (Figure. 1a and 1b). The fabricated scaffolds maintained excellent mechanical stability, making them suitable for long-term cell culture.
ECM Priming and Initial Cell Attachment
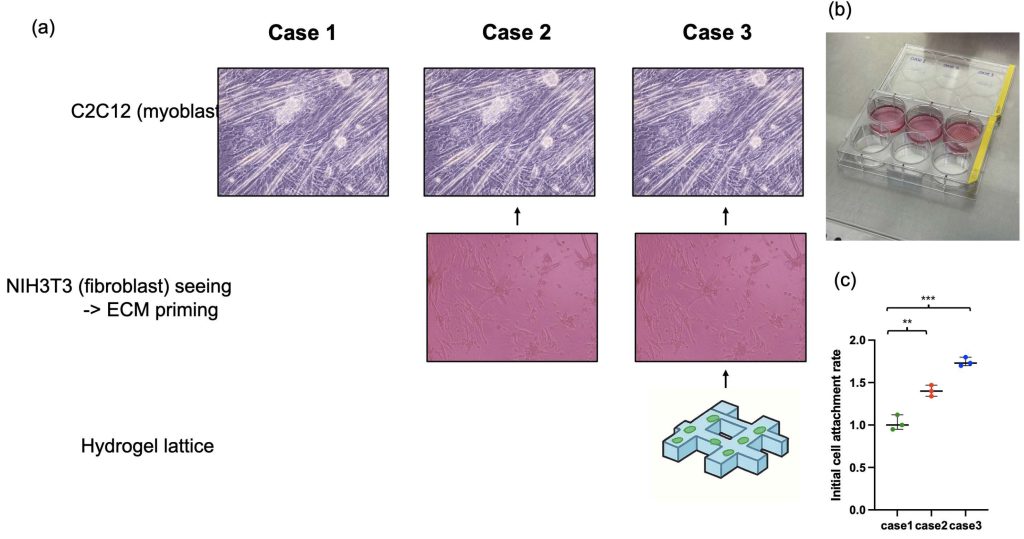
To precisely mimic the in vivo muscle microenvironment, we performed fibroblast-derived extracellular matrix (ECM) priming. This is because fibroblasts secrete various ECM proteins, such as collagen and fibronectin, which are important for muscle tissue maturation and maintenance. These ECM proteins not only act as a physical support for cell attachment and spreading but also serve as catalysts that provide biochemical signals influencing cell differentiation. We divided the experiment into three conditions to investigate the ECM priming capability of fibroblasts and the efficacy of our fabricated 3D hydrogel lattice scaffold (Figure. 2a and 2b). The first condition was seeding only myoblasts (C2C12) without ECM priming. The second condition was inducing fibroblasts (3T3) to naturally secrete ECM by seeding them in a 2D environment for 5 days before seeding C2C12. The third condition was culturing 3T3 cells first on the 3D hydrogel lattice we made to prime the ECM, and then seeding C2C12. After culturing 3T3 for ECM priming, we treated the cells with mitomycin-c to inhibit 3T3 proliferation, preventing complex growth competition between muscle cells and fibroblasts and creating a pure ECM environment. The initial cell attachment rate of C2C12 cells in ECM-primed Case 2 was about 1.4 times higher than in Case 1 (Flat), and it increased to 1.7 times in Case 3 (Figure. 2c). This proves that the ECM secreted by fibroblasts successfully provided an environment favorable for the attachment and survival of myoblasts.
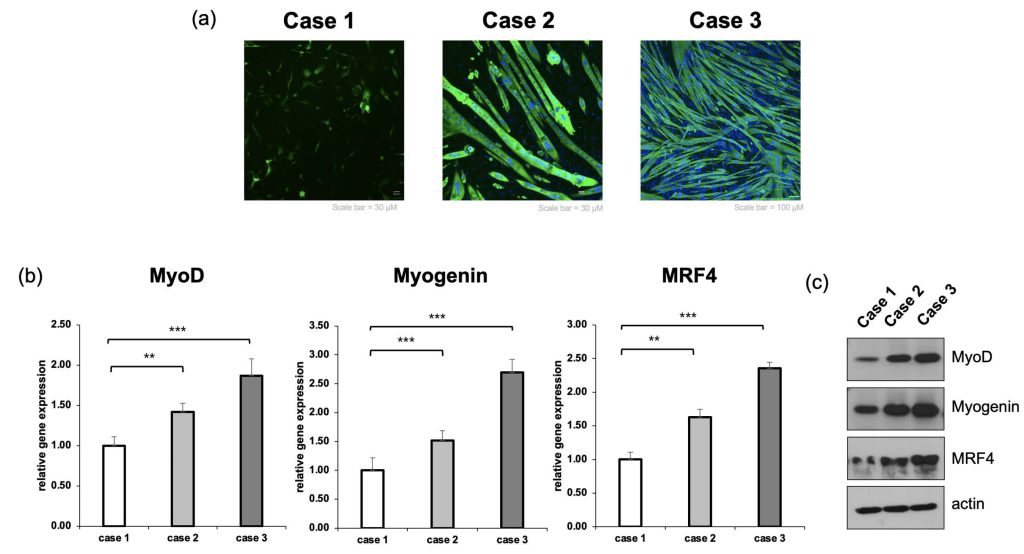
Muscle Cell Differentiation and Molecular Characterization
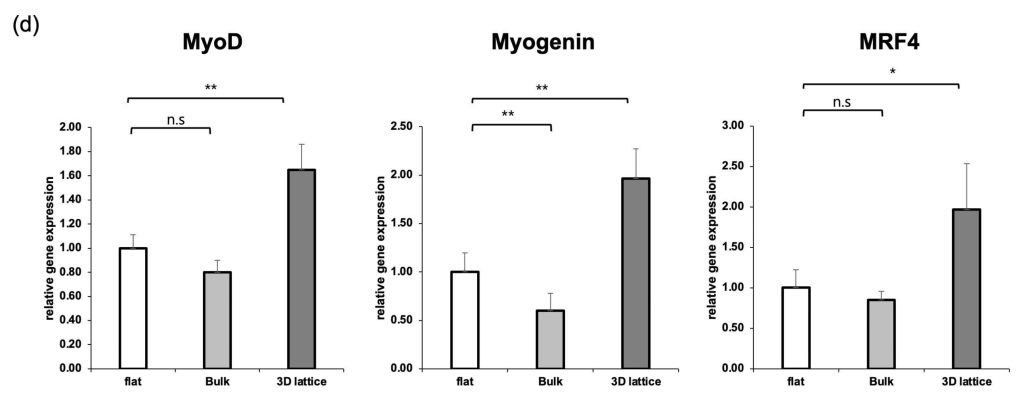
To verify muscle cell differentiation at the morphological and molecular levels, we performed immunofluorescence staining, RT-qPCR, and Western blot analysis. First, MHC immunofluorescence staining results showed that most single-nucleus cells were observed in Case 1, and the formed myotubes were short and arranged in an irregular direction. In contrast, in the 2D condition with ECM priming (Case 2), multinucleated myotubes were distinctly increased, and cell-to-cell connections were strengthened, leading to a significant improvement in the length and thickness of myotubes compared to the Flat condition. Particularly, the most prominent results were observed in the condition where ECM priming was applied on the 3D lattice (Case 3). In Case 3, myotubes grew the longest, and a clear structure of parallel alignment along the direction of the lattice was observed (Figure. 3a). Along with morphological analysis, RT-qPCR confirmed the expression of muscle differentiation-related transcription factors. In the ECM-primed Case 2, the expression of MyoD, Myogenin, and MRF4 increased by approximately 1.4, 1.51, and 1.63 times, respectively, compared to the Flat condition. In Case 3, which combined the 3D lattice and ECM priming, the expression of these genes increased even more significantly, with MyoD, Myogenin, and MRF4 rising by up to 1.87, 2.69, and 2.35 times, respectively (Figure. 3b) (n = 3 independent experiments; one-way ANOVA followed by Tukey’s post hoc test). These results suggest that when ECM and 3D structures are combined, muscle cell differentiation and maturation signals are strongly induced. Furthermore, Western blot analysis confirmed a similar trend at the protein level. The expression of MyoD, Myogenin, and MRF4 proteins showed an increasing tendency from Case 1 to Case 3 (Figure. 3c). Among the GelMA-based conditions, lattice-structured scaffolds consistently exhibited higher expression of MyoD, Myogenin, and MRF4 compared to flat and bulk GelMA controls (Figure. 3d).This supported the finding that muscle cell fusion and maturation were greatly enhanced in the ECM-primed and lattice scaffold environment.
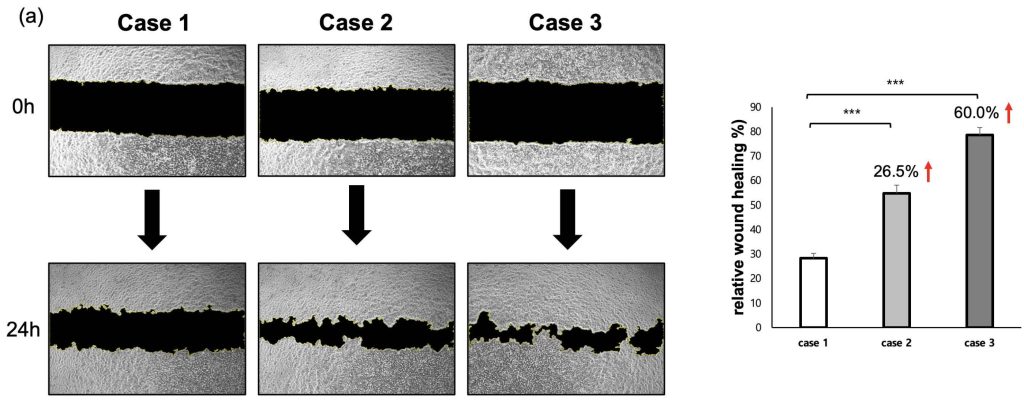
Verification of Regenerative and Functional Properties of the Muscle Patch
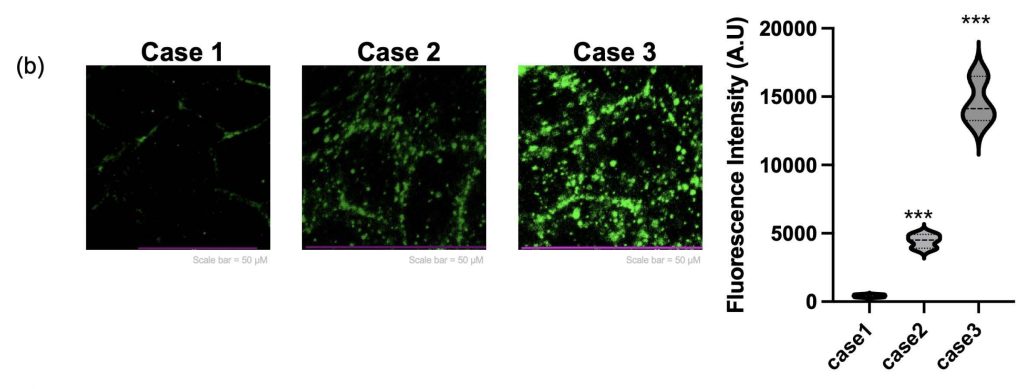
To evaluate cell migratory behavior, a scratch wound assay was performed under each experimental condition. In the Flat condition, the damage area induced by the scratch remained over 70% even after 24 hours, and the recovery speed was relatively slow. In contrast, the damage recovery speed in Case 2 increased by 26.5% compared to Case 1. Notably, the fastest healing ability, exceeding 80%, was observed in Case 3 (Figure 4a). These results are associated with fibroblast-derived ECM priming and indicate enhanced myotube morphology and myogenic gene expression under these conditions. Additionally, to verify that the patch not only has regenerative ability but also possesses functional properties similar to actual muscle, we performed Calcium Imaging. Calcium fluorescence intensity was quantified at a single time point during electrical stimulation to compare relative calcium responsiveness across conditions. Particularly, the lattice+ECM group showed a signal accompanied by strong contractile twitch (Figure 4b), proving that it mimics the electrophysiological properties of muscle tissue.
Discussion
This study successfully demonstrated the potential for developing a functional muscle patch by combining a 3D hydrogel lattice scaffold with fibroblast-derived extracellular matrix (ECM) priming as a new therapeutic strategy for muscle damage. The attempt to precisely mimic the structural and biochemical microenvironment of actual muscle tissue, overcoming the limitations of conventional 2D planar culture and simple bulk 3D models, led to very meaningful results. The lattice-structured GelMA scaffold we fabricated provided an optimal physical environment for muscle cell survival and differentiation. Unlike simple bulk hydrogels, the lattice structure offered a large surface area, enabling efficient diffusion of oxygen and nutrients38,39. Furthermore, the regular struts and spacing of the scaffold guided cells to align naturally along their direction. As shown by the immunofluorescence staining results, this played a crucial role in the formation of long, parallelly aligned multinucleated myotubes. Such structural alignment is an essential factor for restoring the functional properties, such as contractility, of muscle tissue. Fibroblast-derived ECM priming played a core role in creating a biomimetic environment necessary for muscle cell differentiation and maturation40,41. Various ECM proteins (collagen, fibronectin, etc.) secreted by fibroblasts significantly increased the initial attachment and survival rates of myoblasts42. This reconfirmed that the interaction between cells and the matrix is a crucial signal that determines the cell’s fate43,44. In particular, the significant increase in the expression of muscle differentiation-related genes (MyoD, Myogenin, MRF4) in RT-qPCR and Western blot analysis suggests that ECM signals strongly promote the cell’s differentiation pathway. Furthermore, among the tested conditions, the lattice-guided and ECM-primed constructs (case 3) consistently showed higher myogenic marker expression. The physical lattice structure provided the ‘direction’ for cell alignment, while the biochemical ECM signals promoted the ‘speed’ and ‘intensity’ of cell differentiation, thereby constructing an optimized environment for muscle tissue reconstruction. This was clearly demonstrated in the wound healing assay and calcium imaging results. The Case 3 group showed the fastest wound recovery speed and synchronized calcium signals, suggesting that it can secure functional viability similar to actual muscle tissue, going beyond mere cell survival. However, it should be noted that the scratch assay primarily reflects cell migration and proliferation in an in vitro setting and does not represent true skeletal muscle regeneration. Therefore, the observed wound closure is interpreted as enhanced migratory behavior rather than tissue repair. Also, it should be noted that calcium imaging was performed at a single time point, which does not allow direct assessment of synchronous contraction or excitation–contraction coupling dynamics. In addition, measurements of contractile force, twitch kinetics, or time-resolved calcium transients were not included in this study. These functional assessments will be essential in future work to further validate muscle-like behavior of the engineered constructs. The 3D lattice hydrogel scaffold-based muscle tissue engineering platform developed in this study presents an innovative method for the successful in vitro reconstruction of muscle tissue. Although the engineered constructs exhibited improved myotube alignment and increased expression of myogenic markers, this study does not demonstrate the formation of a mechanically integrated, force-generating muscle tissue. Together, these findings suggest that the lattice-structured GelMA scaffold, in combination with fibroblast-derived ECM priming, provides a microenvironment that supports myoblast organization and differentiation in vitro. However, the present data do not demonstrate the formation of a fully functional, force-generating muscle tissue. Nevertheless, this platform can be utilized in the future as a drug screening model for diseases like muscular dystrophy. Future studies should focus on further in vitro validation of functional properties, including measurements of contractile force, time-resolved calcium transients, and refined electrical stimulation protocols. These steps will be essential to better characterize muscle-like behavior before considering more advanced applications. In summary, this study presents an advanced in vitro platform for investigating how structural and biochemical cues influence myogenic differentiation, while highlighting the need for additional functional validation in future work.
Supplementary Tables
| case 1 | case 2 | case 3 | |
| initial cell attachment rate | 1 | 1.4 | 1.7 |
| 1.12 | 1.47 | 1.8 | |
| 0.95 | 1.34 | 1.73 |
| Myo D relative expression | Myogenin relative expression | Mrf4 relative expression | |||||||
| expression | case 1 | case 2 | case 3 | case 1 | case 2 | case 3 | case 1 | case 2 | case 3 |
| average | 1 | 1.42 | 1.866666667 | 1 | 1.513333333 | 2.693333333 | 1.003333333 | 1.63 | 2.353333333 |
| SD | 0.11 | 0.108166538 | 0.21234 | 0.213 | 0.17 | 0.23 | 0.105039675 | 0.117898261 | 0.089628864 |
| Mrf4 relative expression | Mrf4 relative expression | Mrf4 relative expression | |||||||
| expression | flat | bulk | 3d lattice | flat | bulk | 3d lattice | flat | bulk | 3d lattice |
| average | 1 | 0.8 | 1.65 | 1 | 0.6 | 1.966666667 | 1.003333333 | 0.85 | 1.966666667 |
| SD | 0.11 | 0.1 | 0.2322 | 0.197 | 0.18 | 0.305505046 | 0.221 | 0.11 | 0.56862407 |
| case 1 | case 2 | case 3 | |
| relative wound healing (%) | 26.217677 | 57.68087539 | 75.27458319 |
| 28.435 | 51.234 | 79.13 | |
| 30.123 | 55.4 | 81.2 | |
| average | 28.258559 | 54.77162513 | 78.53486106 |
| SD | 1.958631029 | 3.269050596 | 3.007205323 |
| case 1 | case 2 | case 3 | |
| fluorescence intensity (A.U.) | 326 | 4919 | 16500 |
| 424 | 3913 | 13255 | |
| 519 | 4500 | 14119 | |
| average | 423 | 4444 | 14624.66667 |
| SD | 96.50388593 | 505.3325638 | 1680.55953 |
References
- Egan, B., Zierath, Juleen R. Exercise Metabolism and the Molecular Regulation of Skeletal Muscle Adaptation. Cell Metabolism 17, 162-184 (2013). [↩]
- Severinsen, M.C.K., Pedersen, B.K. Muscle–Organ Crosstalk: The Emerging Roles of Myokines. Endocrine Reviews 41, 594-609 (2020). [↩]
- Kirkendall, D.T. & Garrett, W.E. The Effects of Aging and Training on Skeletal Muscle. The American Journal of Sports Medicine 26, 598-602 (1998). [↩]
- Godi, C., et al. Longitudinal MRI quantification of muscle degeneration in Duchenne muscular dystrophy. Annals of Clinical and Translational Neurology 3, 607-622 (2016). [↩]
- De Paepe, B. & De Bleecker, J.L. Cytokines and Chemokines as Regulators of Skeletal Muscle Inflammation: Presenting the Case of Duchenne Muscular Dystrophy. Mediators of Inflammation 2013, 540370 (2013). [↩]
- Corona, B.T., Ward, C.L., Baker, H.B., Walters, T.J. & Christ, G.J. Implantation of In Vitro Tissue Engineered Muscle Repair Constructs and Bladder Acellular Matrices Partially Restore In Vivo Skeletal Muscle Function in a Rat Model of Volumetric Muscle Loss Injury. Tissue Engineering Part A 20, 705-715 (2014). [↩]
- Kern, H., et al. Home-Based Functional Electrical Stimulation Rescues Permanently Denervated Muscles in Paraplegic Patients With Complete Lower Motor Neuron Lesion. Neurorehabilitation and Neural Repair 24, 709-721 (2010). [↩]
- Maffioletti, S.M., Noviello, M., English, K. & Tedesco, F.S. Stem Cell Transplantation for Muscular Dystrophy: The Challenge of Immune Response. BioMed Research International 2014, 964010 (2014). [↩] [↩]
- Greising, S.M., Dearth, C.L. & Corona, B.T. Regenerative and Rehabilitative Medicine: A Necessary Synergy for Functional Recovery from Volumetric Muscle Loss Injury. Cells Tissues Organs 202, 237-249 (2016). [↩]
- Samandari, M., et al. Bioinks and Bioprinting Strategies for Skeletal Muscle Tissue Engineering. Advanced Materials 34, 2105883 (2022). [↩]
- Gilbert-Honick, J. & Grayson, W. Vascularized and Innervated Skeletal Muscle Tissue Engineering. Advanced Healthcare Materials 9, 1900626 (2020). [↩]
- Gao, Y., et al. Engineering of tissue in microphysiological systems demonstrated by modelling skeletal muscle. Regenerative Biomaterials 12(2025). [↩]
- Afshar Bakooshli, M., et al. A 3D culture model of innervated human skeletal muscle enables studies of the adult neuromuscular junction. eLife 8, e44530 (2019). [↩]
- Smith, A.S.T., Passey, S., Greensmith, L., Mudera, V. & Lewis, M.P. Characterization and optimization of a simple, repeatable system for the long term in vitro culture of aligned myotubes in 3D. Journal of Cellular Biochemistry 113, 1044-1053 (2012). [↩]
- Salimath, A.S. & García, A.J. Biofunctional hydrogels for skeletal muscle constructs. Journal of Tissue Engineering and Regenerative Medicine 10, 967-976 (2016). [↩]
- Costantini, M., et al. Engineering Muscle Networks in 3D Gelatin Methacryloyl Hydrogels: Influence of Mechanical Stiffness and Geometrical Confinement. Frontiers in Bioengineering and Biotechnology Volume 5 – 2017(2017). [↩] [↩]
- Jana, S., Levengood, S.K.L. & Zhang, M. Anisotropic Materials for Skeletal-Muscle-Tissue Engineering. Advanced Materials 28, 10588-10612 (2016). [↩]
- Raman, R., Cvetkovic, C. & Bashir, R. A modular approach to the design, fabrication, and characterization of muscle-powered biological machines. Nature Protocols 12, 519-533 (2017). [↩]
- Motohashi, N. & Asakura, A. Muscle satellite cell heterogeneity and self-renewal. Frontiers in Cell and Developmental Biology Volume 2 – 2014(2014). [↩]
- Jin, Y., et al. Functional Skeletal Muscle Regeneration with Thermally Drawn Porous Fibers and Reprogrammed Muscle Progenitors for Volumetric Muscle Injury. Advanced Materials 33, 2007946 (2021). [↩]
- Wang, P.-Y., Yu, H.-T. & Tsai, W.-B. Modulation of alignment and differentiation of skeletal myoblasts by submicron ridges/grooves surface structure. Biotechnology and Bioengineering 106, 285–294 (2010). [↩]
- Cooper, S.T., et al. C2C12 Co-culture on a fibroblast substratum enables sustained survival of contractile, highly differentiated myotubes with peripheral nuclei and adult fast myosin expression. Cell Motility 58, 200–211 (2004). [↩]
- Habanjar, O., Diab-Assaf, M., Caldefie-Chezet, F. & Delort, L. 3D Cell Culture Systems: Tumor Application, Advantages, and Disadvantages. International Journal of Molecular Sciences 22, 12200 (2021). [↩]
- Mayer, U. Integrins: Redundant or Important Players in Skeletal Muscle?*. Journal of Biological Chemistry 278, 14587–14590 (2003). [↩]
- Wang, L., Wu, Y., Guo, B. & Ma, P.X. Nanofiber Yarn/Hydrogel Core–Shell Scaffolds Mimicking Native Skeletal Muscle Tissue for Guiding 3D Myoblast Alignment, Elongation, and Differentiation. ACS Nano 9, 9167–9179 (2015). [↩]
- Chansoria, P., et al. Characterizing the Effects of Synergistic Thermal and Photo-Cross-Linking during Biofabrication on the Structural and Functional Properties of Gelatin Methacryloyl (GelMA) Hydrogels. ACS Biomaterials Science & Engineering 7, 5175–5188 (2021). [↩]
- Figueiredo, L., et al. Assessing glucose and oxygen diffusion in hydrogels for the rational design of 3D stem cell scaffolds in regenerative medicine. Journal of Tissue Engineering and Regenerative Medicine 12, 1238–1246 (2018). [↩]
- De France, K.J., et al. Injectable Anisotropic Nanocomposite Hydrogels Direct in Situ Growth and Alignment of Myotubes. Nano Letters 17, 6487–6495 (2017). [↩]
- Sornkamnerd, S., Okajima, M.K., Matsumura, K. & Kaneko, T. Micropatterned Cell Orientation of Cyanobacterial Liquid-Crystalline Hydrogels. ACS Applied Materials & Interfaces 10, 44834–44843 (2018). [↩]
- Docheva, D., Popov, C., Alberton, P. & Aszodi, A. Integrin signaling in skeletal development and function. Birth Defects Research Part C: Embryo Today: Reviews 102, 13–36 (2014). [↩]
- Osses, N. & Brandan, E. ECM is required for skeletal muscle differentiation independently of muscle regulatory factor expression. American Journal of Physiology-Cell Physiology 282, C383–C394 (2002). [↩]
- Cao, T. & Warren, C.R. From 2D Myotube Cultures to 3D Engineered Skeletal Muscle Constructs: A Comprehensive Review of In Vitro Skeletal Muscle Models and Disease Modeling Applications. Cells 14, 882 (2025). [↩]
- Boyd-Moss, M., et al. Hybrid Self-Assembling Peptide/Gelatin Methacrylate (GelMA) Bioink Blend for Improved Bioprintability and Primary Myoblast Response. Advanced NanoBiomed Research 2, 2100106 (2022). [↩]
- Bakhshandeh, B., et al. Recent progress in the manipulation of biochemical and biophysical cues for engineering functional tissues. Bioengineering & Translational Medicine 8, e10383 (2023). [↩]
- Bandara, G.C., Boudreau, R.D., Wyatt, W. & Caliari, S.R. Heparin-modified aligned collagen scaffolds enhance <em>in vitro</em> myogenesis. bioRxiv, 2025.2010.2015.681268 (2025). [↩]
- Nakada, S. Myocyte culture with decellularized skeletal muscle sheet with observable interaction with extracellular matrix. [↩]
- Sala, A., et al. Engineering 3D cell instructive microenvironments by rational assembly of artificial extracellular matrices and cell patterning. Integrative Biology 3, 1102–1111 (2011). [↩]
- Axpe, E., et al. A Multiscale Model for Solute Diffusion in Hydrogels. Macromolecules 52, 6889-6897 (2019). [↩]
- Lavrentev, F.V., et al. Diffusion-Limited Processes in Hydrogels with Chosen Applications from Drug Delivery to Electronic Components. Molecules 28, 5931 (2023). [↩]
- Osses, N. & Brandan, E. ECM is required for skeletal muscle differentiation independently of muscle regulatory factor expression. American Journal of Physiology-Cell Physiology 282, C383-C394 (2002). [↩]
- Melo, F., Carey, D.J. & Brandan, E. Extracellular matrix is required for skeletal muscle differentiation but not myogenin expression. Journal of Cellular Biochemistry 62, 227-239 (1996). [↩]
- Uchinaka, A., et al. Laminin α2-secreting fibroblasts enhance the therapeutic effect of skeletal myoblast sheets. European Journal of Cardio-Thoracic Surgery 51, 457-464 (2016). [↩]
- Bi, Y., et al. Extracellular Matrix Proteoglycans Control the Fate of Bone Marrow Stromal Cells *. Journal of Biological Chemistry 280, 30481-30489 (2005). [↩]
- Chermnykh, E., Kalabusheva, E. & Vorotelyak, E. Extracellular Matrix as a Regulator of Epidermal Stem Cell Fate. International Journal of Molecular Sciences 19, 1003 (2018). [↩]